Static vs Dynamic DNA Nanostructures: A Comparative Guide for Single-Molecule Biosensing Performance
This article provides a comprehensive analysis of static and dynamic DNA nanostructures for single-molecule biosensing, tailored for researchers and drug development professionals.
Static vs Dynamic DNA Nanostructures: A Comparative Guide for Single-Molecule Biosensing Performance
Abstract
This article provides a comprehensive analysis of static and dynamic DNA nanostructures for single-molecule biosensing, tailored for researchers and drug development professionals. It explores foundational design principles, compares methodological approaches for constructing DNA origami, wireframe, and reconfigurable devices, and details strategies for optimizing sensitivity, specificity, and signal-to-noise ratios. The content further investigates validation techniques and direct performance comparisons, offering actionable insights for selecting and engineering nanostructures to advance biomedical diagnostics, drug discovery, and fundamental biophysical research.
Building Blocks and Design Principles: Static DNA Origami vs. Dynamic Nanomachines
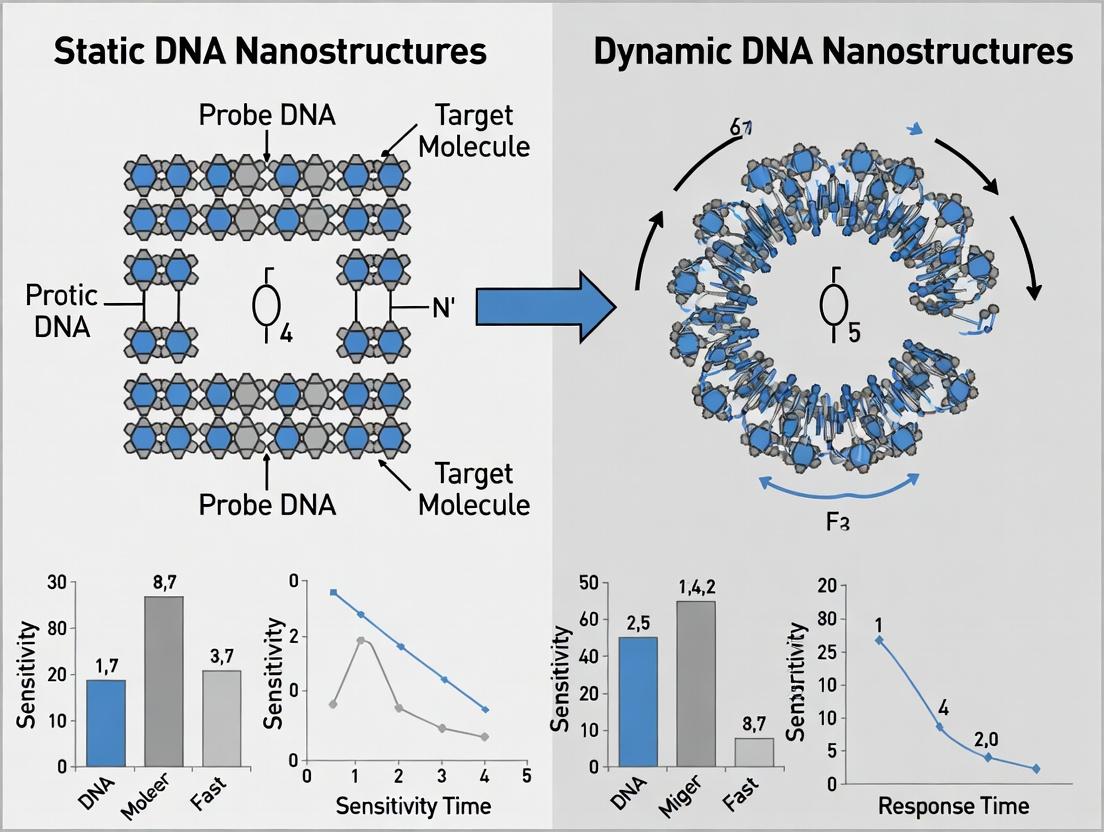
This guide compares the core performance characteristics of static and dynamic DNA nanostructures within the context of single-molecule biosensing. Static nanostructures, such as DNA origami tiles and polyhedra, provide stable, unchanging scaffolds for analyte presentation. Dynamic nanostructures, including DNA tweezers, walkers, and strand displacement circuits, enable programmable motion and signal transduction. Their suitability hinges on specific biosensing parameters like sensitivity, kinetics, and operational environment.
