Sustained Gene Delivery with PEG-PLGA Nanoparticles: A Comprehensive Guide for Research and Development
This article provides a detailed, current analysis of PEG-PLGA nanoparticles for sustained gene release, tailored for researchers and drug development professionals.
Sustained Gene Delivery with PEG-PLGA Nanoparticles: A Comprehensive Guide for Research and Development
Abstract
This article provides a detailed, current analysis of PEG-PLGA nanoparticles for sustained gene release, tailored for researchers and drug development professionals. It explores the foundational science behind this drug delivery system, details advanced fabrication and characterization methodologies, and offers troubleshooting strategies for common formulation challenges. The content further covers critical validation techniques and comparative analyses against other non-viral vectors, synthesizing recent advancements to guide experimental design and translational development.
Understanding PEG-PLGA Nanoparticles: The Science Behind Sustained Gene Release
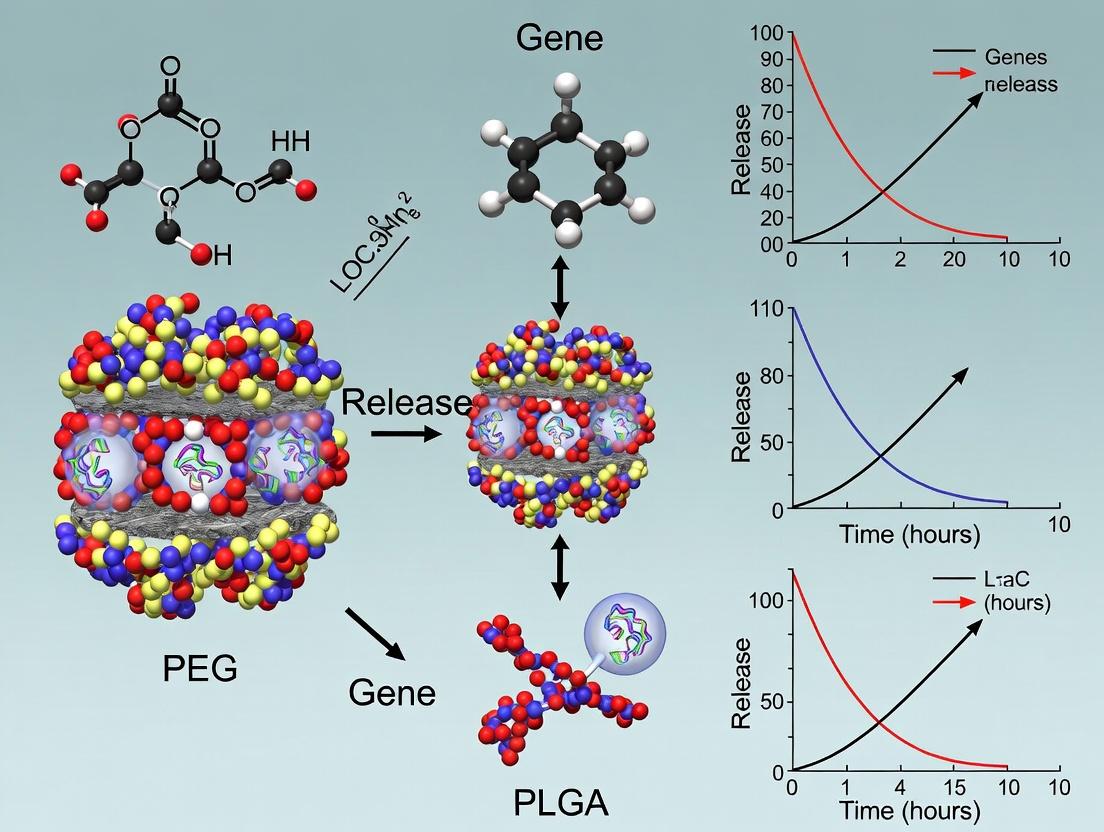
Polymeric nanoparticles (PNPs) represent a pivotal non-viral vector platform for gene therapy, designed to encapsulate and protect nucleic acids (pDNA, siRNA, mRNA) and facilitate their intracellular delivery. Within the specific context of a thesis focused on PEG-PLGA nanoparticles for sustained gene release, these systems offer a tunable, biodegradable, and biocompatible solution. The poly(lactic-co-glycolic acid) (PLGA) core enables controlled, sustained release of genetic payloads over days to weeks, while the poly(ethylene glycol) (PEG) corona ("PEGylation") enhances colloidal stability, prolongs systemic circulation, and reduces opsonization. This architecture is engineered to overcome key extracellular and intracellular barriers in gene delivery.
Table 1: Key Performance Metrics of Representative PEG-PLGA Nanoparticles for Gene Delivery
| Parameter | Typical Range/Value | Key Influencing Factors | Measurement Method |
|---|---|---|---|
| Particle Size | 80 - 200 nm | Polymer MW, PEG ratio, formulation method, nucleic acid loading | Dynamic Light Scattering (DLS) |
| Polydispersity Index (PDI) | < 0.2 (optimal) | Emulsion stability, purification method | DLS |
| Zeta Potential | Slightly negative to neutral (-10 to +5 mV) | PEG shielding, PLGA end groups, cationic adjuvant (e.g., PEI) presence | Electrophoretic Light Scattering |
| Encapsulation Efficiency (EE%) | 50 - 90% for pDNA/siRNA | Complexation method (adsorption vs. encapsulation), N/P ratio | Fluorescence/UV assay of supernatant |
| In Vitro Release Half-life | 3 - 14 days | PLGA lactide:glycolide ratio, polymer MW, nanoparticle porosity | Dialysis in PBS + serum at 37°C |
| In Vivo Circulation Half-life | 4 - 12 hours (mice/rats) | PEG density and length, particle size, surface charge | Fluorescent/blood assay over time |
Table 2: Comparison of Common Polymeric Vectors in Gene Therapy
| Polymer | Advantages | Disadvantages for Sustained Release | Best Suited For |
|---|---|---|---|
| PEG-PLGA | Biodegradable, predictable sustained release, excellent biocompatibility, FDA-approved components, tunable kinetics | Lower transfection efficiency vs. cationic polymers, complexation can be challenging | Sustained/controlled release applications, subcutaneous/intramuscular depot, systemic delivery with stealth |
| Polyethylenimine (PEI) | High transfection efficiency, strong nucleic acid complexation ("proton-sponge" effect) | High cytotoxicity, non-degradable, burst release profile | In vitro transfections, localized in vivo where high efficiency is critical |
| Chitosan | Biodegradable, low toxicity, mucoadhesive | Slow degradation, variable purity, weaker complexation strength, faster release than PLGA | Oral, mucosal, or topical gene delivery |
| Poly(β-amino esters) (PBAEs) | Degradable, high transfection efficiency, tunable structure | More rapid degradation than PLGA, less established for in vivo sustained release | Rapid, efficient transfection in dynamic environments |
Application Notes & Core Protocols
Protocol 3.1: Double Emulsion Solvent Evaporation for pDNA-Loaded PEG-PLGA NPs
Aim: To encapsulate plasmid DNA within a PEG-PLGA matrix for sustained release. Materials: PEG-PLGA copolymer (e.g., 5% PEG, 75:25 PLGA), pDNA solution (0.5 mg/mL in TE buffer), polyvinyl alcohol (PVA, 2% w/v), dichloromethane (DCM), ultrapure water. Procedure:
- Primary Emulsion: Add 1 mL of pDNA solution to 4 mL of DCM containing 100 mg of PEG-PLGA. Probe sonicate on ice (50 W, 30 s) to form a water-in-oil (W/O) emulsion.
- Double Emulsion: Rapidly pour the primary emulsion into 20 mL of 2% PVA solution under high-speed homogenization (10,000 rpm, 2 min) to form a (W/O)/W emulsion.
- Solvent Evaporation: Stir the double emulsion magnetically at room temperature for 4 hours to allow complete DCM evaporation and nanoparticle hardening.
- Purification: Centrifuge at 18,000 rpm for 30 min at 4°C. Wash pellet 3x with water to remove PVA and unencapsulated pDNA.
- Lyophilization: Resuspend in 5% trehalose (cryoprotectant) and lyophilize for 48h to obtain a stable powder.
Protocol 3.2: Characterization of Nanoparticles
A. Size & Zeta Potential: Resuspend lyophilized NPs in 1 mM KCl. Analyze using DLS and electrophoretic light scattering. Report Z-average diameter, PDI, and zeta potential (mean of n=3 measurements). B. Encapsulation Efficiency: Digest 5 mg of NPs in 1 mL of 0.5M NaOH for 2h. Neutralize with HCl. Quantify released pDNA using a Quant-iT PicoGreen assay against a standard curve. EE% = (Mass of DNA in NPs / Total initial mass of DNA) x 100. C. In Vitro Release Study: Place 10 mg of NPs in a dialysis bag (MWCO 100 kDa). Immerse in 50 mL PBS (pH 7.4, 0.1% w/v sodium azide) at 37°C with gentle shaking. At predetermined intervals, sample and replace the release medium. Quantify released pDNA via PicoGreen assay. Plot cumulative release (%) vs. time.
Protocol 3.3: In Vitro Transfection and Assessment
Cell Culture: Seed HEK293 or relevant cell line in 24-well plates at 50,000 cells/well. Transfection: Add NPs equivalent to 0.5 µg pDNA per well in serum-free medium. After 4h, replace with complete medium. Analysis: At 24-72h post-transfection, lyse cells to quantify transgene expression (e.g., luciferase assay, ELISA for encoded protein) and assess cell viability (MTT or AlamarBlue assay).
Visualizations
Diagram Title: Thesis Workflow for PEG-PLGA Gene Delivery Research
Diagram Title: How PEG-PLGA NPs Overcome Gene Delivery Barriers
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for PEG-PLGA Nanoparticle Gene Therapy Research
| Item (Example Supplier) | Function in Research | Critical Specification/Note |
|---|---|---|
| PEG-PLGA Copolymer (LACTEL Absorbable Polymers, Sigma-Aldrich) | Core matrix material forming the nanoparticle. Determines degradation rate, release kinetics, and biocompatibility. | Choose lactide:glycolide ratio (e.g., 50:50 fast, 75:25 slow) and PEG % (e.g., 2-10%) based on desired release profile. |
| PicoGreen dsDNA Quantitation Reagent (Invitrogen) | Ultra-sensitive fluorescent assay for quantifying pDNA encapsulation efficiency and in vitro release profiles. | Essential for working with low nucleic acid concentrations. More sensitive than A260. |
| Polyvinyl Alcohol (PVA, 87-89% hydrolyzed, Sigma-Aldrich) | Common stabilizer/surfactant in double emulsion formulations. Critical for controlling nanoparticle size and polydispersity. | Degree of hydrolysis and MW significantly impact nanoparticle characteristics. |
| Dialysis Membranes (Spectra/Por, 100 kDa MWCO) | Used for in vitro release studies to physically separate released pDNA from encapsulated nanoparticles. | MWCO must be large enough to allow free DNA passage but retain nanoparticles. |
| Trehalose Dihydrate (Pharmaceutical Grade, Pfanstiehl) | Cryoprotectant for lyophilization (freeze-drying) of nanoparticle suspensions to ensure long-term stability as a powder. | Prevents aggregation and maintains particle integrity during drying and reconstitution. |
| Bicinchoninic Acid (BCA) Protein Assay Kit (Pierce) | Quantifies transgene-encoded protein expression in cell lysates post-transfection, a key efficacy readout. | Compatible with detergent-containing cell lysis buffers. |
| Lipofectamine 3000 (Invitrogen) | Commercial cationic lipid transfection reagent. Used as a positive control in in vitro transfection experiments to benchmark PEG-PLGA NP efficiency. | Represents a high-efficiency, but non-sustained, standard for comparison. |
PLGA and PEG are foundational polymers in the development of nanoscale drug and gene delivery systems. Within the context of PEG-PLGA nanoparticles for sustained gene release, each polymer fulfills distinct, complementary roles.
PLGA Core Role: A biodegradable and biocompatible copolymer, PLGA forms the nanoparticle matrix. Its hydrolysis into lactic and glycolic acids allows for controlled, sustained release of encapsulated genetic material (e.g., pDNA, siRNA) over days to weeks. The degradation rate and release kinetics can be tuned by altering the lactide:glycolide ratio, molecular weight, and end-group chemistry.
PEG Shell Role: PEG is typically conjugated to the PLGA surface (PEGylation) or used as a block copolymer (PEG-PLGA). It forms a hydrophilic, steric barrier that reduces opsonization, minimizes non-specific protein adsorption, and increases systemic circulation time by evading the mononuclear phagocyte system (MPS). This "stealth" effect is critical for achieving targeted delivery and sustained gene release in vivo.
Key Data and Properties
Table 1: Tunable Parameters of PLGA and Their Impact on Nanoparticles
| Parameter | Typical Range | Impact on Nanoparticle Properties |
|---|---|---|
| Lactide:Glycolide (L:G) Ratio | 50:50, 65:35, 75:25, 85:15 | 50:50 degrades fastest (~1-2 months). Higher lactide content slows degradation and drug release. |
| Molecular Weight (kDa) | 10 - 100 kDa | Higher MW increases nanoparticle rigidity, slows polymer erosion, and can prolong release. |
| End Group | Ester (-COOH) or capped (e.g., -CH₃) | Acidic end groups increase hydrophilicity and degradation rate vs. capped termini. |
| PEG MW in PEG-PLGA (kDa) | 2 - 5 kDa (common for diblock) | Higher PEG MW enhances stealth but can reduce drug loading efficiency. Optimal ~5 kDa. |
| PEG Density (Surface Coverage) | Variable by synthesis method | Higher density improves circulation time but can hinder cellular uptake; requires optimization. |
Table 2: Quantitative Performance Metrics of PEG-PLGA vs. PLGA NPs
| Performance Metric | PLGA Nanoparticles (Non-PEGylated) | PEG-PLGA Nanoparticles (PEGylated) | Measurement Method |
|---|---|---|---|
| Zeta Potential (in water) | Typically -20 to -40 mV | Typically -5 to -15 mV (near-neutral) | Dynamic Light Scattering |
| Hydrodynamic Diameter | 150 - 250 nm | 150 - 300 nm (slightly larger shell) | Dynamic Light Scattering |
| In Vitro Release Half-life (Gene Payload) | ~3-7 days (burst release common) | ~7-21 days (more sustained, linear phase) | Dialysis in PBS at 37°C, quantification assay |
| In Vivo Circulation Half-life | Minutes to a few hours | 6 - 24 hours (species-dependent) | Blood sampling, fluorescence/PK analysis |
| Macrophage Uptake (In Vitro) | High (60-80% in 2h) | Low to Moderate (10-30% in 2h) | Flow cytometry of J774A.1/THP-1 cells |
Experimental Protocols
Protocol 1: Double Emulsion Solvent Evaporation for Gene-Loaded PEG-PLGA NP Synthesis
Objective: To encapsulate plasmid DNA (pDNA) or siRNA within PEG-PLGA nanoparticles.
Materials (Research Reagent Solutions Toolkit):
- PEG-PLGA Diblock Copolymer: (e.g., Resomer RGP d 50155, 50:50 PLGA-PEG 5kDa). Function: Forms core-shell nanoparticle matrix.
- Dichloromethane (DCM) or Ethyl Acetate: Function: Organic solvent for polymer dissolution.
- Polyvinyl Alcohol (PVA), 1-3% w/v: Function: Surfactant stabilizes the primary water-in-oil emulsion.
- Plasmid DNA or siRNA in Nuclease-Free TE Buffer: Function: Active genetic payload.
- Salmon Sperm DNA or Dextran (Molecular Biology Grade): Function: Carrier molecule to improve nucleic acid encapsulation efficiency.
- 2% w/v Sodium Cholate Solution: Function: Secondary surfactant for the double emulsion.
- Phosphate Buffered Saline (PBS), pH 7.4: Function: Washing and dispersion medium.
Procedure:
- Dissolve 100 mg PEG-PLGA in 2 mL DCM (Organic Phase, O).
- Prepare 200 µL of an aqueous solution containing 100 µg pDNA and 1 mg carrier DNA (Aqueous Phase 1, W1).
- Emulsify W1 in O using a probe sonicator (50 W, 30 s on ice) to form a primary W1/O emulsion.
- Immediately pour this primary emulsion into 4 mL of a cold 2% w/v sodium cholate solution (Aqueous Phase 2, W2). Homogenize at 13,000 rpm for 2 minutes to form a double (W1/O/W2) emulsion.
- Stir the double emulsion gently at room temperature for 3 hours to evaporate the organic solvent.
- Centrifuge the nanoparticles at 21,000 x g for 30 minutes at 4°C. Wash the pellet twice with nuclease-free water or PBS.
- Resuspend the final nanoparticle pellet in 1-2 mL of storage buffer (e.g., 5% w/v sucrose) and lyophilize for long-term storage.
Protocol 2:In VitroGene Release Kinetics Study
Objective: To quantify the sustained release profile of genetic material from nanoparticles.
Materials:
- Dialysis Cassettes (10 kDa MWCO) or Float-A-Lyzer G2 devices.
- Release Medium: PBS (pH 7.4) with 0.02% w/v sodium azide (preservative) and 0.1% w/v BSA (to simulate proteins).
- Quant-iT PicoGreen dsDNA Assay Kit: For quantifying released pDNA. For siRNA, use a fluorescence-based dye like RiboGreen.
Procedure:
- Accurately weigh 10 mg of gene-loaded, lyophilized nanoparticles.
- Resuspend in 1 mL of release medium and place inside the dialysis device.
- Immerse the device in 50 mL of pre-warmed (37°C) release medium under gentle stirring (50 rpm).
- At predetermined time points (1h, 4h, 8h, 1d, 2d, 4d, 7d, 14d, 21d), completely replace the external release medium with fresh, pre-warmed medium.
- Analyze the collected medium samples using the PicoGreen assay (following kit protocol) to determine the cumulative amount of released nucleic acid.
- Plot cumulative release (%) versus time to generate the release profile.
Diagrams
Diagram 1: PEG-PLGA NP Structure & Gene Release Mechanism
Title: Nanoparticle Structure and Gene Release Mechanism
Diagram 2: Workflow for NP Synthesis & Release Testing
Title: Experimental Workflow: Synthesis to Release Assay
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for PEG-PLGA Gene Delivery Research
| Item | Function & Rationale | Example/Catalog Consideration |
|---|---|---|
| PEG-PLGA Diblock Copolymer | The core material. Function: Self-assembles into core-shell NPs, providing both matrix (PLGA) and stealth (PEG) properties. | Merck (Resomer), PolySciTech, Akina (AP series). Select by L:G ratio, MW, PEG length. |
| Biodegradable Polymer Solvent | Function: Dissolves polymer for emulsion formation. Must be volatile for evaporation. | Dichloromethane (DCM) or Ethyl Acetate (less toxic alternative). HPLC grade. |
| Emulsion Stabilizers | Function: PVA stabilizes the W1/O interface; Sodium cholate stabilizes the final W1/O/W2 emulsion, controlling NP size. | Low MW PVA (87-89% hydrolyzed); High-purity bile salts. |
| Nucleic Acid Quantitation Kit | Function: Precisely measures low concentrations of released pDNA or siRNA in complex media. | Quant-iT PicoGreen (dsDNA) or RiboGreen (RNA). Highly sensitive and specific. |
| Dialysis Device (MWCO) | Function: Allows continuous sink conditions for release studies by containing NPs while permitting free nucleic acid diffusion. | Slide-A-Lyzer (10-20 kDa MWCO) or Float-A-Lyzer G2. |
| Lyoprotectant | Function: Prevents nanoparticle aggregation and protects payload during freeze-drying for storage. | Sucrose or Trehalose, molecular biology grade, 5% w/v. |
| Size/Zeta Potential Analyzer | Function: Critical QC instrument. Measures hydrodynamic diameter (DLS) and surface charge (zeta potential) of NPs in suspension. | Malvern Zetasizer Nano series. |
Mechanisms of Gene Encapsulation and Sustained Release Kinetics
Introduction Within the thesis "Engineering PEG-PLGA Nanoparticles for Tunable Sustained Release of Therapeutic Nucleic Acids," understanding the core mechanisms of gene encapsulation and the resultant release kinetics is paramount. This document provides detailed application notes and protocols central to this research, focusing on the double emulsion solvent evaporation technique for PEG-PLGA nanoparticle synthesis. The content is structured for replication and critical evaluation by researchers in drug development.