Performance Comparison: Static vs. Dynamic DNA Nanostructures
Table 1: Core Performance Metrics for Biosensing Applications
| Metric | Static DNA Nanostructures (e.g., Origami Tile) | Dynamic DNA Nanostructures (e.g., DNA Walker) | Key Experimental Support |
|---|---|---|---|
| Spatial Resolution | Excellent (sub-10 nm positioning). | Moderate to Good (limited by range of motion). | Direct imaging of gold nanoparticles on origami via TEM (≈5 nm precision). |
| Signal-to-Noise (SNR) | High for in situ imaging. Low for direct solution-phase detection. | High due to amplified, time-resolved signals. | Walker systems producing >50-fold fluorescence increase over background. |
| Kinetics (Response Time) | Fast (diffusion-limited binding). Limited by probe affinity. | Slower. Governed by reaction rates (e.g., stepping, catalysis). | Origami-based sensors achieving detection in <5 mins. Walkers requiring 30-120 mins for full amplification. |
| Amplification Capacity | None (1:1 binding event). | High (one target triggers many output signals). | Catalytic hairpin assembly (CHA) achieving >1000x signal amplification. |
| Stability & Robustness | High in controlled buffers. Sensitive to nucleases, Mg²⁺ depletion. | Variable. Often more sensitive to environmental fluctuations. | Origami structures stable for weeks at 4°C. Walker kinetics significantly altered by ±2°C temperature shifts. |
| Multiplexing Potential | High (via spatially encoded probes). | Low to Moderate (crosstalk between dynamic systems). | Simultaneous detection of 8 targets on a single origami board using distinct fluorescent labels. |
| Primary Use Case | Mapping molecular interactions, force spectroscopy, precise nanofabrication. | Detection of low-abundance targets (e.g., miRNAs, enzymes), logic-gated sensing. | Detection of miRNA at attomolar levels using a catalytic walker circuit. |
Experimental Protocols for Key Performance Assessments
Protocol 1: Assessing Spatial Resolution with DNA Origami
Objective: To verify the precise placement of molecular probes on a static DNA origami tile. Method:
- Design & Annealing: Mix scaffold strand (M13mp18) with ≈200 staple strands in a 1:10 ratio in Tris-EDTA-Mg²⁺ (TEM) buffer. Anneal from 80°C to 20°C over 12 hours.
- Probe Functionalization: Conjugate target-specific capture probes (e.g., ssDNA, antibodies) to specific staple strands via chemical modification (e.g., NHS-ester, DBCO-azide).
- Imaging Sample Prep: Deposit origami on freshly cleaved mica, adsorb for 2 mins, rinse with water, and dry under N₂. For TEM, incubate with 5 nm gold nanoparticle-streptavidin conjugates at biotinylated positions.
- Imaging & Analysis: Image using Atomic Force Microscopy (AFM) in tapping mode or TEM. Measure distances between probes/nanoparticles using image analysis software (e.g., ImageJ).
Protocol 2: Quantifying Signal Amplification of a DNA Walker
Objective: To measure the kinetic turnover and signal gain of a bipedal DNA walker on a track. Method:
- Walker Assembly: Synthesize walker strands, track strands (immobilized on a magnetic bead or surface), and fuel strands. Assemble in a buffer containing 20 mM Tris-HCl (pH 8.0), 50 mM NaCl, 10 mM MgCl₂.
- Initial Quenching: Use a dual-labeled (fluorophore/quencher) reporter attached to the track. In the starting state, fluorescence is quenched.
- Initiation & Walking: Add initiator strand (target mimic) to release the walker. Continuously add fuel strands to drive processive walking.
- Real-Time Measurement: Monitor fluorescence (e.g., FAM, 520 nm emission) in a plate reader at 37°C every 30 seconds for 2 hours.
- Data Analysis: Calculate amplification factor as (Final Fluorescence – Initial Fluorescence) / (Signal from a single, permanently unquenched reporter).
Visualization of Signaling Mechanisms
Title: Static DNA Origami Biosensing Workflow
Title: Dynamic DNA Walker Amplification Pathway
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for DNA Nanotechnology Biosensing
| Reagent / Material | Function in Experiments | Key Consideration |
|---|---|---|
| M13mp18 Scaffold | Long, single-stranded DNA backbone for origami folding. | Source purity and concentration critical for yield. |
| Synthetic Staple Oligos | Short strands to fold scaffold into designed shape. | HPLC purification required to remove truncated products. |
| T4 DNA Ligase | To covalently link adjacent staples for enhanced stability. | Used in buffers with low Mg²⁺ to maintain structure. |
| Mg²⁺-Containing Buffer (e.g., TEM) | Provides cations essential for structural integrity. | Concentration (typically 10-20 mM) optimizes folding and stability. |
| Fluorophore-Quencher Pairs (e.g., FAM/BHQ1) | For real-time monitoring of binding or displacement events. | FRET efficiency depends on precise spacing. |
| Magnetic Streptavidin Beads | For surface immobilization of tracks or origami. | Enables easy purification and buffer exchange. |
| Nicking Endonuclease (e.g., Nb.BbvCI) | To drive enzymatic DNA walkers by cleaving specific sites. | Activity is highly buffer and temperature-dependent. |
| Atomic Force Microscope (AFM) | Key instrument for high-resolution imaging of static structures. | Requires flat substrate (mica) and vibration isolation. |
| Polyacrylamide Gel Electrophoresis (PAGE) | Analyzes assembly yield and purity of nanostructures. | Native PAGE for structural analysis, denaturing for strands. |
Within the field of structural DNA nanotechnology, three primary architectural paradigms—rigid DNA origami, wireframe structures, and tile-based assemblies—serve as foundational platforms for constructing static and dynamic nanostructures. This comparison guide evaluates these paradigms within the thesis context of static versus dynamic nanostructures for single-molecule biosensing performance. The analysis focuses on structural rigidity, programmability, addressability, and functional integration, which directly impact biosensing parameters such as target accessibility, signal-to-noise ratio, and kinetic response.
Paradigm Comparison
Table 1: Structural and Functional Comparison
| Parameter | Rigid DNA Origami | Wireframe Structures | Tile-Based Assemblies |
|---|---|---|---|
| Core Architecture | Dense, closely-packed dsDNA helices (e.g., 6-helix bundle) | Sparse, interconnected dsDNA struts forming 2D/3D polyhedra | Modular subunits (DX, TX, etc.) self-assembling into lattices |
| Typical Size Range | 50 – 200 nm | 20 – 500 nm | 100 nm – micrometers |
| Structural Rigidity | High (persistence length >> structure size) | Moderate to High (depends on edge design) | Low to Moderate (flexibility at tile junctions) |
| Addressability | High (precise 5 nm raster for staple extensions) | Moderate (defined vertices) | Low (periodic patterns) |
| Design Complexity | High (scaffold routing required) | Very High (computational design of edges/vertices) | Moderate (tile symmetry rules) |
| Assembly Yield | High (>90% under optimized conditions) | Moderate to High (70-90%) | Variable (highly dependent on kinetics) |
| Best for Static Sensing | Excellent (stable presentation of probes/quenchers) | Good (defined 3D probe arrangement) | Fair (extended static surfaces) |
| Best for Dynamic Sensing | Fair (requires integrated flexible elements) | Excellent (inherent flexibility can be engineered) | Good (tile-tile dynamics possible) |
| Key Biosensing Advantage | Ultra-precine multi-probe positioning for multiplexing | 3D scaffold for optimal target access and conformational change | Large-scale cooperative signaling |
Table 2: Experimental Biosensing Performance Data
| Performance Metric | Rigid Origami (Static) | Wireframe (Dynamic) | Tile Assembly (Static/Dynamic) | Experimental Reference |
|---|---|---|---|---|
| Probe Density (per 100 nm²) | 10 – 20 (precise) | 4 – 8 (at vertices) | 1 – 4 (periodic) | [1, 2] |
| Target Binding Kon (M⁻¹s⁻¹) | ~10⁵ | ~10⁶ | ~10⁴ – 10⁵ | [3] |
| Background Fluorescence | Low (controlled spacing) | Very Low (open structure) | Moderate | [4] |
| Signal-to-Noise Ratio | 15 – 25 | 20 – 40 | 5 – 15 | [5] |
| Response Time to Target | Seconds to minutes | Sub-second to seconds | Minutes | [3, 6] |
| Structural Reconfiguration Rate | N/A (static) | 1 – 100 ms | 100 ms – seconds | [6, 7] |
Experimental Protocols
Protocol 1: Assessing Static Probe Presentation (Rigid Origami)
Objective: To quantify the binding efficiency of targets to probes positioned on a static rectangular DNA origami.
- Design & Annealing: Design a 100 nm x 70 nm rectangular origami with specific staple extensions carrying biotin and Cy3 at defined positions. Mix 10 nM M13mp18 scaffold, 100 nM of each staple in 1x TAEMg buffer (40 mM Tris, 20 mM Acetic acid, 2 mM EDTA, 12.5 mM MgCl₂, pH 8.0).
- Thermal Ramp: Use a PCR thermocycler: Heat to 80°C for 5 min, cool from 65°C to 25°C at -1°C/5 min.