1. Mechanisms of Gene Encapsulation: The Double Emulsion (W/O/W) Process The encapsulation of hydrophilic nucleic acids (pDNA, siRNA) within a hydrophobic polyester matrix like PLGA is achieved via a water-in-oil-in-water (W/O/W) double emulsion. The process mechanism involves:
- Primary Emulsion (W₁/O): An aqueous solution of the gene therapeutic (W₁) is emulsified into a dichloromethane (DCM) solution containing dissolved PLGA-PEG copolymer (O) using a probe sonicator. This forms fine water droplets stabilized by the polymer at the interface.
- Secondary Emulsion (W₁/O/W₂): The primary emulsion is poured into a large volume of an external aqueous phase (W₂) containing a stabilizer (e.g., polyvinyl alcohol, PVA) and homogenized. This forms a double emulsion where the primary water droplets (containing the gene) are dispersed within oil droplets, themselves dispersed in the external water.
- Solvent Evaporation & Nanoparticle Hardening: The organic solvent (DCM) is gradually evaporated under reduced pressure or stirring, causing the polymer to precipitate and solidify, trapping the genetic payload within a polymeric matrix. The PEG chains migrate to the surface, creating a hydrophilic corona.
Diagram 1: Double Emulsion Encapsulation Workflow
2. Key Parameters Governing Encapsulation Efficiency (EE%) and Release Kinetics The following parameters critically influence payload encapsulation and the subsequent sustained release profile, a core focus of the thesis.
Table 1: Critical Formulation Parameters and Their Impact
| Parameter | Typical Range Tested | Impact on Encapsulation Efficiency (EE%) | Impact on Release Kinetics |
|---|---|---|---|
| PLGA L:G Ratio | 50:50, 65:35, 75:25, 85:15 | Higher lactide content (e.g., 75:25) increases hydrophobicity, potentially improving EE% for hydrophilic genes. | Higher glycolide content (e.g., 50:50) increases hydration/degradation rate, leading to faster release. |
| PLGA Molecular Weight (kDa) | 10-100 kDa | Higher Mw increases polymer viscosity, often improving EE% by reducing drug leakage. | Higher Mw polymers degrade slower, leading to more sustained release over weeks/months. |
| PEG Chain Length (% wt.) | 5-15% of polymer | >10% PEG can reduce EE% due to increased hydrophilicity and potential pore formation. | Higher PEG content increases initial burst release but can stabilize long-term release by hindering polymer erosion. |
| Gene-to-Polymer Ratio | 1:10 to 1:100 (w/w) | Lower ratios generally yield higher EE%. Excessive gene payload leads to saturation and low EE%. | Higher loading can increase the initial burst release fraction. |
| Primary Emulsion Sonication Energy | 50-150 J/mL | Optimal energy improves W₁ droplet dispersion, increasing EE%. Excessive energy can degrade nucleic acids. | Affects internal droplet size; finer dispersion can modulate release by altering diffusion paths. |
Protocol 1: Preparation of PEG-PLGA Nanoparticles for Gene Encapsulation Objective: Synthesize gene-loaded PEG-PLGA nanoparticles using the double emulsion solvent evaporation method. Materials: See "The Scientist's Toolkit" below. Procedure:
- Primary Emulsion: Dissolve 100 mg PLGA-PEG copolymer in 2 mL of dichloromethane (DCM, organic phase, O). In a separate vial, dissolve 1 mg of plasmid DNA (pDNA) or siRNA in 200 µL of nuclease-free water (internal aqueous phase, W₁). Add the W₁ phase to the O phase. Sonicate the mixture on ice using a probe sonicator at 40% amplitude for 30 seconds (pulse 1 sec on/1 sec off) to form a clear primary emulsion (W₁/O).
- Secondary Emulsion: Immediately pour the primary emulsion into 10 mL of 2% (w/v) polyvinyl alcohol (PVA) solution (external aqueous phase, W₂). Homogenize the mixture at 10,000 rpm for 2 minutes using a high-speed homogenizer to form a double emulsion (W₁/O/W₂).
- Solvent Evaporation: Transfer the double emulsion to a beaker containing 40 mL of 0.2% PVA solution under magnetic stirring. Stir gently for 4 hours at room temperature to allow complete evaporation of DCM and nanoparticle hardening.
- Nanoparticle Recovery: Centrifuge the nanoparticle suspension at 18,000 x g for 30 minutes at 4°C. Discard the supernatant and resuspend the pellet in nuclease-free water. Repeat the wash step twice to remove residual PVA.
- Lyophilization: Resuspend the final pellet in 2 mL of 5% (w/v) sucrose (cryoprotectant) solution. Freeze at -80°C and lyophilize for 48 hours. Store the powder at -20°C.
The Scientist's Toolkit: Essential Research Reagents
| Item | Function & Rationale |
|---|---|
| PLGA-PEG Diblock Copolymer | The core matrix material. PLGA provides biodegradability and controlled release; PEG confers steric stabilization ("stealth" properties) and modulates release kinetics. |
| Dichloromethane (DCM) | A volatile organic solvent that readily dissolves PLGA-PEG and is easily removed by evaporation. |
| Polyvinyl Alcohol (PVA) | A surfactant/stabilizer in the external phase (W₂) that prevents nanoparticle aggregation during formation and hardening. |
| Nuclease-Free Water | Essential for preparing gene solutions to prevent nucleic acid degradation. |
| Sucrose | A cryoprotectant used during lyophilization to maintain nanoparticle integrity and prevent aggregation upon reconstitution. |
| pDNA or siRNA | The model or therapeutic gene payload for encapsulation studies. |
3. Sustained Release Kinetics: Mechanisms and Analysis The release of genetic material from PEG-PLGA nanoparticles is a triphasic process governed by diffusion and erosion mechanisms.
Diagram 2: Triphasic Gene Release from PEG-PLGA NPs
Protocol 2: In Vitro Release Kinetics Study Objective: Quantify the sustained release profile of encapsulated genetic material from PEG-PLGA nanoparticles under physiological conditions. Materials: Gene-loaded nanoparticle powder (from Protocol 1), phosphate-buffered saline (PBS, pH 7.4) with 0.02% sodium azide, dialysis tubes (MWCO 100 kDa), microcentrifuge tubes, fluorescent nucleic acid stain (e.g., Quant-iT PicoGreen for dsDNA), microplate reader. Procedure:
- Sample Preparation: Reconstitute 10 mg of lyophilized nanoparticles in 1 mL of release medium (PBS, pH 7.4, 37°C). Transfer the suspension to a dialysis tube sealed at both ends.
- Release Setup: Immerse the dialysis tube in a 50 mL conical tube containing 30 mL of pre-warmed release medium. Incubate at 37°C under gentle agitation (100 rpm).
- Sampling: At predetermined time points (e.g., 1, 4, 8, 24, 48, 72 hours, then weekly for 4-8 weeks), collect 1 mL of the external release medium from the conical tube and replace it with 1 mL of fresh, pre-warmed medium.
- Quantification: Quantify the released gene in each sample. For pDNA: Mix 100 µL of sample with 100 µL of PicoGreen working solution in a black 96-well plate. Measure fluorescence (ex/em ~480/520 nm). Calculate cumulative release (%) against a standard curve and the total loaded amount (determined by dissolving nanoparticles in DMSO/NaOH).
- Data Modeling: Fit the cumulative release data to mathematical models (e.g., Higuchi, Korsmeyer-Peppas) to determine the dominant release mechanism.
Table 2: Mathematical Models for Release Kinetics Analysis
| Model | Equation | Application & Interpretation |
|---|---|---|
| Zero-Order | Q = k₀t | Describes constant release rate; ideal for sustained delivery. Rarely fits nanoparticle data perfectly. |
| Higuchi | Q = k_H √t | Describes release based on Fickian diffusion through a matrix. Often fits the initial 60% of release. |
| Korsmeyer-Peppas (Power Law) | Mt/M∞ = k tⁿ | Empirically determines release mechanism via n: n=0.45 (Fickian diffusion), 0.45 |
Application Notes: PEG-PLGA Nanoparticles in Sustained Gene Delivery
The pursuit of effective in vivo gene delivery has long been dominated by viral vectors. However, their clinical application is hampered by significant limitations: immunogenicity, insertional mutagenesis risk, limited cargo capacity, and complex, costly manufacturing. This document frames PEG-PLGA (polyethylene glycol-poly(lactic-co-glycolic acid)) nanoparticles (NPs) as a compelling non-viral alternative within sustained gene release research, detailing their advantages through specific application notes and validated protocols.
1. Safety Profile: Unlike viral vectors, PEG-PLGA NPs are synthetic, biodegradable, and non-replicative. They avoid pre-existing immunity, significantly reduce the risk of insertional oncogenesis, and exhibit favorable toxicity profiles. Their composition is hydrolytically degraded into metabolic monomers (lactic and glycolic acids).
2. Tunability: The physicochemical properties of PEG-PLGA NPs are highly engineerable. Key parameters can be precisely modulated to influence biodistribution, release kinetics, and cellular uptake.
Table 1: Tunable Parameters of PEG-PLGA Nanoparticles for Gene Delivery
| Parameter | Typical Range | Impact on Function | Method of Control |
|---|---|---|---|
| NP Size | 80-200 nm | Impacts circulation time, cellular uptake, and tissue penetration. | Homogenization speed/sonication energy, organic:aqueous phase ratio. |
| PEG MW & Density | PEG (2-5 kDa), 1-10% wt. | Reduces opsonization, prolongs circulation ("stealth" effect), modulates targeting. | Use of pre-synthesized PEG-PLGA diblock/triblock copolymers. |
| PLGA LA:GA Ratio | 50:50 to 85:15 | Controls degradation rate & release kinetics (50:50 degrades fastest). | Selection of commercially available PLGA. |
| Zeta Potential | Slightly negative to neutral (-10 to +5 mV) | Reduces non-specific clearance, improves colloidal stability. | Surface modification, choice of stabilizer (e.g., PVA). |
| Drug Loading | 1-5% (w/w) for pDNA | Impacts encapsulation efficiency and release profile. | Double emulsion (W/O/W) method optimization. |
3. Scalability: Formulation relies on established, scalable polymer chemistry and emulsion techniques compatible with Good Manufacturing Practice (GMP). This contrasts with the biological production of viral vectors, which faces challenges in yield, purity, and cost.
Table 2: Quantitative Comparison: Viral Vectors vs. PEG-PLGA Nanoparticles
| Feature | Viral Vectors (e.g., AAV, Lentivirus) | PEG-PLGA Nanoparticles |
|---|---|---|
| Immunogenicity | High (neutralizing antibodies, cellular immunity) | Low (PEGylated, synthetic) |
| Insertional Mutagenesis Risk | Low (AAV) to High (Retrovirus) | None |
| Cargo Capacity | Limited (<5 kb for AAV) | High (>10 kb possible) |
| Manufacturing Scalability | Complex, low-yield, high-cost | Straightforward, high-yield, lower cost |
| Production Time | Weeks to months | Days |
| Storage Stability | Requires -80°C, sensitive | Generally stable at 2-8°C or -20°C |
| Release Kinetics | Typically rapid, burst expression | Sustained release (days to weeks) tunable |
| Targeting Flexibility | Requires re-engineering of capsid | Easy surface functionalization with ligands |
Experimental Protocols
Protocol 1: Formulation of pDNA-Loaded PEG-PLGA Nanoparticles via Double Emulsion (W/O/W) Objective: To encapsulate plasmid DNA (pDNA) in PEG-PLGA NPs for sustained release. Materials: See "The Scientist's Toolkit" below. Procedure:
- Primary Emulsion (W1/O): Dissolve 50 mg PEG-PLGA (e.g., 5% PEG, 75:25 LA:GA) in 2 mL dichloromethane (DCM). In a separate tube, dilute 100 µg of pDNA (in TE buffer) in 100 µL of nuclease-free water. Combine the aqueous pDNA solution with the polymer/DCM solution. Sonicate the mixture on ice using a microtip probe sonicator at 40-50 W for 30-60 seconds to form a water-in-oil (W1/O) emulsion.
- Secondary Emulsion (W1/O/W2): Quickly pour the primary emulsion into 4 mL of a 2% (w/v) polyvinyl alcohol (PVA) solution. Homogenize at 10,000 rpm for 2 minutes to form the double emulsion (W1/O/W2).
- Solvent Evaporation: Stir the double emulsion magnetically at room temperature for 4-6 hours to evaporate the organic solvent.
- Nanoparticle Harvesting: Centrifuge the suspension at 18,000 x g for 30 minutes at 4°C. Wash the pellet twice with nuclease-free water to remove excess PVA.
- Lyophilization: Resuspend the final nanoparticle pellet in 1 mL of 5% (w/v) trehalose solution (cryoprotectant). Freeze at -80°C and lyophilize for 48 hours. Store dried NPs at -20°C.
Protocol 2: In Vitro Sustained Release Kinetics of pDNA Objective: To quantify the release of pDNA from NPs over time. Procedure:
- Weigh 10 mg of lyophilized pDNA-NPs and suspend in 1 mL of PBS (pH 7.4) containing 0.02% sodium azide in a microcentrifuge tube. Incubate at 37°C under gentle agitation.
- At predetermined time points (1, 6, 24 hours, then days 3, 5, 7, 10, 14), centrifuge the sample at 18,000 x g for 30 minutes.
- Carefully collect 800 µL of the supernatant for analysis and replace with 800 µL of fresh release medium.
- Quantify the released pDNA in the supernatant using a Picogreen dsDNA assay against a standard curve. Calculate cumulative release percentage.
- Data Modeling: Fit the release data to models (e.g., Higuchi, Korsmeyer-Peppas) to determine release mechanisms.
Visualizations
Title: Core Advantages of PEG-PLGA NPs vs. Viral Vectors
Title: Workflow for pDNA NP Formulation
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| PEG-PLGA Copolymer (e.g., 5% PEG, 75:25 LA:GA) | Amphiphilic polymer forming the NP matrix. PEG confers stealth; PLGA ratio controls degradation. |
| Endotoxin-Free Plasmid DNA | Therapeutic gene cargo. Must be high purity and concentration (>1 mg/mL). |
| Polyvinyl Alcohol (PVA) | Emulsifier and stabilizer. Critical for forming stable double emulsions and controlling NP size. |
| Dichloromethane (DCM) | Organic solvent for dissolving PLGA polymer. Volatile for easy evaporation. |
| Trehalose Dihydrate | Lyoprotectant. Prevents NP aggregation during freeze-drying, ensuring long-term stability. |
| Picogreen Assay Kit | Ultrasensitive fluorescent nucleic acid stain. Quantifies trace amounts of released pDNA. |
| Nuclease-Free Water & Buffers | Prevents degradation of the pDNA cargo during formulation and analysis. |
| Dynamic Light Scattering (DLS) Instrument | For characterizing NP hydrodynamic diameter, polydispersity index (PDI), and zeta potential. |
Within the broader thesis on developing PEG-PLGA nanoparticles for sustained gene release, addressing the tripartite barrier of cellular uptake, endosomal escape, and nuclear entry is paramount for achieving therapeutic efficacy. These application notes consolidate current strategies and provide actionable protocols.
Table 1: Common Ligands for Receptor-Mediated Uptake of PEG-PLGA Nanoparticles
| Ligand | Target Receptor | Typical Conjugation Method | Reported Uptake Increase* | Key Considerations |
|---|---|---|---|---|
| Transferrin | Transferrin Receptor (TfR) | NHS-PEG-Maleimide | 3-5 fold | Ubiquitous but overexpressed in many cancers. |
| Folic Acid | Folate Receptor (FR) | PEG-Folate conjugates | 4-8 fold | Highly specific; FRα overexpressed in ovarian, lung, breast cancers. |
| RGD Peptide | αvβ3 Integrin | Carbodiimide (EDC/NHS) | 2-4 fold | Targets tumor angiogenesis; promotes cell adhesion. |
| Anti-EGFR Antibody | EGFR | Thiol-maleimide or streptavidin-biotin | 5-10 fold | High specificity; larger size may affect nanoparticle properties. |
*Baseline: Non-targeted PEG-PLGA nanoparticles.
Table 2: Endosomal Escape Agents and Their Mechanisms
| Agent | Mechanism | Typical Incorporation Method | Reported Escape Efficiency* | Potential Cytotoxicity |
|---|---|---|---|---|
| Chloroquine | Endosomotropic (buffering) | Co-incubation with NPs | 20-35% | High at effective doses. |
| HA2 Peptide | pH-dependent membrane fusion | Covalent attachment to PLGA or PEG | 40-60% | Low. |
| Cell-Penetrating Peptides (e.g., TAT) | Multiple (pore formation, fusion) | Covalent attachment or physical mix | 30-50% | Variable; depends on sequence and density. |
| Protonable Polymers (e.g., PEI) | Proton Sponge Effect | Co-encapsulation or surface coating | 50-70% | High for high Mw PEI; lower for short chains. |
*Quantified as % of internalized NPs/genes reaching the cytosol.
Detailed Experimental Protocols
Protocol 1: Conjugation of Targeting Ligands (Folate) to PEG-PLGA Nanoparticles
Objective: Synthesize folate-PEG-PLGA for active targeting. Materials: PLGA-COOH, NH2-PEG-COOH, Folic Acid, EDC, NHS, DCC, DMSO, Dialysis tubing. Procedure:
- Activate Folate: Dissolve folic acid (10 mg) and DCC (15 mg) in anhydrous DMSO (2 mL). Stir under N2 for 6h. Filter to remove dicyclohexylurea.
- Conjugate to PEG: Add NH2-PEG-COOH (100 mg) to the activated folate solution. React for 24h at RT, protected from light. Dialyze (1 kDa MWCO) against water to obtain Folate-PEG-COOH.
- Form Nanoparticles: Mix Folate-PEG-COOH with PLGA-COOH (e.g., 1:10 w/w) in acetone. Use nanoprecipitation or emulsion-solvent evaporation to form targeted NPs.
- Validate: Confirm conjugation via 1H-NMR; quantify surface folate by UV-Vis.
Protocol 2: Quantifying Endosomal Escape Efficiency via Gal8-mCherry Assay
Objective: Visualize and quantify endosomal membrane rupture. Materials: HeLa cells (Gal8-mCherry reporter), Nanoparticles, Hoechst 33342, Confocal microscope, ImageJ. Procedure:
- Seed Cells: Plate Gal8-mCherry HeLa cells in 8-well chamber slides at 60% confluency.
- Treat: Incubate with fluorescently-labeled NPs (e.g., Cy5-DNA loaded) for 4h.
- Stain & Image: Stain nuclei with Hoechst. Acquire Z-stack images at 60x.
- Analyze: Count cytosolic Cy5 puncta not co-localized with Gal8-mCherry foci. Calculate escape ratio: (Free Cy5 puncta / Total intracellular Cy5 puncta) x 100%.
Protocol 3: Assessing Nuclear Entry via Fractionation & qPCR
Objective: Quantify plasmid DNA delivery to the nucleus. Materials: Cells, NPs, Nuclear/Cytoplasmic Fractionation Kit, qPCR system, primers for plasmid gene (e.g., GFP). Procedure:
- Treat & Harvest: Treat cells with plasmid-loaded NPs. Harvest at 24h post-transfection.
- Fractionate: Use the kit to separate cytoplasmic and nuclear fractions. Validate purity via Lamin B1 (nuclear) and GAPDH (cytoplasmic) Western blot.
- Extract DNA: Isolate total DNA from both fractions.
- qPCR: Run qPCR with plasmid-specific primers. Use a standard curve of pure plasmid to calculate copy numbers. Express as % of total intracellular plasmid copies in the nuclear fraction.
Visualization Diagrams
Diagram 1: Gene Delivery Pathway from NP to Nucleus (76 chars)
Diagram 2: Three Primary Endosomal Escape Mechanisms (79 chars)
The Scientist's Toolkit
Table 3: Essential Research Reagents for Investigating Delivery Barriers
| Reagent / Material | Function / Application | Key Consideration |
|---|---|---|
| NH2-PEG-COOH / COOH-PEG-NHS | Linker for covalent ligand conjugation to NP surface. | PEG length (e.g., 2k vs 5k Da) affects ligand presentation & stealth. |
| Fluorescent Dyes (Cy5, FITC, DIR) | Label polymers or nucleic acids to track cellular uptake & localization via flow cytometry/imaging. | Choose dye with minimal pH sensitivity for endosomal tracking. |
| LysoTracker & Early Endosome Dyes | Fluorescent markers for specific organelles to assess co-localization. | Use with fixed or live cells depending on protocol. |
| Chloroquine Diposphate | Positive control for endosomotropic activity. | High concentrations are cytotoxic; titrate carefully. |
| Bafilomycin A1 | V-ATPase inhibitor; blocks endosomal acidification. Negative control for pH-dependent escape. | Validates mechanism but completely inhibits escape for some agents. |
| WGA (Wheat Germ Agglutinin) | Inhibitor of nuclear import; negative control for nuclear entry assays. | Reversible; use at non-cytotoxic doses. |
| Nuclear/Cytoplasmic Fractionation Kit | Isolates subcellular compartments to quantify nuclear delivery (e.g., via qPCR). | Check for cross-contamination with organelle-specific markers. |
| Gal8-mCherry Reporter Cell Line | Visualizes endosomal rupture via Galectin-8 recruitment. | Gold-standard direct escape assay; requires confocal imaging. |
Fabrication and Functionalization: Protocols for PEG-PLGA Gene Nanoparticle Synthesis
Within the context of developing PEG-PLGA nanoparticles for sustained gene release, the selection of a synthesis method is critical. It dictates key nanoparticle characteristics such as size, polydispersity, encapsulation efficiency, and release kinetics. This document provides detailed application notes and protocols for three primary synthesis methods, optimized for encapsulating genetic material (e.g., pDNA, siRNA) within PEG-PLGA nanoparticles to achieve controlled, sustained release profiles essential for in vivo applications.
Application Notes & Comparative Analysis
Double Emulsion (W/O/W)
Best for: High encapsulation efficiency of hydrophilic macromolecules like DNA/RNA. Principle: A primary water-in-oil (W/O) emulsion, containing the aqueous genetic material in an organic PLGA solution, is emulsified into a secondary aqueous phase containing a stabilizer (e.g., PVA). Key Considerations: The two emulsification steps introduce shear stress, which can fragment nucleic acids. Sustained release is achieved by the slow, hydrolytic degradation of the PLGA matrix, with PEG chains providing steric stabilization and reduced opsonization in vivo.
Nanoprecipitation (Solvent Displacement)
Best for: Small, monodisperse nanoparticles with moderate encapsulation efficiency for hydrophobic or some hydrophilic agents. Principle: A water-miscible organic solvent containing the polymer and drug is added to an aqueous phase under moderate stirring. The rapid diffusion of the solvent into the water phase causes instantaneous polymer precipitation into nanoparticles. Key Considerations: Gentle on nucleic acid structure but initial encapsulation efficiency for hydrophilic genes can be low without modification. Often used with cationic lipids or polymers co-encapsulated to complex genetic material. Release is governed by diffusion and polymer erosion.
Microfluidics
Best for: Ultra-uniform nanoparticles with precise control over size and composition; excellent reproducibility. Principle: Laminar flow streams of the organic (polymer) and aqueous phases meet in a microfluidic chip, enabling controlled mixing via diffusion at the interface, leading to highly reproducible nanoprecipitation. Key Considerations: Ideal for screening formulation parameters. Enables precise tuning of nanoparticle characteristics critical for predictable gene release kinetics. Scalability can require parallelized chip setups.
Table 1: Comparative Analysis of PEG-PLGA Nanoparticle Synthesis Methods for Gene Delivery
| Parameter | Double Emulsion (W/O/W) | Nanoprecipitation | Microfluidics |
|---|---|---|---|
| Typical Size (nm) | 150 - 300 | 80 - 200 | 50 - 200 (tight PDI) |
| Polydispersity (PDI) | 0.15 - 0.25 | 0.10 - 0.20 | 0.05 - 0.10 |
| Encapsulation Efficiency (EE%) for DNA/RNA | High (60-90%) | Moderate to Low (20-50%)* | Tunable (30-80%) |
| Process Scalability | High (batch volume) | High (batch volume) | Moderate (requires scaling-out) |
| Key Advantage | High EE for hydrophilic genes | Simplicity, small size | Precision & Reproducibility |
| Key Limitation | Shear stress on nucleic acids | Lower EE for hydrophilic genes | Initial setup complexity |
| Primary Release Mechanism | Polymer degradation-led | Diffusion & erosion | Precisely engineered diffusion/degradation |
*Can be improved using modified techniques (e.g., ionic complexation prior to nanoprecipitation).
Detailed Experimental Protocols
Protocol: Double Emulsion (W/O/W) for pDNA-Loaded PEG-PLGA NPs
Objective: Synthesize PEG-PLGA nanoparticles encapsulating plasmid DNA for sustained release.
Materials:
- PEG-PLGA copolymer (e.g., PLGA-PEG-COOH, 50:50, MW 15k-5k).
- Plasmid DNA (pDNA) in nuclease-free TE buffer or water.
- Dichloromethane (DCM), HPLC grade.
- Polyvinyl alcohol (PVA, 2% w/v in water).
- Diethyl ether.
- Ultra-pure water.
- Probe sonicator.
- Magnetic stirrer.
Procedure:
- Primary Emulsion (W1/O): Dissolve 50 mg PEG-PLGA in 2 mL DCM. Add 100 µL of aqueous pDNA solution (1 mg/mL) to the organic phase. Sonicate this mixture on ice using a probe sonicator at 40 W for 60 seconds to form a stable water-in-oil (W1/O) emulsion.
- Secondary Emulsion (W1/O/W2): Quickly pour the primary emulsion into 10 mL of a continuously stirring (600 rpm) 2% PVA solution. Sonicate this mixture on ice at 60 W for 120 seconds to form the double emulsion (W1/O/W2).
- Solvent Evaporation: Stir the double emulsion at room temperature, uncovered, for 4 hours to allow complete evaporation of the organic solvent.
- Nanoparticle Recovery: Centrifuge the suspension at 20,000 x g for 30 minutes at 4°C. Wash the pellet with water to remove excess PVA. Resuspend the final nanoparticle pellet in 5 mL of nuclease-free water or buffer for characterization.
- Characterization: Measure size and PDI by DLS. Determine pDNA encapsulation efficiency using a Quant-iT PicoGreen assay after nanoparticle dissolution in 0.1N NaOH/1% SDS.
Protocol: Modified Nanoprecipitation for siRNA-PEG-PLGA NPs
Objective: Form small, stable nanoparticles for siRNA delivery.
Materials:
- PEG-PLGA (e.g., PLGA-PEG-NH2 for surface functionalization).
- siRNA (target sequence).
- Cationic lipid (e.g., DOTAP) or polymer (e.g., chitosan).
- Acetone, HPLC grade.
- Tween 80.
- Ultra-pure water.
- Syringe pump or pipette.
- Magnetic stirrer.
Procedure:
- Organic Phase: Dissolve 25 mg PEG-PLGA and 2 mg cationic lipid (DOTAP) in 5 mL of acetone.
- Aqueous Phase: Prepare 10 mL of a 0.3% (v/v) Tween 80 solution in nuclease-free water. Optional: Pre-complex siRNA with a portion of cationic lipid in a separate tube for 15 minutes.
- Nanoprecipitation: Using a syringe pump, add the organic phase to the magnetically stirred aqueous phase (600 rpm) at a constant rate of 1 mL/min.
- Solvent Removal: Stir the resulting suspension for 2 hours to allow complete evaporation of acetone.
- Concentration & Purification: Concentrate the nanoparticle suspension using centrifugal filter units (MWCO 100kDa). Wash twice with water.
- Characterization: Measure size/PDI by DLS. Determine siRNA EE using RiboGreen assay.
Protocol: Microfluidic Synthesis using a Hydrodynamic Flow-Focusing Chip
Objective: Produce monodisperse PEG-PLGA nanoparticles with precise control over size.
Materials:
- PEG-PLGA copolymer.
- Acetonitrile (ACN) or ethanol.
- Sterile PBS or water.
- Microfluidic chip (e.g., Dolomite Part # 3000427).
- Precision syringe pumps (2-3).
- Gas-tight glass syringes.
- Tubing and connectors.
Procedure:
- Phase Preparation: Organic Phase: Dissolve PEG-PLGA in ACN at 10 mg/mL. Aqueous Phase: Use sterile PBS.
- Chip Setup: Load the organic phase into a syringe connected to the center inlet. Load the aqueous phase into syringes connected to the two side inlets. Connect an output tube from the outlet to a collection vial.
- Flow Rate Optimization: Set the aqueous phase flow rate (Qaq) to 1.0 mL/min *per side channel*. Set the organic phase flow rate (Qorg) to 0.2 mL/min. The Total Flow Ratio (TFR = Qaq total / Qorg) is 10:1, typically yielding smaller nanoparticles.
- Run Synthesis: Start the pumps simultaneously. Collect the effluent in a vial placed on a stir plate. The rapid mixing within the chip precipitates nanoparticles instantly.
- Solvent Dialysis: Immediately transfer the collected suspension to a dialysis membrane (MWCO 3.5 kDa) against water for 2 hours to remove ACN.
- Characterization: Analyze size, PDI, and concentration. The PDI should be consistently below 0.1.
Diagrams
Title: Double Emulsion Workflow for pDNA NPs
Title: Nanoprecipitation Mechanism
Title: Gene Release Pathway from PEG-PLGA NPs
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for PEG-PLGA Nanoparticle Synthesis in Gene Delivery
| Item/Category | Example & Function |
|---|---|
| PEG-PLGA Copolymers | Lactel Absorbable Polymers (Birmingham Polymers): Provides core matrix (PLGA) and stealth/functionalization (PEG). MW and LA:GA ratio control degradation rate. |
| Genetic Material | Sigma-Aldrich (pDNA) or Dharmacon (siRNA): The therapeutic payload. Requires high purity and integrity. |
| Biocompatible Solvents | Dichloromethane (DCM), Acetonitrile (ACN), Ethyl Acetate: Organic solvents for dissolving polymer. Must be easily removable. |
| Surfactants/Stabilizers | Polyvinyl Alcohol (PVA), Tween 80, Pluronic F-68: Stabilize the emulsion during formation and prevent aggregation. |
| Cationic Complexing Agents | DOTAP Lipids (Avanti Polar Lipids), Polyethylenimine (PEI): Improve encapsulation and loading of negatively charged nucleic acids. |
| Purification Tools | Amicon Ultra Centrifugal Filters (MilliporeSigma): For concentrating and washing nanoparticles via size exclusion. |
| Characterization Kits | Quant-iT PicoGreen/RiboGreen Assay (Thermo Fisher): Specifically quantifies encapsulated or free DNA/RNA. |
| Microfluidic Hardware | Dolomite Microfluidic Chips & Syringe Pumps: For precise, reproducible nanoparticle synthesis via microfluidics. |
Critical Process Parameters Affecting Particle Size, PDI, and Encapsulation Efficiency
Within a broader thesis on developing PEG-PLGA nanoparticles for sustained gene release, controlling Critical Process Parameters (CPPs) is essential for producing nanocarriers with optimal physicochemical properties. Particle size, polydispersity index (PDI), and encapsulation efficiency (EE%) are critical quality attributes that directly influence in vivo biodistribution, cellular uptake, and release kinetics. This Application Note details the key CPPs, their mechanistic effects, and standardized protocols for systematic optimization in gene delivery formulations.
Key Critical Process Parameters & Quantitative Effects
The following table summarizes the primary CPPs and their quantified impact on nanoparticle characteristics, based on current literature and experimental data.
Table 1: Critical Process Parameters and Their Impact on Nanoparticle Attributes
| Critical Process Parameter | Typical Range Studied | Effect on Particle Size (nm) | Effect on PDI | Effect on Encapsulation Efficiency (%) | Primary Mechanistic Reason |
|---|---|---|---|---|---|
| Organic Phase : Aqueous Phase Volume Ratio | 1:1 to 1:10 | 120 → 85 (as ratio increases) | 0.25 → 0.15 (as ratio increases) | 75 → 92 (as ratio increases) | Enhanced diffusion rate and solvent displacement with larger aqueous volume. |
| PEG-PLGA Polymer Concentration | 0.5% to 5% (w/v) | 80 → 220 (as concentration increases) | 0.1 → 0.3 (as concentration increases) | 50 → 95 (as concentration increases) | Increased viscosity and polymer mass available for encapsulation. |
| Sonication Energy / Time (Emulsification) | 30-120 sec at 40-80W | 200 → 100 (as energy increases) | 0.4 → 0.12 (as energy increases) | 65 → 80 (as energy increases) | Increased shear forces disrupt larger droplets, creating finer emulsion. |
| Polymer to Gene (pDNA/siRNA) Mass Ratio | 10:1 to 100:1 | 110 → 150 (as ratio increases) | 0.15 → 0.22 (as ratio increases) | 40 → 95 (as ratio increases) | Greater polymer mass encapsulates nucleic acid more effectively. |
| Surfactant Concentration (e.g., PVA) | 0.1% to 5% (w/v) | 250 → 110 (as [PVA] increases to opt.) | 0.3 → 0.1 (as [PVA] increases to opt.) | 70 → 90 (as [PVA] increases to opt.) | Stabilizes emulsion interface, preventing coalescence. Excess can cause secondary nucleation. |
Experimental Protocols
Protocol 3.1: Standardized Double Emulsion (W/O/W) Method for pDNA-Loaded PEG-PLGA NPs
Objective: To reproducibly prepare nanoparticles with controlled size, low PDI, and high encapsulation efficiency for plasmid DNA (pDNA).
Materials (See Toolkit Section 5)
- Reagents: PEG-PLGA (e.g., PLGA-PEG-COOH), Dichloromethane (DCM), Polyvinyl Alcohol (PVA, Mw 30-70 kDa), pDNA in TE buffer, Ethyl Acetate, Ultra-pure Water.
- Equipment: Probe Sonicator, Magnetic Stirrer, Centrifuge, Zetasizer, HPLC system.
Procedure:
- Primary W/O Emulsion: Dissolve 50 mg PEG-PLGA in 2 mL DCM (organic phase). Add 200 µL of aqueous pDNA solution (1 mg/mL) to the polymer solution. Sonicate this mixture on ice using a probe sonicator at 40W for 60 seconds (pulse: 5 sec on, 2 sec off) to form the primary water-in-oil (W/O) emulsion.
- Secondary W/O/W Emulsion: Inject the primary emulsion into 10 mL of a 2% (w/v) PVA solution under vigorous magnetic stirring (800 rpm). Sonicate this secondary mixture on ice at 35W for 90 seconds.
- Solvent Evaporation & Hardening: Transfer the double emulsion to 40 mL of a 0.1% PVA solution. Stir for 4 hours at room temperature to allow complete solvent evaporation and nanoparticle hardening.
- Nanoparticle Recovery: Centrifuge the suspension at 18,000 x g for 30 minutes at 4°C. Wash the pellet twice with ultrapure water to remove excess PVA and unencapsulated pDNA.
- Resuspension: Resuspend the final nanoparticle pellet in 5 mL of sterile phosphate-buffered saline (PBS, pH 7.4) or lyophilization buffer for storage.
Protocol 3.2: Determination of Encapsulation Efficiency
Objective: To quantify the amount of gene therapeutic successfully encapsulated within the nanoparticles.
Indirect Method (Measuring Unencapsulated Gene):
- After the final wash step (Protocol 3.1, Step 4), carefully collect all supernatant and wash fractions.
- Quantify the amount of free, unencapsulated pDNA/siRNA in the pooled supernatants using a fluorescent nucleic acid binding assay (e.g., Quant-iT PicoGreen for dsDNA).
- Calculate EE% using the formula: EE% = [(Total amount of gene added – Amount of free gene in supernatant) / Total amount of gene added] x 100.
Visualized Pathways and Workflows
Title: Relationship Between CPPs, Nanoparticle Properties, and Performance
Title: Double Emulsion Workflow for PEG-PLGA Gene Nanoparticles
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for PEG-PLGA Gene Nanoparticle Research
| Item | Function in Research | Example/Specification |
|---|---|---|
| PEG-PLGA Copolymer | Forms the nanoparticle matrix; PEG confers stealth properties, PLGA controls biodegradability and release. | Lactel Polymers: PLGA-PEG-COOH, 50:50 PLGA, 5k-10k PEG block. Resomer series. |
| Nucleic Acid (Gene Payload) | The active therapeutic agent to be encapsulated and delivered. | pDNA (e.g., gWIZ GFP), siRNA (e.g., anti-GAPDH), or mRNA. |
| Volatile Organic Solvent | Dissolves the polymer to form the organic phase. | Dichloromethane (DCM), Ethyl Acetate (less toxic alternative). |
| Aqueous Phase Surfactant | Stabilizes the emulsion droplets during formation, prevents aggregation. | Polyvinyl Alcohol (PVA, Mw 30-70 kDa, 87-89% hydrolyzed). |
| Sonication Probe | Applies high shear energy to create fine, homogeneous emulsions. | Tip diameter: 3-6 mm, with pulse capability to prevent overheating. |
| Ultracentrifuge | Pelletizes nanoparticles for washing and concentration. | Capable of ≥ 18,000 x g, with temperature control. |
| Dynamic Light Scattering (DLS) Instrument | Measures hydrodynamic particle size, size distribution (PDI), and zeta potential. | Malvern Zetasizer Nano ZS. |
| Fluorescent Nucleic Acid Stain | Quantifies free or encapsulated nucleic acid for encapsulation efficiency. | Quant-iT PicoGreen (dsDNA) or RiboGreen (RNA). |
| Lyophilizer | Preserves nanoparticle integrity for long-term storage by removing water. | Bench-top freeze dryer with condenser capacity for aqueous samples. |
Surface Functionalization Strategies for Targeted Delivery (e.g., Peptides, Antibodies)
Application Notes
This section details the application of surface functionalization strategies to PEG-PLGA nanoparticles (NPs) within a thesis focused on sustained gene release. Functionalization is critical for achieving targeted delivery to specific cell types, enhancing cellular uptake, and improving the therapeutic index of encapsulated nucleic acids (e.g., pDNA, siRNA).
Key Rationale: The "stealth" property conferred by PEG in PEG-PLGA NPs reduces non-specific uptake and extends circulation time. However, this can also passively hinder interaction with target cells. Active targeting ligands grafted onto the NP surface overcome this by binding to receptors overexpressed on target cells, promoting receptor-mediated endocytosis.
Primary Ligand Classes:
- Peptides: Short sequences (e.g., RGD for αvβ3 integrin) offer small size, ease of synthesis, and modular design.
- Antibodies/Fragments: Provide high specificity and affinity (e.g., anti-HER2, anti-EGFR). Smaller fragments (scFv, Fab) are often preferred to minimize steric hindrance.
- Other Ligands: Aptamers, small molecules (folic acid), and proteins (transferrin) are also commonly used.
Critical Consideration: The conjugation chemistry must preserve ligand activity and NP stability. The "density" of ligands on the NP surface is a crucial parameter that directly influences binding avidity and cellular outcomes—too low may be ineffective, while too high may cause aggregation or non-specific binding.
Table 1: Comparison of Targeting Ligands on PEG-PLGA Nanoparticles for Gene Delivery In Vitro
| Ligand Type | Specific Target | Conjugation Method | Ligand Density (approx. molecules/NP) | Reported Gene Transfection Efficiency Increase (vs. non-targeted NP) | Key Cell Line / Model |
|---|---|---|---|---|---|
| cRGDfK Peptide | αvβ3/αvβ5 Integrin | Maleimide-thiol (PEG terminus) | 50-100 | 3.5 to 5-fold | Human Umbilical Vein Endothelial Cells (HUVECs) |
| Anti-EGFR scFv | Epidermal Growth Factor Receptor | NHS Ester-Amine | 20-40 | 4 to 6-fold | EGFR+ A431 Epidermal Carcinoma |
| TAT Peptide | Cell Membrane (Nonspecific) | Carbodiimide (EDC/NHS) | >200 | 2 to 3-fold (uptake) | HeLa (general cell penetration) |
| Transferrin | Transferrin Receptor (TfR) | EDC/NHS Chemistry | 30-60 | 4-fold | Brain Capillary Endothelial Cells (BCECs) |
Table 2: Impact of Ligand Density on Nanoparticle Pharmacokinetics (Representative In Vivo Data)
| Nanoparticle Formulation | Ligand Density (Peptides/NP) | Circulation Half-life (t1/2, h) | Tumor Accumulation (%ID/g) | Tumor-to-Liver Ratio |
|---|---|---|---|---|
| Non-targeted PEG-PLGA | 0 | ~12 | 1.2 | 0.3 |
| cRGDfK-PEG-PLGA (Low) | ~30 | ~10 | 2.8 | 0.8 |
| cRGDfK-PEG-PLGA (Medium) | ~70 | ~8.5 | 4.5 | 1.5 |
| cRGDfK-PEG-PLGA (High) | ~150 | ~6 | 3.9 | 1.1 |
Detailed Protocols
Protocol 1: Synthesis of Maleimide-Terminated PEG-PLGA Copolymer for Thiol-Based Conjugation
Objective: To synthesize a functionalized copolymer where the PEG terminus is available for covalent conjugation to thiol-containing ligands (e.g., cysteine-terminated peptides, reduced antibody fragments).
Materials:
- PLGA-COOH (e.g., Resomer RG 503H, 50:50, 24-38 kDa)
- HO-PEG-NH₂ (MW 3400 Da)
- N,N'-Dicyclohexylcarbodiimide (DCC)
- N-Hydroxysuccinimide (NHS)
- N-(2-Aminoethyl)maleimide, trifluoroacetic acid salt
- Anhydrous Dichloromethane (DCM) and Dimethylformamide (DMF)
- Diethyl Ether (cold)
- Dialysis tubing (MWCO 3.5 kDa)
- Freeze dryer
Procedure:
- Activate PLGA-COOH: Dissolve PLGA-COOH (1 mmol) and NHS (1.2 mmol) in 20 mL anhydrous DCM. Stir under argon. Add DCC (1.2 mmol) in DCM dropwise at 0°C. React for 6 h at room temperature (RT). Filter to remove dicyclohexylurea precipitate.
- Conjugate PEG: Add HO-PEG-NH₂ (1.1 mmol) and a catalytic amount of triethylamine to the NHS-activated PLGA solution. Stir for 24 h at RT under argon.
- Recover PLGA-PEG-NH₂: Precipitate the polymer in cold diethyl ether, centrifuge, and wash 3x. Dry under vacuum.
- Maleimide Functionalization: Dissolve PLGA-PEG-NH₂ (0.5 mmol) and N-(2-aminoethyl)maleimide (1 mmol) in 10 mL anhydrous DMF. Add EDC·HCl (1.2 mmol) and NHS (1.2 mmol). React for 24 h at RT, protected from light.
- Purification: Dialyze the reaction mixture against DI water (pH 6.5, 4°C, 48 h, with frequent water changes) to remove unreacted reagents. Lyophilize to obtain the final PLGA-PEG-MAL copolymer as a white solid. Confirm via ¹H NMR (CDCl₃): δ 6.7 ppm (s, 2H, maleimide).
Protocol 2: Conjugation of cRGDfK Peptide to PLGA-PEG-MAL Nanoparticles & siRNA Encapsulation
Objective: To prepare targeted siRNA-loaded nanoparticles using a pre-functionalized copolymer.
Materials:
- PLGA-PEG-MAL copolymer (from Protocol 1)
- Plain PLGA-COOH
- cRGDfK peptide (Cyclo(Arg-Gly-Asp-D-Phe-Lys), thiolated or with terminal Cys)
- siRNA (e.g., anti-GFP siRNA)
- Polyvinyl alcohol (PVA, MW 30-70 kDa)
- Double-emulsion solvent evaporation equipment (sonicator, homogenizer)
- Nitrogen gas stream
- Tris(2-carboxyethyl)phosphine (TCEP)
- Zeta potential & DLS analyzer
Procedure: A. Ligand Conjugation & Nanoparticle Preparation (w/o/w double emulsion):
- Peptide Reduction (if needed): Dissolve cRGDfK-SH in degassed PBS (pH 7.0) with 5x molar excess of TCEP. Incubate 1 h at RT to reduce disulfide bonds. Purify via desalting column.
- Prepare Organic Phase: Dissolve a polymer blend (e.g., 85:15 plain PLGA:PLGA-PEG-MAL, 50 mg total) in 2 mL DCM.
- Prepare First Aqueous Phase (W1): Dissolve siRNA (100 µg) in 100 µL nuclease-free water.
- Form Primary Emulsion (W1/O): Add the siRNA solution to the organic phase. Sonicate on ice (50% amplitude, 30 s) using a probe sonicator under a nitrogen atmosphere to form a water-in-oil emulsion.
- Form Secondary Emulsion (W1/O/W2): Add the primary emulsion to 4 mL of 2% (w/v) PVA solution. Homogenize (10,000 rpm, 1 min) or sonicate again (30 s) to form a double emulsion.
- Conjugate Ligand: Immediately add the reduced cRGDfK peptide (10x molar excess to maleimide groups) to the W2 phase. Stir gently for 6 h at 4°C, protected from light, allowing conjugation to occur at the nanoparticle interface.
- Solvent Evaporation & Harvest: Stir the emulsion overnight at RT to evaporate DCM. Collect nanoparticles by ultracentrifugation (21,000 rpm, 30 min, 4°C). Wash 3x with water to remove PVA and unreacted peptide. Resuspend in buffer and lyophilize with a cryoprotectant (e.g., 2% trehalose).
B. Characterization:
- Size & Zeta Potential: Measure by DLS in 1 mM KCl.
- Ligand Density Quantification: Use a fluorescence-based assay (e.g., OPA assay for residual lysine on peptide) or radiolabeled peptide to calculate surface ligand number.
- siRNA Loading: Quantify via HPLC after nanoparticle dissolution in DMSO or using a dye displacement assay (e.g., RiboGreen).
Protocol 3: In Vitro Evaluation of Targeted Cellular Uptake and Gene Knockdown
Objective: To validate the functional activity of ligand-conjugated nanoparticles.
Materials:
- Target cell line (e.g., U87MG glioblastoma for αvβ3 integrin)
- Control cell line (low receptor expression)
- Fluorescently labeled siRNA (e.g., Cy5-siRNA)
- Flow cytometer
- Confocal microscope
- qRT-PCR reagents for target mRNA quantification
- Western blot reagents for target protein quantification
Procedure:
- Cellular Uptake (Flow Cytometry):
- Seed cells in 12-well plates (2x10⁵ cells/well). Grow overnight.
- Treat with Cy5-labeled siRNA loaded in targeted (cRGDfK) and non-targeted NPs (equivalent siRNA dose, e.g., 100 nM). Include free Cy5-siRNA as control.
- Incubate 4 h at 37°C.
- Wash cells extensively with cold PBS, trypsinize, and resuspend in PBS+2% FBS.
- Analyze Cy5 fluorescence per cell using a flow cytometer (Ex/Em 649/670 nm). Calculate mean fluorescence intensity (MFI) shift.
Cellular Uptake (Confocal Microscopy):
- Seed cells on glass-bottom dishes.
- Treat with NPs as above. After 2-4 h, wash, fix with 4% PFA, and stain actin/phalloidin and nuclei/DAPI.
- Image using a confocal microscope to visualize intracellular NP localization.
Gene Knockdown Efficacy (qRT-PCR/Western Blot):
- Seed cells as above.
- Treat with NPs loaded with therapeutic siRNA (e.g., anti-GFP, anti-survivin). Use a scrambled siRNA NP control.
- After 48-72 h, harvest cells.
- Extract total RNA for cDNA synthesis and qRT-PCR analysis of target mRNA levels.
- Or, lyse cells for Western blot analysis to quantify reduction in target protein levels. Normalize to GAPDH or β-actin.
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for Targeted PEG-PLGA Nanoparticle Fabrication
| Reagent / Material | Supplier Examples | Function in Experiment |
|---|---|---|
| PLGA-COOH (Resomer series) | Merck, Evonik, Lactel Absorbable Polymers | The core biodegradable polymer providing nanoparticle structure and sustained release properties. |
| Heterobifunctional PEG (e.g., HO-PEG-NH₂, MAL-PEG-NHS) | Iris Biotech, JenKem Technology, Creative PEGWorks | The linker/spacer that provides stealth properties and a functional handle for ligand conjugation. |
| Targeting Peptides (cRGDfK, iRGD, TAT) | Peptide Specialty Laboratories, GenScript, Bachem | The active targeting ligand that confers specificity to cell-surface receptors. |
| siRNA (custom, gene-specific) | Horizon Discovery, Integrated DNA Technologies, Qiagen | The therapeutic nucleic acid cargo for gene silencing applications. |
| Polyvinyl Alcohol (PVA) | Merck, Sigma-Aldrich | A common stabilizer/surfactant used in emulsion methods to control nanoparticle size and prevent aggregation. |
| Tris(2-carboxyethyl)phosphine (TCEP) | Thermo Fisher Scientific | A reducing agent used to cleave disulfide bonds and generate free thiols on ligands for maleimide conjugation. |
| NHS/EDC or DCC Coupling Reagents | Tokyo Chemical Industry, Merck | Carbodiimide-based catalysts for forming amide bonds between carboxyl and amine groups during polymer modification. |
| Dialysis Tubing (MWCO 3.5-14 kDa) | Repligen, Spectrum Labs | For purifying polymers or nanoparticles from organic solvents and small-molecule impurities. |
Visualization Diagrams
Diagram 1: Mechanism of Ligand-Targeted Nanoparticle Uptake
Diagram 2: Workflow for Preparing Targeted siRNA NPs
Within the broader thesis investigating PEG-PLGA nanoparticles for sustained gene release, the effective encapsulation and protection of distinct genetic payloads—plasmid DNA (pDNA), small interfering RNA (siRNA), and messenger RNA (mRNA)—is a foundational challenge. Each molecule presents unique physicochemical characteristics and stability requirements that directly influence nanoparticle formulation strategy, loading efficiency, and ultimately, the sustained release profile and therapeutic efficacy.
Comparative Payload Characteristics & Loading Data
The following table summarizes key attributes and quantitative loading data for the three genetic payload types using double emulsion (W/O/W) solvent evaporation methods with PEG-PLGA.
Table 1: Characteristics and Loading Efficiencies of Genetic Payloads in PEG-PLGA Nanoparticles
| Payload Type | Typical Size (nt/bp) | Charge (at pH 7) | Key Stability Challenge | Avg. Loading Efficiency (%)* | Avg. Encapsulation Efficiency (%)* | Sustained Release Duration (Days)* |
|---|---|---|---|---|---|---|
| Plasmid DNA | 3000-10000 bp | Negative | Shear degradation, nuclease activity | 2.5 - 4.0 | 65 - 85 | 14 - 28 |
| siRNA | 19-23 bp (duplex) | Negative | Nuclease activity, rapid renal clearance | 1.0 - 2.5 | 70 - 90 | 7 - 21 |
| mRNA | 500-5000 nt | Negative | Hydrolysis, nuclease activity, innate immune activation | 1.5 - 3.5 | 60 - 80 | 5 - 14 |
*Data compiled from recent literature (2023-2024) using standard PEG(5k)-PLGA(50:50) formulations. Efficiency ranges account for variations in payload size, encapsulation method optimization, and PLGA molecular weight.
Detailed Protocols
Protocol 1: Primary Water-in-Oil (W/O) Emulsion Formation for pDNA
Objective: To efficiently incorporate pDNA into the hydrophobic PLGA polymer phase.
- Dissolve PEG-PLGA: Dissolve 100 mg of PEG(5k)-PLGA(50:50, 24-38 kDa) in 2 mL of dichloromethane (DCM) in a glass vial (Organic Phase).
- Prepare Aqueous Payload: Dilute 100 µg of purified pDNA (e.g., gWiz GFP) in 100 µL of nuclease-free water. For complexation, mix with 10 µL of 1 mg/mL poly-L-lysine (PLL, 15-30 kDa) solution and incubate 15 min at room temperature to form coacervates (Aqueous Phase 1).
- Primary Emulsification: Add the Aqueous Phase 1 to the Organic Phase. Immediately probe sonicate (e.g., 70% amplitude, 30 seconds, pulse mode 5 sec on/2 sec off) over an ice bath.
- Product: A stable, milky white W/O primary emulsion.
Protocol 2: Double Emulsion (W/O/W) Solvent Evaporation for siRNA/mRNA
Objective: To form nanoparticles with high encapsulation efficiency for siRNA or mRNA.
- Prepare Primary W/O Emulsion: Follow Protocol 1, but replace pDNA solution with 100 µL of nuclease-free water containing 50 µg of siRNA (e.g., targeting GFP) or 50 µg of mRNA. Cationic complexation agents (e.g., PLL) can be omitted for mRNA to minimize interference with translation.
- Prepare External Aqueous Phase: Add 4 mL of 2% (w/v) polyvinyl alcohol (PVA, 30-70 kDa) solution in a 50 mL beaker.
- Secondary Emulsification: Pour the primary W/O emulsion into the stirring (500 rpm) external PVA solution. Immediately homogenize at 13,000 rpm for 2 minutes using a high-speed homogenizer.
- Solvent Evaporation & Harvest: Stir the double emulsion gently (300 rpm) overnight at room temperature to evaporate DCM. Concentrate and wash nanoparticles via centrifugation (21,000 x g, 30 min, 4°C) three times with nuclease-free water. Resuspend the final pellet in 1 mL of PBS or trehalose solution (5% w/v) for lyophilization.
Protocol 3: In Vitro Nuclease Protection Assay
Objective: To validate the protective capacity of PEG-PLGA nanoparticles.
- Treat Samples: Incubate 10 µg of each (free payload vs. encapsulated payload) with 1 µL of DNase I (for pDNA) or RNase A (for siRNA/mRNA) in 1x reaction buffer at 37°C for 30 minutes.
- Release Payload: Add 100 µL of 0.1 N NaOH with 1% SDS to nanoparticle samples, vortex, and incubate for 1 hour at 4°C to degrade polymer and release payload.
- Analyze Integrity: Neutralize with 1 M Tris-HCl (pH 7.4). Run all samples on a 1% agarose gel (for pDNA) or a 2% agarose/EtBr gel (for siRNA/mRNA) alongside untreated controls.
- Expected Result: Free payloads show complete degradation, while encapsulated payloads show intact bands, confirming protection.
Visualizations
Diagram Title: Nanoparticle Formulation & Release Workflow
Diagram Title: Nanoparticle Protection & Intracellular Action Pathway
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for PEG-PLGA Genetic Payload Formulation
| Item | Function/Benefit | Example (Supplier) |
|---|---|---|
| PEG-PLGA Copolymer | Forms nanoparticle matrix; PEG provides stealth, PLGA controls biodegradation & release kinetics. | PEG(5k)-PLGA(50:50), Acid-terminated (Akina, Inc.) |
| Cationic Complexation Agent | Condenses negatively charged nucleic acids, improves loading efficiency and stability. | Poly-L-lysine hydrobromide (PLL, 15-30 kDa) (Sigma-Aldrich) |
| Stabilizing Surfactant | Forms stable emulsion during nanoparticle fabrication, prevents aggregation. | Polyvinyl Alcohol (PVA, 30-70 kDa, 87-89% hydrolyzed) (Sigma-Aldrich) |
| Nuclease Inhibitors | Critical for protecting payloads during formulation before encapsulation. | Recombinant RNase Inhibitor (Takara Bio) |
| Lyophilization Protectant | Prevents nanoparticle aggregation and payload degradation during freeze-drying for storage. | Trehalose, Dihydrate (Fisher Scientific) |
| Purified Genetic Payloads | High-purity, endotoxin-free inputs are essential for reproducible loading and biological activity. | siRNA (Horizon Discovery), CleanCap mRNA (TriLink BioTechnologies) |
This application note provides detailed protocols for evaluating the sustained release of genetic material (e.g., plasmid DNA, siRNA) from Poly(ethylene glycol)-poly(lactic-co-glycolic acid) (PEG-PLGA) nanoparticles. These methodologies are integral to the broader thesis research aiming to develop and optimize PEG-PLGA nanocarriers for prolonged gene delivery in regenerative medicine and cancer therapy. The protocols are designed to characterize release kinetics in vitro and therapeutic efficacy in vivo.
Research Reagent Solutions Toolkit
| Reagent/Material | Function in Sustained Release Research |
|---|---|
| PEG-PLGA Copolymer (e.g., PLGA-PEG-COOH) | Forms the nanoparticle matrix; PLGA controls biodegradation rate, PEG provides "stealth" properties to reduce opsonization. |
| Double-Emulsion Solvent Evaporation | Key synthesis method for encapsulating hydrophilic genetic material within PEG-PLGA nanoparticles. |
| PicoGreen / RiboGreen Assay | Fluorescent quantification of encapsulated or released DNA/RNA without interference from nanoparticles. |
| Dialysis Sack (MWCO 50-100 kDa) | Standard apparatus for conducting in vitro release studies under sink conditions. |
| HEPES-buffered saline (pH 7.4) | Standard release medium that maintains physiological pH and ionic strength. |
| Murine Model (e.g., BALB/c nude mice) | Common in vivo model for evaluating sustained gene expression or tumor suppression over weeks. |
| IVIS Imaging System | Enables non-invasive, longitudinal tracking of luciferase reporter gene expression in vivo. |
| Enzyme-Linked Immunosorbent Assay (ELISA) | Quantifies sustained protein expression (e.g., therapeutic protein) from the delivered gene over time. |
In Vitro Release Protocol
Protocol 3.1: Direct Sampling Method for Release Kinetics
Objective: To quantify the cumulative release of genetic material from PEG-PLGA nanoparticles over time in a controlled buffer.
Materials:
- Synthesized PEG-PLGA nanoparticles loaded with DNA/siRNA.
- Release medium: 1x PBS or HEPES-buffered saline (pH 7.4) with 0.01% w/v sodium azide.
- Thermostated shaking water bath (37°C, 100 rpm).
- Centrifuge with cooling (4°C).
- Microfuge tubes, PicoGreen reagent, plate reader.
Procedure:
- Dispersion: Suspend 5 mg of freeze-dried nanoparticles in 5 mL of pre-warmed release medium in a sterile tube. This is your release vessel.
- Incubation: Place the vessel in a shaking water bath at 37°C ± 0.5°C.
- Sampling: At predetermined time points (e.g., 1, 4, 8, 24, 48 hours, then daily for 2-4 weeks), centrifuge 500 µL aliquot from the vessel at 21,000 x g for 30 minutes at 4°C.
- Quantification: Carefully collect 300 µL of the supernatant without disturbing the pellet. Analyze the supernatant for released nucleic acid using the PicoGreen (for DNA) assay per manufacturer's instructions.
- Replenishment: After each sampling, add 500 µL of fresh, pre-warmed release medium to the main vessel to maintain sink conditions.
- Data Calculation: Calculate cumulative release percentage against a standard curve of known nucleic acid concentration and the total loaded amount.
Protocol 3.2: Dialysis Bag Method
Objective: To physically separate nanoparticles from the release medium, allowing for complete medium change.
Procedure:
- Bag Preparation: Hydrate a dialysis membrane (MWCO 50-100 kDa) in release medium for 30 minutes.
- Loading: Place 2 mL of nanoparticle suspension (containing ~1 mg nanoparticles) inside the bag. Seal both ends securely.
- Immersion: Immerse the bag in 50 mL of release medium in a glass bottle. Place the bottle in the shaking water bath (37°C, 100 rpm).
- Sampling: At each time point, completely replace the external release medium with 50 mL of fresh, pre-warmed medium. Store the collected medium for analysis.
- Analysis: Quantify the nucleic acid in the collected release medium using the PicoGreen/RiboGreen assay.
Table 1: Representative In Vitro Release Data for PEG-PLGA-DNA Nanoparticles
| Time Point (Day) | Cumulative Release % (Direct Method) | Cumulative Release % (Dialysis Method) | Key Phase Identified |
|---|---|---|---|
| 0.5 | 18.5 ± 3.2 | 15.8 ± 2.7 | Initial Burst |
| 1 | 25.1 ± 4.1 | 22.4 ± 3.5 | - |
| 3 | 41.7 ± 5.3 | 38.9 ± 4.8 | Lag/Diffusion Phase |
| 7 | 65.3 ± 6.8 | 60.2 ± 5.9 | - |
| 14 | 82.4 ± 7.5 | 78.6 ± 7.1 | Erosion-Controlled Phase |
| 21 | 94.8 ± 8.1 | 91.3 ± 8.3 | - |
Diagram 1: In Vitro Release Testing Workflow
In Vivo Efficacy Protocol
Protocol 4.1: Longitudinal Gene Expression in a Tumor Xenograft Model
Objective: To assess the sustained production of a therapeutic protein (via gene delivery) over time following a single administration of PEG-PLGA nanoparticles.
Materials:
- Athymic nude mice with established subcutaneous tumors (~100 mm³).
- PEG-PLGA nanoparticles loaded with plasmid DNA encoding firefly luciferase (for imaging) and a therapeutic gene (e.g., p53).
- In vivo imaging system (IVIS).
- ELISA kits for therapeutic protein.
- Animal scale, calipers, injection supplies.
Procedure:
- Dosing: Randomize mice into groups (n=5-8). Administer a single intratumoral or intravenous injection of nanoparticle formulation. Control groups receive empty NPs or free plasmid.
- Longitudinal Imaging: For luciferase expression, inject mice intraperitoneally with D-luciferin (150 mg/kg) 10 minutes prior to imaging. Anesthetize and image mice using IVIS at days 1, 3, 7, 14, 21, and 28 post-injection.
- Tumor Monitoring: Measure tumor dimensions with calipers every 2-3 days. Calculate volume using the formula: V = (length x width²)/2.
- Terminal Analysis: At endpoint (e.g., day 28), euthanize animals. Collect tumors and major organs. Homogenize tissues to quantify therapeutic protein levels via ELISA and perform histology.
Table 2: Representative In Vivo Efficacy Data (Tumor Volume & Gene Expression)
| Day Post-Injection | Tumor Volume (mm³)\nFree DNA | Tumor Volume (mm³)\nPEG-PLGA-DNA NPs | Luminescence (p/s/cm²/sr)\nPEG-PLGA-DNA NPs |
|---|---|---|---|
| 0 | 105 ± 12 | 108 ± 10 | 5.2e3 ± 1.1e3 |
| 3 | 180 ± 25 | 155 ± 18 | 2.8e5 ± 4.5e4 |
| 7 | 320 ± 40 | 210 ± 22 | 1.9e6 ± 3.1e5 |
| 14 | 650 ± 85 | 280 ± 35 | 8.5e5 ± 1.2e5 |
| 21 | 1200 ± 150 | 410 ± 55 | 3.2e5 ± 5.5e4 |
| 28 | >1500 (Endpoint) | 520 ± 70 | 1.1e5 ± 2.1e4 |
Diagram 2: In Vivo Sustained Gene Delivery & Action Pathway
Optimizing Formulation Stability and Efficacy: Troubleshooting Common Issues
Addressing Poor Encapsulation Efficiency and Rapid Burst Release
Within the broader thesis on developing PEG-PLGA nanoparticles for sustained gene release, two critical barriers impede clinical translation: Poor Encapsulation Efficiency (EE) of nucleic acids (e.g., pDNA, siRNA) and a Rapid Burst Release profile. This document provides application notes and detailed protocols to diagnose, troubleshoot, and mitigate these issues, enabling the development of nanoparticles with high payload retention and near-zero-order release kinetics suitable for gene therapy.
Key Challenges and Diagnostic Data
Table 1: Common Causes and Quantitative Impact on Encapsulation Efficiency (EE) and Burst Release
| Factor | Typical Impact on EE (%) | Typical Impact on Burst Release (0-24h, %) | Key Diagnostic Assay |
|---|---|---|---|
| Inadequate Nucleic Acid-Complexant Ratio | 10-40 | 60-90 | Gel Retardation / PicoGreen |
| Large Aqueous Phase Volume (w/o/w) | 15-50 | 50-85 | Volume Optimization Series |
| High Surfactant Concentration (e.g., PVA %) | 20-60 | 40-75 | Particle Size vs. EE Analysis |
| Low Molecular Weight PLGA | 30-70 | 30-60 | GPC Analysis of Polymer |
| Improper Solvent Removal Rate | 25-55 | 50-80 | Release Kinetics Profile |
| Absence of Cationic/Complexing Agent | 5-25 | 70-95 | Zeta Potential Measurement |
| Poor Polymer:PEG Ratio | 40-80 | 20-50 | Surface Chemistry (XPS) |
Core Protocols for Optimization
Protocol 3.1: Optimized Double Emulsion (w/o/w) for High EE
Aim: Encapsulate pDNA/siRNA with EE > 80%. Reagents: PLGA-PEG (e.g., 15kDa PLGA-5kDa PEG), Dichloromethane (DMC), Polyvinyl Alcohol (PVA, 1% w/v), Spermidine or Chitosan (as complexant), Tris-EDTA buffer. Procedure:
- Inner Aqueous Phase (W1): Dilute 50 µg of nucleic acid in 100 µL of nuclease-free water containing 0.25 mg spermidine. Vortex gently.
- Oil Phase (O): Dissolve 100 mg PLGA-PEG in 2 mL of DCM.
- Primary Emulsion (W1/O): Add W1 to O. Probe sonicate (20% amplitude, 30 s, pulse 2s on/1s off) in an ice bath.
- External Aqueous Phase (W2): Add the primary emulsion to 4 mL of 1% w/v PVA solution. Homogenize at 13,000 rpm for 2 minutes.
- Solvent Evaporation: Stir the double emulsion at 500 rpm for 4 hours at room temperature to evaporate DCM.
- Collection: Ultracentrifuge at 30,000 x g for 25 min at 4°C. Wash pellet 3x with water. Resuspend in formulation buffer.
- EE Quantification: Use a Quant-iT PicoGreen assay. Measure free nucleic acid in supernatant vs. total.
Protocol 3.2: Minimizing Burst Release via Cross-linking or Coatings
Aim: Achieve <20% release in first 24h, sustained release over >14 days. Reagents: Ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC), N-Hydroxysuccinimide (NHS), Chitosan (low MW), Phosphate Buffered Saline (PBS, pH 7.4). Procedure:
- Prepare nanoparticles per Protocol 3.1.
- Surface Cross-linking: Resuspend NP pellet in 2 mL MES buffer (pH 6.0). Add EDC and NHS (molar ratio 2:1 to surface -COOH groups). React for 2h on a shaker.
- Secondary Coating: Alternatively, incubate NPs with 0.1% w/v chitosan in acetate buffer (pH 5.5) for 30 min.
- Release Study: Place 5 mg of NPs in 1 mL of PBS + 0.01% Tween 20 in a dialysis tube (MWCO 100kDa). Immerse in 30 mL release medium at 37°C with gentle shaking.
- Sampling: At predetermined intervals, collect 1 mL of external medium and replace with fresh pre-warmed medium.
- Quantification: Assay samples for nucleic acid content (PicoGreen for dsDNA, RiboGreen for RNA). Plot cumulative release.
Visualization of Workflows and Mechanisms
Diagram 1: Problem Analysis & Solution Pathways
Diagram 2: Optimized Double Emulsion Workflow
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions
| Reagent / Material | Function & Rationale | Key Consideration |
|---|---|---|
| PLGA-PEG (di-block copolymer) | Forms nanoparticle core (PLGA) and provides "stealth" hydrophilic corona (PEG) to reduce opsonization and burst release. | PEGylation ratio (5-20% w/w) is critical for stealth vs. EE balance. |
| Cationic Complexant (Spermidine, Chitosan, PEI) | Electrostatically condenses negatively charged nucleic acids, preventing aqueous phase leakage during emulsification. | Charge ratio (N/P) must be optimized to balance complexation and cytotoxicity. |
| Polyvinyl Alcohol (PVA) | Most common surfactant/stabilizer in emulsion methods. Controls particle size and polydispersity. | Residual PVA affects surface properties; cold washing reduces it. |
| Dichloromethane (DCM) / Ethyl Acetate | Organic solvent for PLGA. DCM offers higher EE; Ethyl Acetate is less toxic. | Evaporation rate directly impacts nanoparticle porosity and burst release. |
| Carbodiimide Cross-linkers (EDC/NHS) | Activates surface -COOH groups (from PLGA terminal acid or added polymers) for covalent cross-linking to reduce initial burst. | Must be performed in MES buffer (pH 5-6) for optimal efficiency. |
| Quant-iT PicoGreen Assay | Ultra-sensitive fluorescent nucleic acid quantitation. Essential for measuring EE% (via supernatant) and release kinetics. | Measures only double-stranded DNA; use Ribogreen for RNA. |
| Dialysis Tubing (MWCO 100 kDa) | Standard method for in vitro release studies, allowing sink conditions to be maintained. | MWCO must be significantly larger than NPs but smaller than free nucleic acids. |
Strategies to Enhance Colloidal Stability and Prevent Aggregation
Application Notes Within the context of developing PEG-PLGA nanoparticles (NPs) for sustained gene release, colloidal stability is paramount. Aggregation compromises particle size distribution, cellular uptake kinetics, biodistribution, and release profiles. The primary strategies involve modulating surface properties and the dispersion medium to counteract van der Waals forces, the main driver of aggregation. Steric stabilization via PEGylation is the cornerstone, but it must be optimized and complemented by other approaches.
Quantitative Data Summary: Key Stabilizing Agents & Their Impact on PEG-PLGA NPs
Table 1: Stabilizers for PEG-PLGA Nanoparticle Formulations
| Stabilizer Class/Agent | Typical Conc. Range | Key Mechanism | Impact on Zeta Potential (mV) | Effect on PDI |
|---|---|---|---|---|
| Non-ionic Surfactant (Poloxamer 188) | 0.1 - 1.0 % w/v | Steric Hindrance | -1 to -5 (minimal change) | Reduces to <0.1 |
| Ionic Surfactant (SDS) | 0.01 - 0.1 % w/v | Electrostatic Repulsion | -30 to -50 | Can increase if >CMC |
| Polymeric Stabilizer (PVP K30) | 0.5 - 5 % w/v | Steric Hindrance | 0 to -10 | Reduces to <0.15 |
| Sugar (Trehalose) | 2 - 10 % w/v | Cryo/Lyoprotection | Maintains initial value | Maintains initial value |
| Charge Modifier (DOTAP) | 1 - 10 mol% (to polymer) | Electrostatic & Steric | +20 to +40 | Can increase if >10% |
Table 2: Formulation & Storage Stability Parameters
| Storage Condition | Key Stability Indicator | Target Value | Measurement Frequency |
|---|---|---|---|
| 4°C (Aqueous) | Hydrodynamic Diameter | <200 nm & Δ < 10% | Weekly for 1 month |
| 25°C (Aqueous) | Polydispersity Index (PDI) | <0.2 | Weekly for 1 month |
| Lyophilized | Zeta Potential | Δ < 5 mV from baseline | Pre & post-lyophilization |
| Reconstitution | Gene Loading Efficiency | >85% retention | Post-reconstitution |
Experimental Protocols
Protocol 1: Optimized Double-Emulsion Solvent Evaporation for Stable PEG-PLGA Gene-Loaded NPs Objective: To formulate colloidally stable, gene-loaded PEG-PLGA nanoparticles using a steric and electrostatic stabilization strategy. Materials: PEG-PLGA copolymer (e.g., 5% PEG), PLGA, Dichloromethane (DCM), Poloxamer 188, Chitosan HCl, pDNA or siRNA, Polyvinyl alcohol (PVA), Ultra-pure water. Procedure:
- Primary W1/O Emulsion: Dissolve 50 mg PEG-PLGA/PLGA blend in 2 mL DCM. Add 100 µL of an aqueous gene solution (e.g., 1 mg/mL pDNA in 5 mM chitosan solution) to the polymer solution. Sonicate on ice (70% amplitude, 30 s).
- Secondary W1/O/W2 Emulsion: Pour the primary emulsion into 8 mL of 2% w/v PVA and 0.5% w/v Poloxamer 188 aqueous solution. Homogenize (15,000 rpm, 2 min).
- Solvent Evaporation: Stir the double emulsion at room temperature for 4 hours to evaporate DCM.
- Purification: Centrifuge the suspension at 21,000 x g for 30 min. Wash pellet twice with ultra-pure water.
- Resuspension/Storage: Resuspend NPs in 2 mL of 5% w/v trehalose solution. Characterize size, PDI, and zeta potential. Store at 4°C or lyophilize for long-term storage.
Protocol 2: Assessment of Colloidal Stability via Accelerated Aggregation Studies Objective: To rapidly screen formulation stability under stress conditions. Materials: NP suspension, Phosphate Buffered Saline (PBS), 1 M NaCl solution, Thermonixer. Procedure:
- Prepare 1 mL aliquots of purified NP suspension in three media: (A) 5% trehalose, (B) 1X PBS, (C) 0.15 M NaCl.
- Place aliquots in a thermomixer at 37°C with constant shaking (500 rpm).
- Measure the hydrodynamic diameter (Dh) and PDI of each aliquot at t=0, 2, 6, 24, and 48 hours using dynamic light scattering.
- Stability Criterion: A formulation is considered stable if the change in Dh (ΔDh) is <20% and PDI remains <0.25 over 48 hours in high-ionic-strength media (PBS/NaCl).
Visualizations
Title: Protocol for Stable PEG-PLGA Nanoparticle Formulation
Title: Mechanisms Preventing Nanoparticle Aggregation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Formulating Stable PEG-PLGA Gene Vectors
| Reagent/Material | Primary Function | Critical Consideration for Stability |
|---|---|---|
| Diblock PEG-PLGA Copolymer | Provides inherent steric stabilization via surface-grafted PEG chains. | PEG molecular weight (2-5 kDa) and density (5-10% wt) are key. |
| Poloxamer 188 (Pluronic F-68) | Non-ionic surfactant; adsorbs to NP surface, providing additional steric barrier. | Prevents Oswald ripening and aggregation during emulsification. |
| Polyvinyl Alcohol (PVA) | Emulsifier and stabilizer during fabrication; forms residual layer on NP surface. | Degree of hydrolysis (87-89%) offers optimal stability. |
| Chitosan HCl | Cationic polyelectrolyte; condenses nucleic acids and imparts positive surface charge. | Low molecular weight (3-10 kDa) minimizes aggregation risks. |
| Trehalose Dihydrate | Lyoprotectant and bulking agent; forms glassy matrix, preventing fusion during freeze-drying. | 5-10% w/v ratio to solids is optimal for cake formation and re-dispersion. |
| DOTAP (cationic lipid) | Co-formulant; enhances gene complexation and adds electrostatic stabilization. | Molar ratio to polymer must be optimized to balance stability and cytotoxicity. |
| Sterile, Nuclease-Free Water | Dispersion medium for all aqueous phases and final resuspension. | Eliminates ionic contaminants and nucleases that degrade payloads. |
1. Introduction Within the context of a thesis on developing PEG-PLGA nanoparticles (NPs) for sustained gene delivery, precise control over the in vitro release kinetics of genetic payloads (e.g., pDNA, siRNA) is paramount. Two critical material parameters govern this control: the molecular weight (MW) of the PLGA block and the density of the PEG corona. This document provides application notes and detailed protocols for systematically investigating these parameters to achieve a target release profile.
2. Quantitative Parameter Impact Summary The following table synthesizes current research data on the effects of PEG density and PLGA MW on key nanoparticle characteristics and release kinetics.
Table 1: Impact of PEG Density and PLGA MW on Nanoparticle Properties & Release
| Parameter | Typical Range Tested | Effect on Nanoparticle Characteristics | Effect on In Vitro Release Profile (Model Payloads) |
|---|---|---|---|
| PEG Density | 2-10% (PEG-PLGA w/w) | Increased Density: Reduces particle size, improves colloidal stability, decreases protein opsonization, increases steric hindrance. | Higher Density: Prolongs initial lag phase, slows diffusion-based release, leads to a more sustained, linear profile. Burst release is minimized. |
| PLGA MW (kDa) | 10 - 100 kDa | Higher MW: Increases particle size, enhances mechanical integrity/rigidity of polymer matrix, slows degradation rate. | Higher MW: Significantly extends release duration. Reduces initial burst due to denser matrix. Primary release mechanism shifts toward bulk erosion. |
| Combined Effect | N/A | High PLGA MW with moderate PEG density (5-8%) optimizes for stability and controlled matrix erosion. | A high MW PLGA (e.g., 50-75 kDa) coupled with a PEG density of 5-8% is typically optimal for linear, sustained release over 3-6 weeks. |
3. Experimental Protocols
Protocol 3.1: Formulation of PEG-PLGA NPs with Varied PEG Density Objective: To prepare gene-loaded NPs using a series of PEG-PLGA copolymers with identical PLGA MW but varying PEGylation density (e.g., 2%, 5%, 10%). Materials: PEG-PLGA polymers (e.g., PEG5k-PLGA45k, PEG2k-PLGA45k, mPEG-PLGA), plasmid DNA (pDNA) or siRNA, polyvinyl alcohol (PVA), dichloromethane (DCM), deionized water, probe sonicator, magnetic stirrer. Procedure:
- Aqueous Phase: Dissolve 2% (w/v) PVA in DI water. Dissolve the gene payload in a separate portion of this PVA solution.
- Organic Phase: Dissolve 100 mg of the selected PEG-PLGA polymer in 2 mL of DCM.
- Primary Emulsion: Combine the organic phase with the aqueous gene/PVA solution. Emulsify using a probe sonicator (70% amplitude, 60 s on ice).
- Secondary Emulsion: Pour the primary emulsion into 20 mL of a 0.5% (w/v) PVA solution under vigorous stirring. Sonicate again (50% amplitude, 90 s on ice).
- Solvent Evaporation: Stir the double emulsion overnight at room temperature to evaporate DCM.
- Purification: Centrifuge the NP suspension at 21,000 x g for 30 min at 4°C. Wash the pellet three times with DI water.
- Characterization: Re-suspend NPs in buffer. Determine size (PDI) via DLS, zeta potential, and encapsulation efficiency (using a dye-binding assay for pDNA or HPLC for siRNA).
Protocol 3.2: Formulation of PEG-PLGA NPs with Varied PLGA MW Objective: To prepare NPs using PEG-PLGA copolymers with similar PEG density but different PLGA MWs (e.g., PEG5k-PLGA20k, PEG5k-PLGA50k). Materials: PEG-PLGA polymers with varying PLGA blocks, other materials as in Protocol 3.1. Procedure: Follow Protocol 3.1, substituting polymers. Critical Note: Adjust the organic-to-aqueous phase ratio or sonication energy to achieve consistent particle size across different MW batches, as higher MW PLGA tends to form larger NPs.
Protocol 3.3: In Vitro Release Study & Kinetic Modeling Objective: To quantify payload release over time and fit data to mathematical models. Materials: Dialysis cassettes (e.g., 10 kDa MWCO), release medium (PBS, pH 7.4 ± enzymes), microplate reader or spectrophotometer. Procedure:
- Place 2 mL of purified NP suspension (known payload amount) into a dialysis cassette.
- Immerse the cassette in 200 mL of release medium at 37°C with gentle shaking.
- At predetermined time points, collect 1 mL of the external medium and replace with fresh pre-warmed medium.
- Quantify the released payload concentration using an appropriate assay (e.g., fluorescence for labeled nucleic acids, picogreen for pDNA).
- Data Analysis: Calculate cumulative release (%) and plot vs. time. Fit data to models:
- Zero-Order:
Q = k0 * t(ideal for sustained release). - First-Order:
ln(100 - Q) = ln(100) - k1 * t. - Higuchi:
Q = kH * sqrt(t)(for diffusion-controlled release). - Korsmeyer-Peppas:
Q / Q∞ = k * t^n(determine release mechanism vianvalue).
- Zero-Order:
4. The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for PEG-PLGA Gene Delivery Optimization
| Item | Function/Explanation |
|---|---|
| PEG-PLGA Copolymers | Core material. Libraries with defined PEG% (2k-5k Da) and PLGA MW (10-100k Da) & LA:GA ratio (e.g., 50:50, 75:25) are essential for systematic study. |
| Polyvinyl Alcohol (PVA) | A common stabilizer/emulsifier in double emulsion formulations. MW and degree of hydrolysis affect NP size and surface properties. |
| Dialysis Cassettes (3.5-10 kDa MWCO) | Standard tool for in vitro release studies, allowing buffer exchange while retaining NPs. |
| Fluorescent Nucleic Acid Labels (e.g., Cy5, FITC) | Enable rapid tracking of encapsulation efficiency and release kinetics via fluorescence assays. |
| PicoGreen / RiboGreen Assay Kits | Quantify double-stranded DNA or RNA with high sensitivity, crucial for measuring encapsulation and release. |
| Size Exclusion Chromatography (SEC) Columns | For precise purification of NPs from free polymer, unencapsulated payload, and emulsifier. |
5. Visualization of Experimental Workflow & Release Mechanisms
Title: Workflow for Optimizing Nanoparticle Release Profiles
Title: Material Parameters Influence Release Mechanisms
Within the thesis research focused on developing PEG-PLGA nanoparticles for sustained gene release, a central challenge is achieving efficient transfection. This process is hindered by multiple biological barriers, including serum degradation, cellular uptake, endosomal entrapment, and nuclear entry. This application note details current techniques and protocols to overcome these barriers, specifically tailored for non-viral, polymeric nanoparticle systems.
Key Biological Barriers and Overcoming Strategies
The journey of a gene vector from administration to functional expression involves navigating a series of obstacles. The table below summarizes the primary barriers and corresponding techniques to enhance PEG-PLGA nanoparticle transfection.
Table 1: Biological Barriers and Techniques for PEG-PLGA Nanoparticle Transfection
| Biological Barrier | Impact on Transfection | Overcoming Technique | Typical Efficacy Improvement (vs. baseline) |
|---|---|---|---|
| Serum & Enzymatic Degradation | Nuclease degradation of payload; opsonization. | PEGylation; chitosan coating; tune PEG density. | ~50-80% payload protection increase. |
| Cellular Uptake | Low internalization rate; non-specific uptake. | Ligand conjugation (e.g., RGD, folate); charge optimization (+15 to +25 mV). | 2-5 fold increase in cellular association. |
| Endosomal Entrapment | Lysosomal degradation of nucleic acid. | Proton-sponge polymers (PEI); endosomolytic peptides (HA2); chloroquine. | 3-10 fold boost in gene expression. |
| Cytoplasmic Transport & Nuclear Entry | Poor cytoplasmic mobility; inefficient nuclear import. | NLS peptide conjugation; microtubule-targeting agents. | 2-4 fold increase for non-dividing cells. |
Detailed Experimental Protocols
Protocol 1: Formulation of Ligand-Targeted PEG-PLGA Nanoparticles
Objective: Synthesize folate-conjugated PEG-PLGA nanoparticles for enhanced receptor-mediated uptake in cancer cells.
Materials:
- PLGA (50:50, acid-terminated)
- NH2-PEG-COOH
- Folic Acid
- N-Hydroxysuccinimide (NHS)
- N-(3-Dimethylaminopropyl)-N′-ethylcarbodiimide (EDC)
- Dichloromethane (DCM)
- Polyvinyl Alcohol (PVA)
- Probe sonicator
- Centrifugal filter devices (100 kDa MWCO)
Method:
- Synthesis of Folate-PEG-PLGA Conjugate:
- Activate 10 mg of folic acid with 20 mg EDC and 12 mg NHS in 2 mL DMSO for 2 hrs.
- Add the activated solution dropwise to 100 mg of NH2-PEG-COOH in 3 mL PBS (pH 7.4). Stir for 12 hrs at RT.
- Purify via dialysis (MWCO 3.5 kDa) against water. Lyophilize to obtain Folate-PEG-COOH.
- Activate Folate-PEG-COOH with EDC/NHS and conjugate to PLGA-NH2 (pre-formed) in DCM. Precipitate, wash, and dry the final Folate-PEG-PLGA copolymer.
- Nanoparticle Preparation (Double Emulsion):
- Dissolve 50 mg of Folate-PEG-PLGA and 50 mg standard PLGA in 2 mL DCM.
- Add 200 µL of nuclease-free water containing 100 µg of plasmid DNA (pDNA) as the first aqueous phase.
- Emulsify using a probe sonicator (50 W, 30 s) on ice to form the primary W/O emulsion.
- Add this emulsion to 8 mL of 4% (w/v) PVA solution. Sonicate again (60 W, 60 s) to form a W/O/W double emulsion.
- Stir overnight to evaporate DCM. Centrifuge at 18,000 × g for 30 min, wash pellets 3x with water, and resuspend in PBS. Filter through a 0.45 µm filter. Characterize size, zeta potential, and DNA encapsulation efficiency (PicoGreen assay).
Protocol 2: Assessing Endosomal Escape via Fluorescence Microscopy
Objective: Quantify the endosomal escape efficiency of nanoparticles using a split-GFP reporter assay.
Materials:
- HeLa cells
- pDNA encoding GFP11-tagged protein
- Nanoparticles with/without endosomolytic agent (e.g., 10% w/w PEI blended)
- Confocal microscope
- LysoTracker Deep Red
- Image analysis software (e.g., ImageJ)
Method:
- Seed HeLa cells stably expressing GFP1-10 in a glass-bottom dish.
- Treat cells with nanoparticles (loaded with pDNA-GFP11) at 1 µg pDNA/mL. Include a chloroquine (100 µM) positive control.
- After 4 hrs, replace with fresh media. At 24 hrs post-transfection, add LysoTracker (75 nM) for 1 hr.
- Image using confocal microscopy (488 nm for GFP, 647 nm for LysoTracker).
- Analyze colocalization using Pearson's correlation coefficient. Lower coefficient indicates superior endosomal escape.
Visualizing Key Pathways and Workflows
Title: Sequential Biological Barriers and Transfection Techniques
Title: Workflow for Optimizing Transfection Efficiency
The Scientist's Toolkit
Table 2: Key Research Reagent Solutions for Transfection Enhancement
| Reagent/Material | Supplier Examples | Function in Transfection Enhancement |
|---|---|---|
| PLGA (50:50, acid term.) | Sigma-Aldrich, Lactel Absorbable Polymers | Core biodegradable polymer for nanoparticle formation and sustained release. |
| NH2-PEG-COOH | JenKem Technology, Sigma-Aldrich | Provides reactive groups for ligand conjugation and imparts steric stability (stealth effect). |
| Branched PEI (25 kDa) | Polysciences, Sigma-Aldrich | "Proton-sponge" polymer blended into NPs to disrupt endosomes and promote escape. |
| Cell-Penetrating Peptides (e.g., TAT) | AnaSpec, GenScript | Conjugated to NPs to enhance cellular internalization and nuclear import. |
| PicoGreen Assay Kit | Thermo Fisher Scientific | Quantifies encapsulated or released DNA with high sensitivity, critical for EE% calculation. |
| LysoTracker Dyes | Thermo Fisher Scientific | Fluorescent probes to label acidic organelles (lysosomes/endosomes) for colocalization studies. |
| Fetal Bovine Serum (FBS) | Gibco, Sigma-Aldrich | Used in stability and transfection media to simulate in vivo protein interactions. |
| Polyvinyl Alcohol (PVA) | Sigma-Aldrich | Common surfactant/stabilizer used in emulsion-based nanoparticle formulation. |
Application Notes
This document details a systematic pathway for scaling up the production of PEG-PLGA nanoparticles (NPs) for sustained gene release, transitioning from benchtop research to Good Manufacturing Practice (GMP)-compliant processes. The core challenge lies in maintaining critical quality attributes (CQAs)—such as particle size, polydispersity index (PDI), zeta potential, encapsulation efficiency (EE%), and sustained release kinetics—while increasing batch size and implementing rigorous controls.
1. Key Scale-Up Parameters and Process Analytical Technology (PAT) Successful scale-up requires identifying and monitoring parameters that influence CQAs. The table below contrasts bench and GMP-scale considerations.
Table 1: Comparison of Bench-Scale and GMP-Scale Production Parameters for PEG-PLGA Gene Nanoparticles
| Parameter | Bench-Scale (Lab) | GMP-Scale (Pilot/Production) | Impact on CQAs |
|---|---|---|---|
| Batch Volume | 10 - 100 mL | 10 - 100 L | Affects mixing dynamics, shear forces, and heat transfer. |
| Method | Single-/Double-Emulsion Solvent Evaporation | Homogenization (HPH/Microfluidizer) & Evaporation | Determines particle size distribution and encapsulation efficiency. |
| Equipment | Probe Sonicator, Magnetic Stirrer | High-Pressure Homogenizer (HPH), In-Line Mixers, Reactor Vessels | Provides reproducible shear and energy input. |
| Process Control | Manual, discrete sampling | Automated, in-line PAT (e.g., DLS, NIR) | Ensures batch-to-batch consistency and real-time monitoring. |
| Purification | Centrifugation, Dialysis | Tangential Flow Filtration (TFF) | Efficient, scalable buffer exchange and concentration. |
| Environment | Laboratory Hood (Class II) | Cleanrooms (Grade B/C) | Controls bioburden and endotoxin levels. |
| Raw Materials | Research-grade reagents | GMP-grade, certified reagents with TSE/BSE statements | Ensures safety, traceability, and quality. |
| Documentation | Laboratory notebook | Batch Manufacturing Record (BMR), Standard Operating Procedures (SOPs) | Ensures regulatory compliance and traceability. |
2. Protocol: Scale-Up of PEG-PLGA Nanoparticles Loaded with pDNA via Double Emulsion (W/O/W) Objective: To produce a 10-liter GMP-compliant batch of PEG-PLGA nanoparticles encapsulating plasmid DNA (pDNA) for sustained release.
Materials (Research Reagent Solutions Toolkit): Table 2: Essential Research Reagent Solutions for PEG-PLGA Nanoparticle Scale-Up
| Reagent/Material | Function | GMP Consideration |
|---|---|---|
| GMP-grade PLGA-PEG | Copolymer forming the nanoparticle matrix, PEG ensures stealth properties. | Certificate of Analysis (CoA) required for MW, LA:GA ratio, PEG length, endotoxin. |
| GMP-grade pDNA | The therapeutic gene payload. | Must be produced in a GMP facility, with CoA for identity, purity, and potency. |
| Dichloromethane (DCM) or Ethyl Acetate | Organic solvent for polymer dissolution. | Residual solvent must be controlled to ICH limits; ethyl acetate is often preferred for safety. |
| Polyvinyl Alcohol (PVA) | Surfactant stabilizing the primary and secondary emulsion. | GMP-grade, defined degree of hydrolysis and MW. |
| Sterile Water for Injection (WFI) | Aqueous phase for emulsions and purification. | Must be pyrogen-free. |
| Tangential Flow Filtration (TFF) System | For diafiltration and concentration of the nanoparticle suspension. | Validated system with appropriate molecular weight cut-off (MWCO) membranes. |
| Lyoprotectant (e.g., Sucrose, Trehalose) | Protects nanoparticle integrity during freeze-drying. | GMP-grade, included in final formulation buffer. |
Detailed Protocol:
A. Pre-Process Preparation (in Grade C Cleanroom)
- Solution Preparation: Dissolve GMP-grade PLGA-PEG in the chosen organic solvent (e.g., 5% w/v) under mixing. Prepare the inner aqueous phase containing the pDNA in WFI. Prepare a 1-2% w/v solution of PVA in WFI. Pre-cool/heater as needed.
- Equipment Setup & Sanitization: Assemble and sanitize the high-pressure homogenizer (HPH), in-line mixer, and reactor vessel according to SOPs. Calibrate in-line PAT probes (e.g., for pressure, temperature).
B. Primary Emulsion (W1/O) Formation
- Transfer the pDNA solution (W1) to a sealed reactor containing the polymer solution (O).
- Using an in-line high-shear mixer or initial HPH pass at low pressure (e.g., 2000 psi), create a coarse primary emulsion (W1/O). Maintain temperature control (<25°C).
C. Secondary Emulsion (W1/O/W2) Formation & Solvent Evaporation
- Transfer the primary emulsion into a larger reactor containing the PVA solution (W2) under controlled agitation.
- Pump the mixture through the HPH for 3-5 passes at a defined, optimized pressure (e.g., 8000-12000 psi) to form fine nanoparticles. Monitor particle size in-line after the final pass.
- Transfer the nano-emulsion to a temperature-controlled evaporation vessel with gentle stirring and vacuum application for 2-4 hours to remove organic solvent.
D. Purification & Formulation
- Concentrate and diafilter the nanoparticle suspension against WFI using a TFF system (e.g., 300 kDa MWCO) to remove free PVA, unencapsulated pDNA, and residual solvent. Achieve a target exchange volume of 10-15 diavolumes.
- Concentrate the retentate to the desired final nanoparticle concentration.
- Adjust the formulation by adding a lyoprotectant (e.g., 5% sucrose) and homogenize gently.
E. Sterile Filtration & Lyophilization
- Pass the final suspension through a 0.22 µm sterilizing-grade filter into sterile containers.
- Fill vials and lyophilize using a validated cycle to obtain a stable powder.
3. Analytical Control Strategy Implement a quality-by-design (QbD) approach. In-process controls (IPC) include in-line size monitoring. Release testing includes:
- Particle Size & PDI: Dynamic Light Scattering (DLS).
- Zeta Potential: Electrophoretic light scattering.
- Encapsulation Efficiency: Quantify unencapsulated pDNA in TFF permeate using a fluorescence assay (e.g., PicoGreen) and calculate EE%.
- DNA Integrity: Gel electrophoresis post-extraction from NPs.
- In Vitro Release: Incubate NPs in PBS (pH 7.4) at 37°C under sink conditions; sample and quantify pDNA release over 4-8 weeks.
- Sterility & Endotoxin: USP <71> Sterility Test and <85> Bacterial Endotoxins Test.
Visualizations
Scale-Up Workflow from Research to GMP
QbD: Relationship Between CMA, CPP, and CQA
Benchmarking Performance: Validation, Characterization, and Comparative Analysis
Application Notes: Characterization of PEG-PLGA Nanoparticles for Sustained Gene Release
The efficacy, safety, and release kinetics of PEG-PLGA nanoparticles (NPs) for gene delivery are critically dependent on their physicochemical properties. This suite of techniques provides a comprehensive analysis essential for rational formulation development.
- Dynamic Light Scattering (DLS): Determines the hydrodynamic diameter, polydispersity index (PDI), and zeta potential of the NP suspension. Size and PDI influence cellular uptake and biodistribution, while zeta potential indicates colloidal stability and potential for non-specific protein adsorption.
- Transmission Electron Microscopy (TEM): Provides direct visualization of NP morphology, core-shell structure (PEG coating), and confirms size measurements from DLS. It is crucial for assessing aggregation state and structural integrity.
- High-Performance Liquid Chromatography (HPLC): Quantifies the encapsulation efficiency (EE%) of the genetic payload (e.g., pDNA, siRNA) and analyzes in vitro release profiles. It also monitors polymer degradation products and payload integrity.
- Differential Scanning Calorimetry (DSC): Investigates the thermal properties of the NP matrix. It identifies the glass transition temperature (Tg) of PLGA, which affects drug release kinetics, and confirms the successful incorporation of PEG into the NP structure.
Table 1: Summary of Key Quantitative Parameters and Their Significance
| Technique | Key Parameters | Target Range for PEG-PLGA Gene NPs | Significance for Sustained Gene Release |
|---|---|---|---|
| DLS | Hydrodynamic Diameter (Z-Avg) | 80-200 nm | Optimizes cellular uptake and circulation time. |
| Polydispersity Index (PDI) | < 0.2 | Indicates a monodisperse, reproducible formulation. | |
| Zeta Potential (in water) | -30 mV to +10 mV | Negative surface (PLGA) or near-neutral (PEG-shielded) enhances stability and reduces opsonization. | |
| TEM | Core Diameter | Correlates with DLS size | Validates DLS data, confirms spherical morphology, and visualizes PEG corona. |
| HPLC | Encapsulation Efficiency (EE%) | > 85% | Maximizes therapeutic payload delivery, reduces waste. |
| Sustained Release Duration | Days to weeks | Core metric for sustained release formulation; depends on PLGA molecular weight & lactide:glycolide ratio. | |
| DSC | Glass Transition Temp. (Tg) | 40-50°C (for PLGA 50:50) | A lower Tg accelerates polymer erosion and gene release. Monitoring Tg shift confirms PEGylation. |
Experimental Protocols
Protocol 1: DLS Analysis of PEG-PLGA NPs
Objective: Determine hydrodynamic size distribution, PDI, and zeta potential. Materials: NP suspension (1 mg/mL in filtered DI water or 1mM KCl), disposable sizing cuvettes, disposable folded capillary zeta cells, DLS/Zeta Potential analyzer. Procedure:
- Dilute the NP suspension appropriately to achieve an optimal scattering intensity.
- Filter the sample through a 0.45 µm or 1 µm syringe filter to remove dust.
- For size measurement, load 1 mL into a sizing cuvette. Equilibrate at 25°C for 2 minutes.
- Measure with a backscatter detector angle (e.g., 173°). Perform minimum 3 runs.
- For zeta potential, load 0.8 mL into a folded capillary cell. Measure electrophoretic mobility and calculate zeta potential via the Smoluchowski equation. Perform minimum 6 runs.
Protocol 2: TEM Sample Preparation (Negative Staining)
Objective: Visualize NP morphology and size. Materials: Formvar/carbon-coated copper grids, 2% (w/v) aqueous uranyl acetate solution, filter paper. Procedure:
- Apply a 5-10 µL droplet of diluted NP suspension onto a glow-discharged TEM grid for 60 seconds.
- Wick away excess liquid with filter paper.
- Immediately apply a 5-10 µL droplet of 2% uranyl acetate stain for 30 seconds.
- Wick away excess stain and allow the grid to air-dry thoroughly.
- Image using a TEM operated at 80-100 kV.
Protocol 3: HPLC Analysis of siRNA Encapsulation Efficiency
Objective: Quantify the amount of siRNA encapsulated within PEG-PLGA NPs. Materials: Cationic ion-pairing HPLC system, C18 column, mobile phases (A: 0.1M TEAA, B: Acetonitrile), siRNA standard solutions, 1% (v/v) Triton X-100 in water. Procedure:
- Total siRNA Content: Dissolve 1 mg of lyophilized NPs in 1 mL of DMSO. Add 9 mL of 1% Triton X-100 to precipitate polymer and release siRNA. Vortex, centrifuge, and filter the supernatant for HPLC analysis.
- Free siRNA Content: Centrifuge a fresh NP suspension at high speed (e.g., 21,000 x g, 30 min). Filter the supernatant (0.22 µm) and analyze directly by HPLC.
- HPLC Conditions: Flow rate: 1 mL/min. Gradient: 10-25% B over 15 min. Detect at 260 nm.
- Calculation: EE% = [(Total siRNA - Free siRNA) / Total siRNA] x 100.
Protocol 4: DSC Analysis of NP Matrix
Objective: Determine the thermal transitions of PEG-PLGA NPs. Materials: Sealed aluminum DSC pans, lyophilized NP powder (~5 mg), DSC instrument. Procedure:
- Accurately weigh 3-5 mg of lyophilized NPs into a pre-tared aluminum pan. Crimp the lid.
- Use an empty sealed pan as a reference.
- Run a heat-cool-heat cycle: Equilibrate at -20°C, heat to 100°C at 10°C/min, cool to -20°C at 20°C/min, and re-heat to 100°C at 10°C/min under nitrogen purge.
- Analyze the second heating curve. Report the midpoint Tg of PLGA and the melting point (Tm) of crystalline PEG if present.
Visualizations
Characterization Workflow for PEG-PLGA Nanoparticles
Mechanism of Sustained Gene Release from PEG-PLGA NPs
Research Reagent Solutions Toolkit
Table 2: Essential Materials for PEG-PLGA Nanoparticle Characterization
| Item | Function/Application | Example/Notes |
|---|---|---|
| PLGA (50:50, acid-terminated) | Core biodegradable polymer. Molecular weight (7-38 kDa) controls degradation & release rate. | Resomer RG 502H |
| Methoxy-PEG-OH | Forms stealth corona, prolongs circulation, enhances stability. MW: 2-5 kDa. | Sigma-Aldrich 729076 |
| Double-filtered DI Water | Diluent for DLS to avoid particulate interference. Essential for reliable size/zeta data. | 0.1 µm filtered, from a purification system. |
| Uranyl Acetate (2% aq.) | Negative stain for TEM; provides contrast by staining background and NP surface. | Handle as radioactive waste; prepare fresh. |
| C18 Reverse-Phase Column | HPLC separation of nucleic acids (siRNA, pDNA) using ion-pairing chromatography. | Waters XBridge C18, 4.6 x 150 mm, 3.5 µm. |
| Triethylammonium Acetate (TEAA) Buffer | Ion-pairing agent for HPLC, enabling separation of nucleic acids on C18 columns. | 0.1 M, pH 7.0, sterile filtered. |
| Triton X-100 Detergent | Used in siRNA EE% protocol to disrupt NPs and release all encapsulated genetic material. | 1% (v/v) solution in nuclease-free water. |
| Hermetic Aluminum DSC Pans | Ensures no mass loss during DSC heating scans, crucial for accurate thermal analysis. | Tzero pans and lids (TA Instruments). |
This document details protocols for validating PEG-PLGA nanoparticle formulations designed for sustained gene (e.g., plasmid DNA, siRNA) delivery. The primary goal is to establish a robust correlation between in vitro release kinetics and the preserved bioactivity of the released genetic cargo, a critical step in preclinical development.
Core Validation Strategy: Successful sustained-release systems must demonstrate two concurrent outcomes: (1) a prolonged, controlled release profile that fits established kinetic models, and (2) the retention of the genetic material's structural integrity and functional activity post-release from the polymer matrix. This requires complementary analytical and biological assays.
Key Research Reagent Solutions
The following table lists essential materials for conducting these validation studies.
| Research Reagent / Material | Function in Validation |
|---|---|
| PEG-PLGA Copolymer (e.g., 50:50 PLGA-PEG) | The biodegradable, amphiphilic matrix forming the nanoparticle core (PLGA) and stealth corona (PEG). Defines degradation rate and release kinetics. |
| Heparin Sodium Salt | A polyanion used to disrupt ionic interactions and ensure complete displacement of encapsulated nucleic acids from the polymer for release quantification. |
| SYBR Gold Nucleic Acid Gel Stain | Ultrasensitive fluorescent dye for quantifying intact, double-stranded DNA released from nanoparticles, minimizing background. |
| PicoGreen Assay Kit | Alternative high-sensitivity fluorescent assay for dsDNA quantification in release media. |
| RiboGreen Assay Kit | High-sensitivity fluorescent assay for quantifying RNA (e.g., siRNA) in release samples. |
| RNase Inhibitor | Essential for siRNA release studies to prevent degradation of released RNA in media during the assay period. |
| Cell Line with Reporter Gene (e.g., Luciferase, GFP) | Enables functional bioactivity assay by measuring transfection efficiency and duration of gene expression. |
| Lipofectamine 3000 | Positive control transfection reagent for comparing the bioactivity of released genetic material vs. fresh, non-encapsulated material. |
| Dialysis Membranes (Float-A-Lyzer) | With precise molecular weight cut-off (e.g., 100 kDa), used for sample-and-separate release studies under sink conditions. |
Protocols
Protocol 3.1: In Vitro Release Kinetics under Sink Conditions
Objective: To quantify the cumulative release of nucleic acids from PEG-PLGA nanoparticles over time in a physiologically relevant buffer.
Materials: Nanoparticle suspension, Release medium (1x PBS, pH 7.4, ± 0.1% w/v heparin), Dialysis devices (e.g., Float-A-Lyzer, 100 kDa MWCO), Microplate reader, Fluorescent nucleic acid stain (SYBR Gold/PicoGreen/RiboGreen).
Procedure:
- Preparation: Load 1 mL of nanoparticle suspension (containing a known amount of nucleic acid) into a pre-hydrated dialysis device.
- Incubation: Immerse the device in a sink volume of release medium (e.g., 20 mL) at 37°C under gentle agitation (50 rpm). The use of heparin in the external medium ensures sink conditions by dissociating any re-adsorbed nucleic acid.
- Sampling: At predetermined time points (e.g., 1, 4, 8, 24, 48, 72 hours, then weekly), completely replace the external sink medium with fresh, pre-warmed medium.
- Quantification: a. Mix 100 µL of the collected release medium with 100 µL of assay buffer and appropriate dye per manufacturer instructions. b. Measure fluorescence (e.g., PicoGreen: Ex/Em ~480/520 nm). c. Determine nucleic acid concentration from a standard curve of free nucleic acid in the same release medium.
- Data Analysis: Calculate cumulative release percentage. Plot release vs. time and fit data to kinetic models (e.g., Higuchi, Korsmeyer-Peppas, zero-order).
Protocol 3.2: Bioactivity Assay for Released Genetic Material
Objective: To assess the functional integrity and transfection capability of nucleic acids released from nanoparticles over time.
Materials: Release medium samples from Protocol 3.1, Relevant cell line (e.g., HEK293), Complete cell culture medium, Transfection reagent (Lipofectamine 3000 as control), Bioactivity readout system (e.g., luciferase assay kit, flow cytometry for GFP).
Procedure:
- Sample Preparation: Pool and concentrate nucleic acids from specific time-point release samples (e.g., Day 1, Day 7, Day 28) using centrifugal filters if necessary. Adjust to a known concentration in sterile buffer.
- Cell Seeding: Seed cells in a 24-well plate to reach 70-80% confluence at the time of transfection.
- Transfection: For each sample, perform transfection in triplicate using a standard protocol with a commercial reagent (e.g., Lipofectamine 3000). Controls must include:
- Positive Control: Fresh, non-encapsulated nucleic acid.
- Negative Control: Naked nucleic acid incubated in release medium for the longest time point.
- Nanoparticle Control: Cells treated with empty nanoparticles.
- Incubation: Incubate cells for 24-48 hours (for protein expression) or relevant timeframe for functional knockdown (siRNA).
- Bioactivity Measurement:
- For plasmid DNA: Lyse cells and measure reporter protein activity (e.g., luciferase luminescence, GFP fluorescence via flow cytometry).
- For siRNA: Extract total RNA, perform qRT-PCR to measure target gene knockdown relative to a scrambled siRNA control.
- Data Analysis: Normalize bioactivity of released samples to the positive control (fresh nucleic acid), expressed as a percentage of retained bioactivity.
Data Presentation & Analysis
Table 1: Representative In Vitro Release Kinetics Data of pDNA-Loaded PEG-PLGA Nanoparticles
| Time Point (Days) | Cumulative Release (%) ± SD | Best-Fit Kinetic Model | R² Value | Model Parameter (e.g., n from Korsmeyer-Peppas) |
|---|---|---|---|---|
| 1 | 18.5 ± 2.1 | Higuchi | 0.98 | - |
| 3 | 35.2 ± 3.4 | Higuchi | 0.97 | - |
| 7 | 58.7 ± 4.2 | Korsmeyer-Peppas | 0.99 | n = 0.63 |
| 14 | 78.9 ± 5.1 | Korsmeyer-Peppas | 0.98 | n = 0.63 |
| 28 | 95.3 ± 3.8 | Zero-Order | 0.96 | - |
SD: Standard Deviation; n: release exponent indicating mechanism (n>0.85 = Case II transport, 0.45
Table 2: Bioactivity of Released Plasmid DNA Over Time
| Release Sample (Time Point) | Luciferase Activity (RLU/µg protein) ± SD | Bioactivity Retained (% vs. Fresh pDNA) |
|---|---|---|
| Fresh pDNA (Positive Control) | 1.2 x 10⁸ ± 1.5 x 10⁷ | 100% |
| Day 1 Release | 1.1 x 10⁸ ± 9.8 x 10⁶ | 92% |
| Day 7 Release | 9.4 x 10⁷ ± 8.2 x 10⁶ | 78% |
| Day 28 Release | 6.8 x 10⁷ ± 7.1 x 10⁶ | 57% |
| pDNA in Medium, Day 28 (Negative Ctrl) | 5.1 x 10⁵ ± 6.3 x 10⁴ | 0.4% |
RLU: Relative Light Units.
Experimental Visualizations
Title: Workflow for Sustained Release Validation
Title: Mechanism of Sustained Release from PEG-PLGA NPs
Comparative Analysis with Other Non-Viral Vectors (e.g., Liposomes, PEI)
Within the broader thesis on PEG-PLGA nanoparticles for sustained gene release, it is critical to situate this platform's performance against established non-viral vectors. This application note provides a comparative analysis focusing on physicochemical properties, transfection efficiency, cytotoxicity, and in vivo pharmacokinetics. Detailed protocols for head-to-head evaluation are included to enable reproducible research.
Quantitative Comparison of Key Non-Viral Vectors
Table 1: Comparative Properties of Major Non-Viral Gene Delivery Vectors
| Property | PEG-PLGA Nanoparticles | Cationic Liposomes (e.g., Lipofectamine) | Cationic Polymers (e.g., PEI, 25 kDa) | Inorganic Nanoparticles (e.g., Gold NPs) |
|---|---|---|---|---|
| Typical Size Range (nm) | 80-200 | 50-150 | 50-200 (complex) | 10-100 |
| Surface Charge (Zeta Potential, mV) | Slightly negative to +10 | +20 to +60 | +30 to +50 | Variable (-30 to +50) |
| Primary Loading Mechanism | Encapsulation (matrix) | Complexation/ encapsulation | Electrostatic complexation (polyplex) | Surface adsorption/ conjugation |
| Transfection Efficiency (in vitro, %) | 15-40% (cell-dependent) | 70-95% (high in many lines) | 50-80% (cell-dependent) | 5-30% (varies widely) |
| Sustained Release Capability | High (days to weeks) | Low to moderate (burst release) | Moderate (complex dissociation) | Low to moderate |
| Cytotoxicity (MTT assay, Cell Viability %) | >85% (highly biocompatible) | 60-80% (dose-dependent) | 40-70% (highly dose/type-dependent) | 70-95% (surface-dependent) |
| In Vivo Circulation Half-life (t½, h) | 8-24 h (PEGylated) | 1-4 h (rapid clearance) | <1 h (rapid clearance) | 2-12 h (PEGylated) |
| Scalability & GMP Production | Excellent (solvent evaporation) | Good (extrusion) | Excellent (polymer synthesis) | Good (chemical synthesis) |
| Primary Limitation | Moderate transfection efficiency | Toxicity, immunogenicity | High cytotoxicity, instability in blood | Low loading, potential long-term toxicity |
Experimental Protocols for Comparative Analysis
Protocol 3.1: Parallel Formulation & Characterization
Objective: To prepare and characterize PEG-PLGA NPs, liposomes, and PEI polyplexes loaded with a reporter gene (e.g., pDNA encoding GFP) for direct comparison. Materials: See "The Scientist's Toolkit" below. Procedure:
- PEG-PLGA NPs: Use double emulsion solvent evaporation. Dissolve 50 mg PEG-PLGA (50:50, 10% PEG) in 2 mL dichloromethane. Add 200 µL aqueous pDNA solution (0.1 mg/mL) and sonicate (tip, 40 W, 30 s) to form primary W/O emulsion. Pour into 10 mL 1% PVA solution, homogenize (10,000 rpm, 2 min). Stir overnight to evaporate solvent. Centrifuge (21,000 x g, 30 min), wash, lyophilize.
- Cationic Liposomes: Prepare using thin-film hydration. Dissolve DOTAP and DOPE (1:1 molar ratio) in chloroform. Rotary evaporate to form thin film. Hydrate with HEPES buffer (pH 7.4) to 1 mg/mL lipid. Extrude 11x through 100 nm polycarbonate membrane. Complex with pDNA at N/P ratio 5:1 (incubate 20 min RT).
- PEI Polyplexes: Dilute branched PEI (25 kDa) in HEPES buffer. Mix with pDNA solution at N/P ratio 10:1. Vortex, incubate 30 min at RT.
- Characterization: Measure particle size (dynamic light scattering), zeta potential (laser Doppler velocimetry), and DNA encapsulation/loading efficiency (PicoGreen assay for supernatant).
Protocol 3.2: In Vitro Transfection & Cytotoxicity Assay
Objective: To compare transfection efficiency and cytotoxicity in a standardized cell line (e.g., HEK-293). Procedure:
- Seed cells in 24-well plate at 5x10⁴ cells/well. Incubate 24 h (37°C, 5% CO₂).
- Prepare vectors containing 0.5 µg pDNA-GFP per well in serum-free medium.
- Replace medium with vector complexes. After 4 h, replace with complete medium.
- Transfection Analysis: At 48 h, trypsinize cells, resuspend in PBS, and analyze GFP-positive cells via flow cytometry. Report as % transfection.
- Cytotoxicity (Parallel Plate): Using same conditions (step 3), incubate for 24 h. Add MTT reagent (0.5 mg/mL) for 4 h. Solubilize formazan crystals with DMSO. Measure absorbance at 570 nm. Calculate viability relative to untreated cells.
Protocol 3.3: In Vivo Pharmacokinetics and Sustained Release
Objective: To compare plasma circulation and tissue retention of DNA delivered by different vectors. Procedure:
- Label pDNA with Cy5 fluorophore using Label IT kit.
- Formulate Cy5-pDNA into PEG-PLGA NPs, liposomes, and PEI polyplexes.
- Intravenously inject mice (n=5 per group) with equivalent DNA dose (20 µg).
- Collect blood samples at 5 min, 30 min, 2 h, 8 h, 24 h, 48 h.
- Isolate plasma, measure Cy5 fluorescence. Calculate pharmacokinetic parameters (t½, AUC).
- At 48 h, sacrifice, image major organs ex vivo via fluorescence imager. Quantify signal in liver, spleen, kidneys, and lungs.
Visualization of Experimental Workflows & Mechanisms
Title: Workflow for Comparative Vector Analysis
Title: Vector Uptake and Release Mechanisms
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions
| Item | Function in Comparative Analysis | Example Product/Catalog # |
|---|---|---|
| PEG-PLGA Copolymer | Forms core nanoparticle matrix for sustained release. Provides "stealth" properties. | RESOMER RGP d 50155 (Merck) |
| Cationic Lipid (DOTAP) | Key component of liposomal vectors for DNA complexation via charge interaction. | Avanti Polar Lipids, #890890 |
| Branched PEI (25 kDa) | Gold-standard cationic polymer for polyplex formation; high transfection benchmark. | Sigma-Aldrich, #408727 |
| PicoGreen dsDNA Assay Kit | Quantifies encapsulated vs. free DNA for loading efficiency calculations. | Thermo Fisher, #P11496 |
| Label IT Nucleic Acid Labeling Kit (Cy5) | Fluorescently labels pDNA for tracking in vitro and in vivo. | Mirus Bio, #MIR 3700 |
| In Vivo-JetPEI | Linear PEI formulation optimized for in vivo studies; a relevant comparator. | Polyplus, #201-50G |
| GMP-grade pDNA (e.g., gWiz GFP) | Standardized, high-quality reporter gene for consistent transfection assays. | Aldevron, #gWiz-GFP |
| Polyvinyl Alcohol (PVA) | Stabilizer/emulsifier in PEG-PLGA NP formulation. | Sigma-Aldrich, #363138 |
| Size Exclusion Columns (e.g., Sephadex G-50) | Purifies DNA complexes from unbound components. | Cytiva, #45-001-527 |
| MTT Cell Viability Assay Kit | Standardized colorimetric assay for cytotoxicity comparison. | Abcam, #ab211091 |
Within the thesis research on developing PEG-PLGA nanoparticles for sustained gene release, rigorous evaluation of safety parameters is paramount for translational success. This document provides detailed application notes and standardized protocols for assessing cytotoxicity and immunogenicity—two critical pillars of the biocompatibility profile. These assessments are essential for de-risking novel nanocarrier formulations prior to in vivo studies and clinical translation.
Cytotoxicity Assessments: Protocols and Data
Cytotoxicity evaluation determines the potential adverse effects of PEG-PLGA nanoparticles on cell viability and function. Standardized in vitro tests are the first line of screening.
Table 1: Summary of Standard In Vitro Cytotoxicity Assays
| Assay Name | Measured Endpoint | Key Reagents/ Kits | Typical Incubation Time with Nanoparticles | Data Output | Interpretation Threshold (Typical) |
|---|---|---|---|---|---|
| MTT Assay | Metabolic activity (mitochondrial reductase function) | MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide), DMSO | 24 - 72 hours | Absorbance at 570 nm | Cell viability < 70-80% of control indicates potential cytotoxicity. |
| CCK-8 Assay | Metabolic activity (dehydrogenase activity) | CCK-8 (Cell Counting Kit-8) solution | 1 - 4 hours after 24-72h incubation | Absorbance at 450 nm | More sensitive than MTT. Same threshold applies. |
| LDH Release Assay | Membrane integrity (cytotoxicity) | LDH assay kit (measures lactate dehydrogenase leakage) | 24 - 48 hours | Absorbance at 490 nm | Increased LDH in supernatant correlates with necrotic cell death. |
| Live/Dead Staining | Direct viability count (membrane integrity) | Calcein-AM (live, green), Propidium Iodide or EthD-1 (dead, red) | 30-45 min staining after incubation | Fluorescence microscopy/ quantification | Provides visual confirmation and ratio of live/dead cells. |
| Annexin V/PI Flow Cytometry | Apoptosis vs. Necrosis | Annexin V-FITC, Propidium Iodide (PI) | After 24-48h incubation | Flow cytometry scatter plots (Annexin V vs. PI) | Quantifies early/late apoptotic and necrotic populations. |
Detailed Protocol: MTT Assay for PEG-PLGA Nanoparticles
Objective: To assess the metabolic activity of cells (e.g., HEK293, HepG2, or primary cells relevant to the target tissue) after exposure to PEG-PLGA gene-loaded nanoparticles.
Materials:
- Cell line of choice
- Complete cell culture medium
- PEG-PLGA nanoparticle suspensions (in sterile PBS or culture medium) at various concentrations (e.g., 0.1, 0.5, 1.0 mg/mL).
- MTT reagent (5 mg/mL in PBS, sterile-filtered).
- Dimethyl sulfoxide (DMSO).
- 96-well tissue culture plate.
- Multi-channel pipette.
- Microplate reader.
Procedure:
- Cell Seeding: Seed cells in a 96-well plate at an optimal density (e.g., 5,000-10,000 cells/well) in 100 µL of complete medium. Incubate for 24 hours to allow cell adherence.
- Nanoparticle Treatment: Prepare serial dilutions of the nanoparticle suspension in culture medium. Aspirate the medium from the plate and add 100 µL of each nanoparticle concentration to respective wells. Include a negative control (medium only) and a positive control (e.g., 1% Triton X-100). Perform replicates (n=4-6).
- Incubation: Incubate the plate for the desired period (24h, 48h, 72h) at 37°C, 5% CO₂.
- MTT Addition: After incubation, carefully add 10 µL of MTT solution (5 mg/mL) to each well. Return plate to incubator for 2-4 hours.
- Solubilization: Gently remove the medium containing MTT. Add 100 µL of DMSO to each well to solubilize the formed purple formazan crystals.
- Measurement: Agitate the plate gently for 5 minutes. Measure the absorbance of each well at 570 nm using a microplate reader, with a reference wavelength of 630-650 nm.
- Data Analysis: Calculate cell viability as a percentage: (Mean Absorbance of Test Well / Mean Absorbance of Control Well) * 100%. Plot viability vs. nanoparticle concentration.
Cytotoxicity Assessment Workflow
Immunogenicity Assessments: Protocols and Data
Immunogenicity assessment evaluates the potential of PEG-PLGA nanoparticles to trigger unwanted immune responses, which can lead to rapid clearance, reduced efficacy, and adverse effects.
Table 2: Summary of Key Immunogenicity Assays for Nanoparticles
| Assay Name | Immune Component Assessed | Key Reagents/ Cells | Sample Type (In Vitro/Ex Vivo) | Readout |
|---|---|---|---|---|
| Cytokine Profiling | Innate & adaptive immune activation (e.g., TNF-α, IL-6, IL-1β, IFN-γ) | ELISA or Multiplex Luminex kits; PBMCs or macrophage cell lines (RAW 264.7, THP-1) | Cell culture supernatant | Cytokine concentration (pg/mL) |
| Complement Activation | Activation of complement cascade (innate immunity) | C3a, C5a, or SC5b-9 ELISA kits | Human/animal serum incubated with NPs | Complement split product levels |
| Hemolysis Assay | Erythrocyte membrane damage (non-specific immune trigger) | Human/animal RBCs, PBS, Triton X-100 | RBC suspension + NPs | Absorbance at 540 nm (% Hemolysis) |
| Differential Scanning Calorimetry (DSC) | Protein corona-induced conformational changes (immunogenicity predictor) | NP-protein corona complex | Ex vivo serum-incubated NPs | Thermogram shift (ΔTm) |
| Anti-PEG Antibody Detection | Humoral response to PEG component | PEGylated antigen, serum from treated animals | Animal serum | ELISA titer (anti-PEG IgM/IgG) |
Detailed Protocol: Cytokine Release Assay Using Human PBMCs
Objective: To evaluate the pro-inflammatory potential of PEG-PLGA nanoparticles by measuring cytokine secretion from primary human immune cells.
Materials:
- Fresh or cryopreserved Human Peripheral Blood Mononuclear Cells (PBMCs).
- RPMI-1640 medium supplemented with 10% FBS and 1% Pen/Strep.
- PEG-PLGA nanoparticle suspensions (sterile, in PBS).
- Positive control: Lipopolysaccharide (LPS, 1 µg/mL).
- Negative control: PBS or "empty" PEG-PLGA nanoparticles.
- 96-well U-bottom plate.
- Human cytokine ELISA kits (e.g., for TNF-α and IL-6).
- Centrifuge, CO₂ incubator, microplate reader.
Procedure:
- PBMC Preparation: Isolate PBMCs from buffy coats using density gradient centrifugation (Ficoll-Paque) or thaw cryopreserved vials. Resuspend cells in complete RPMI at 1x10⁶ cells/mL.
- Cell Stimulation: Aliquot 100 µL of cell suspension (1x10⁵ cells) into wells of a 96-well U-bottom plate. Add 100 µL of nanoparticle suspension (at 2x final concentration in medium) to triplicate wells. Include LPS-positive control and PBS/empty NP-negative controls. Final nanoparticle concentrations should span the expected therapeutic range.
- Incubation: Incubate plate for 24 hours (for early cytokines like TNF-α, IL-6) at 37°C, 5% CO₂.
- Supernatant Collection: Centrifuge plate at 300 x g for 5 minutes. Carefully collect 150 µL of supernatant from each well without disturbing the cell pellet. Store at -80°C if not used immediately.
- Cytokine ELISA: Perform ELISA according to the manufacturer's instructions. Typically involves coating plates with capture antibody, blocking, adding samples and standards, detection antibody, enzyme conjugate, and substrate.
- Data Analysis: Calculate cytokine concentrations from the standard curve. Express data as mean ± SEM (pg/mL). A statistically significant increase in cytokine levels compared to the negative control indicates immunostimulatory potential.
Immunogenicity Assessment Workflow & Key Pathways
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Safety Assessment of PEG-PLGA Nanoparticles
| Item Name | Supplier Examples (Current) | Function in Assessment | Critical Notes for PEG-PLGA NPs |
|---|---|---|---|
| Cell Counting Kit-8 (CCK-8) | Dojindo, Sigma-Aldrich, Abcam | One-step, highly sensitive measurement of cell metabolic activity (dehydrogenases). | Preferred over MTT for high-throughput; more soluble formazan. Check for NP interference at 450 nm. |
| LDH Cytotoxicity Assay Kit | Promega, Thermo Fisher (Pierce), Cayman Chemical | Quantifies lactate dehydrogenase released upon cell membrane damage (necrosis). | Use serum-free medium during NP exposure to avoid background LDH. |
| Annexin V-FITC / PI Apoptosis Kit | BioLegend, BD Biosciences, Invitrogen | Distinguishes early apoptotic (Annexin V+/PI-), late apoptotic/necrotic (Annexin V+/PI+) cells via flow cytometry. | Essential for gene delivery NPs, as cargo (e.g., pDNA, siRNA) can induce apoptosis. |
| Human/Mouse Cytokine ELISA Kits | R&D Systems, BioLegend, Invitrogen | Quantifies specific cytokine proteins (e.g., TNF-α, IL-6, IL-1β) in cell supernatants or serum. | Use multiplex panels (Luminex) for broader screening from limited in vivo samples. |
| Human Complement C3a ELISA Kit | Quidel, MyBioSource, Abcam | Measures C3a levels as a marker of complement activation via the alternative pathway. | Incubate NPs in 10-50% human serum for 1h at 37°C before measurement. |
| Calcein-AM / Ethidium Homodimer-1 (EthD-1) | Invitrogen (Live/Dead Kit), Sigma | Fluorescent live/dead cell double stain for microscopy. Calcein-AM (green, live), EthD-1 (red, dead). | Direct visualization of NP toxicity; confirm results from plate-based assays. |
| THP-1 Human Monocyte Cell Line | ATCC, Sigma | Differentiable to macrophage-like cells, a standard model for innate immune response (cytokine release). | Differentiate with PMA (phorbol ester) before NP treatment for macrophage phenotype. |
| Purified Human Serum | Commercial vendors or local blood bank | Provides native complement proteins and opsonic proteins for protein corona and immunogenicity studies. | Use freshly prepared or properly stored (-80°C) serum to maintain complement activity. |
1. Introduction Within the thesis framework of developing PEG-PLGA nanoparticles (NPs) for sustained gene release, preclinical studies across diverse therapeutic areas validate the platform's versatility. This document details key application notes and protocols for cancer therapy, vaccinology, and regenerative medicine, emphasizing quantitative outcomes and reproducible methods.
2. Case Study 1: Cancer Therapy – Sustained siRNA Delivery for Oncogene Silencing
2.1 Application Note PEG-PLGA NPs loaded with siRNA targeting the KRASG12D oncogene were evaluated in a pancreatic ductal adenocarcinoma (PDAC) xenograft model (MIA PaCa-2 cells). The formulation enabled sustained siRNA release over 14 days in vitro.
Table 1: Efficacy Data of KRAS siRNA-Loaded PEG-PLGA NPs in a PDAC Xenograft Model
| Parameter | PBS Control | Naked siRNA | PEG-PLGA/siRNA NPs |
|---|---|---|---|
| Tumor Volume (Day 21) | 1250 ± 210 mm³ | 1180 ± 195 mm³ | 520 ± 115 mm³ |
| KRAS mRNA Expression (Relative) | 1.00 ± 0.15 | 0.92 ± 0.18 | 0.32 ± 0.09 |
| Apoptotic Index (TUNEL+ %) | 4.5 ± 1.2% | 5.1 ± 1.5% | 28.7 ± 4.8% |
| Peak Serum Cytokine (IL-6) | <10 pg/mL | 12 pg/mL | <10 pg/mL |
p<0.01 vs. PBS Control
2.2 Protocol: Preparation and In Vivo Evaluation of siRNA-Loaded PEG-PLGA NPs
A. Nanoparticle Formulation via Double Emulsion (W/O/W)
- Inner Aqueous Phase: Dissolve 100 µg KRASG12D siRNA in 0.5 mL of nuclease-free water.
- Organic Phase: Dissolve 100 mg of PEG(5k)-PLGA(50k) (50:50 LA:GA) in 2 mL of dichloromethane (DCM).
- Primary Emulsion: Add the aqueous phase to the organic phase. Emulsify using a probe sonicator (70% amplitude, 30 s on ice) to form a W/O emulsion.
- Secondary Emulsion: Pour the primary emulsion into 8 mL of 2% (w/v) polyvinyl alcohol (PVA) solution. Sonicate again (50% amplitude, 60 s) to form a W/O/W emulsion.
- Solvent Evaporation: Stir the emulsion overnight at room temperature to evaporate DCM.
- Purification: Centrifuge at 18,000 x g for 30 min. Wash pellet with water twice and resuspend in sterile PBS. Filter through a 0.8/0.2 µm syringe filter.
- Characterization: Measure particle size (Zetasizer), encapsulation efficiency (RiboGreen assay), and in vitro release profile (PBS, 37°C).
B. In Vivo Efficacy Study in PDAC Xenografts
- Establish subcutaneous MIA PaCa-2 tumors (~100 mm³) in NOD/SCID mice.
- Randomize animals into groups (n=8): PBS, Naked siRNA, PEG-PLGA/siRNA NPs.
- Administer 2 mg/kg siRNA dose via intratumoral injection every 7 days for 3 weeks.
- Measure tumor dimensions bi-weekly with calipers. Calculate volume: V = (L x W²)/2.
- On day 22, euthanize, excise tumors, and process for qRT-PCR (KRAS mRNA) and TUNEL immunohistochemistry.
2.3 Diagram: Mechanism of Action for siRNA NPs in Cancer Cells
Title: siRNA NP Mechanism for Oncogene Silencing
3. Case Study 2: Vaccinology – Sustained Antigen & Adjuvant Co-Delivery for Cellular Immunity
3.1 Application Note PEG-PLGA NPs co-encapsulating ovalbumin (OVA) model antigen and the TLR7/8 agonist R848 induced potent, sustained cytotoxic T lymphocyte (CTL) responses. In a prophylactic murine melanoma (B16-OVA) challenge model, NPs elicited complete protection in 80% of animals.
Table 2: Immunogenicity of OVA/R848 PEG-PLGA NP Vaccine
| Parameter | Soluble OVA+R848 | PEG-PLGA/OVA+R848 NPs |
|---|---|---|
| OVA-Specific CD8+ T Cells (Day 7, % of CD8+) | 2.3 ± 0.5% | 12.8 ± 2.1% |
| IFN-γ Secretion (Spot/10⁶ Splenocytes) | 45 ± 15 | 320 ± 55 |
| Serum Anti-OVA IgG Titer (Day 14) | 1:2,500 | 1:51,200 |
| Tumor-Free Survival (Day 60) | 0% | 80% |
| Antigen+DC in DLN (Day 3, Fold Increase) | 1x | 8.5x |
p<0.01 vs. Soluble group
3.2 Protocol: Preparation of Co-Encapsulated Vaccine NPs and Immunogenicity Assessment
A. Antigen/Adjuvant NP Formulation
- Inner Aqueous Phase: Dissolve 1 mg OVA protein and 0.1 mg R848 in 0.5 mL of 10 mM sodium acetate buffer (pH 5.0).
- Follow steps 2-7 from Protocol 2.2.A, using the same polymer and method.
- Characterization: Measure size, PDI, zeta potential. Determine OVA encapsulation (BCA assay) and R848 encapsulation (HPLC).
B. In Vivo Immunization and Challenge
- Immunize C57BL/6 mice (n=10/group) subcutaneously with 50 µg OVA and 5 µg R848 equivalents on days 0 and 14.
- On day 21, isolate splenocytes for ELISpot (IFN-γ) and flow cytometry (tetramer staining for OVA257-264).
- For challenge, immunize on days 0 and 14. On day 28, inject 5x10⁵ B16-OVA cells subcutaneously. Monitor tumor growth and survival for 60 days.
3.3 Diagram: Immune Activation Pathway by Vaccine NP
Title: Vaccine NP Induces Cellular Immunity
4. Case Study 3: Regenerative Medicine – Sustained miRNA Delivery for Bone Regeneration
4.1 Application Note PEG-PLGA NPs delivering pro-osteogenic miRNA-26a were applied in a critical-size calvarial defect model in rats. Sustained release over 21 days enhanced bone marrow stromal cell (BMSC) osteogenesis in vitro and led to ~85% bone defect coverage in vivo at 8 weeks.
Table 3: Bone Regeneration with miRNA-26a Loaded PEG-PLGA NPs
| Parameter | Empty NP | PEG-PLGA/miRNA-26a NPs |
|---|---|---|
| In Vitro ALP Activity (Day 7, Fold Change) | 1.0 ± 0.2 | 3.5 ± 0.4 |
| In Vitro Calcium Deposition (Day 21, Alizarin Red, Fold) | 1.0 ± 0.3 | 4.2 ± 0.6 |
| In Vivo New Bone Volume (BV/TV, % at 8 wks) | 22.5 ± 5.5% | 84.7 ± 6.3% |
| In Vivo Bone Mineral Density (mg HA/cm³) | 350 ± 45 | 680 ± 60 |
| Defect Closure (µCT, %) | 25% | 85% |
p<0.01 vs. Empty NP
4.2 Protocol: miRNA-Loaded NP Formulation and In Vivo Bone Defect Study
A. miRNA NP Formulation and In Vitro Testing
- Formulate NPs using Protocol 2.2.A, replacing siRNA with miRNA-26a mimic.
- Transfection: Seed human BMSCs (hBMSCs) in 24-well plates. Treat with NPs containing 50 nM miRNA-26a. Replace medium after 24h.
- Assays: Measure alkaline phosphatase (ALP) activity at day 7 (pNPP assay) and mineralized matrix at day 21 (Alizarin Red S staining, quantitative elution with cetylpyridinium chloride).
B. Calvarial Defect Model in Rats
- Create a 5mm critical-size calvarial defect in Sprague-Dawley rats.
- Implant a collagen sponge soaked with either PBS, Empty NPs, or miRNA-26a NPs (2 µg miRNA dose) into the defect.
- At 4 and 8 weeks post-op, euthanize animals and analyze defects via micro-CT for bone volume/total volume (BV/TV). Process for histology (H&E, Masson's Trichrome).
4.3 Diagram: miRNA-26a Pro-Osteogenic Signaling Pathway
Title: miRNA-26a NP Drives Osteogenesis
5. The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for PEG-PLGA Nanoparticle Research
| Reagent/Material | Supplier Examples | Function in Research |
|---|---|---|
| PEG-PLGA Copolymers (e.g., PEG5k-PLGA50k) | Sigma-Aldrich, Lactel Absorbable Polymers, PolySciTech | Core biodegradable polymer providing sustained release and stealth properties. |
| Therapeutic Nucleic Acids (siRNA, miRNA mimic) | Horizon Discovery, Qiagen, Sigma-Aldrich | Active pharmaceutical ingredient for gene silencing (siRNA) or regulation (miRNA). |
| Model Antigen & Adjuvant (OVA, R848) | InvivoGen, Sigma-Aldrich | Key components for vaccine studies to evaluate humoral and cellular immune responses. |
| Polyvinyl Alcohol (PVA) (MW 31-50k, 87-89% hydrolyzed) | Sigma-Aldrich, Merck | Critical emulsifier and stabilizer during NP formulation via double emulsion. |
| RiboGreen/Quant-iT Assay Kit | Thermo Fisher Scientific | Highly sensitive, specific quantification of encapsulated nucleic acid. |
| Dichloromethane (DCM), HPLC Grade | Sigma-Aldrich, Fisher Scientific | Organic solvent for dissolving PLGA polymer during emulsion preparation. |
| Zetasizer Nano System | Malvern Panalytical | Key instrument for measuring nanoparticle hydrodynamic diameter (DLS), PDI, and zeta potential. |
| Cell Lines (e.g., MIA PaCa-2, B16-OVA, hBMSCs) | ATCC, MilliporeSigma | Essential for in vitro efficacy, cytotoxicity, and mechanism studies. |
Conclusion
PEG-PLGA nanoparticles represent a versatile and highly tunable platform for sustained gene delivery, offering a favorable balance between efficacy, safety, and manufacturability. Mastery of the foundational principles, coupled with robust methodological protocols and systematic troubleshooting, is essential for developing successful formulations. While challenges in transfection efficiency and scale-up persist, ongoing innovations in surface engineering, material science, and process control continue to advance the field. Future directions will likely focus on smart, stimuli-responsive systems, combinatorial therapies, and navigating the regulatory pathway toward clinical translation, solidifying the role of these nanoparticles in the next generation of genetic medicines.