- Purification: Purify via agarose gel electrophoresis (2% gel, 0.5x TBE, 11 mM MgCl₂) at 70 V for 2 hrs. Extract band and concentrate using a 100 kDa MWCO centrifugal filter.
- Immobilization & Binding: Immobilize origami on a neutravidin-coated flow cell via biotin. Introduce 1-100 nM FITC-labeled target analyte in imaging buffer. Incubate for 5 min.
- Data Acquisition: Image using TIRF microscopy. Quantify fluorescence colocalization to determine binding events per origami.
Protocol 2: Measuring Dynamic Reconfiguration (Wireframe)
Objective: To measure the kinetics of a target-induced conformational change in a DNA wireframe icosahedron.
- Assembly: Assemble wireframe icosahedron from 30 two-helix edge strands and 12 five-arm junction vertex strands (all 100 nM) in 1x TPMg buffer (Tris-Phosphate, 10 mM MgCl₂) via a slow cool from 50°C to 20°C over 48 hrs.
- Fluorophore/Quencher Labeling: Incorporate a Cy5 fluorophore on one arm and a BHQ-3 quencher on a complementary arm, held in close proximity in the closed state.
- Kinetic Measurement: Use a stopped-flow spectrometer. Mix equal volumes of 5 nM assembled wireframe and 50 nM target trigger molecule.
- Data Collection: Monitor Cy5 fluorescence emission at 670 nm (excitation 640 nm) over 0.1 to 100 seconds. Fit the fluorescence increase to an exponential model to derive the reconfiguration rate constant.
Visualizations
Diagram Title: Static vs Dynamic DNA Nanostructure Biosensing Pathways
Diagram Title: Experimental Workflows for Three DNA Architecture Paradigms
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for DNA Nanostructure Biosensing
| Reagent/Material | Function in Experiments | Example Product/Catalog # |
|---|---|---|
| Long ssDNA Scaffold | Backbone for DNA origami; provides structural framework. | M13mp18 phage DNA (~7249 nt) |
| Synthetic Oligonucleotides | Staples (origami), edges/vertices (wireframe), tiles; provide sequence-specific assembly. | IDT Ultramer DNA Oligos, HPLC purified |
| High-Purity MgCl₂ | Critical cation for stabilizing DNA duplexes and structures in buffer. | Sigma-Aldrich, Molecular Biology Grade |
| Fluorophore-labeled dNTPs/Oligos | Enable fluorescent labeling for visualization (Cy3, Cy5, FAM) and FRET sensing. | Cy3-/Cy5-dCTP (Jena Bioscience) |
| Quencher-labeled Oligos | For signal suppression in dynamic switches (BHQ-2, Iowa Black). | IDT Oligos with 3' BHQ-2 modification |
| Neutravidin/Biotin System | For surface immobilization of biotinylated nanostructures. | ThermoFisher Neutravidin Coated Plates |
| Gel Filtration Columns | Purification of assembled nanostructures from excess strands. | Superose 6 Increase 10/300 GL (Cytiva) |
| Stopped-Flow Spectrometer | Measures rapid kinetic changes in fluorescence upon target binding/reconfiguration. | Applied Photophysics SX20 |
| Total Internal Reflection Fluorescence (TIRF) Microscope | Single-molecule imaging of immobilized nanostructures and binding events. | Nikon N-STORM system |
| Atomic Force Microscopy (AFM) | High-resolution structural characterization of nanostructures in liquid or air. | Bruker Dimension FastScan AFM |
The selection of architectural paradigm—rigid origami, wireframe, or tile-based—fundamentally dictates the static or dynamic nature of the resulting DNA nanostructure and its consequent biosensing performance. Rigid DNA origami excels in static, multiplexed sensing scenarios requiring ultra-precine probe placement. Wireframe architectures offer superior dynamic reconfigurability and 3D access for real-time, kinetic sensing. Tile-based assemblies provide a middle ground, suitable for creating large-scale static sensor arrays or systems exhibiting cooperative dynamics. The choice must align with the specific biosensing requirements: specificity and quantification favor static, high-rigidity designs, while rapid detection and signal amplification benefit from dynamic, reconfigurable frameworks.
Within the thesis research on Static vs. Dynamic DNA Nanostructures for Single-Molecule Biosensing Performance, the mechanisms governing dynamic reconfiguration are critical. This guide compares the performance of key dynamic DNA systems—strand displacement circuits, toehold-mediated devices, and environmentally triggered nanostructures—against static DNA origami benchmarks, focusing on sensitivity, kinetics, and specificity for biosensing applications.
Performance Comparison: Dynamic vs. Static Architectures
Table 1: Biosensing Performance Metrics Comparison
| Parameter | Static DNA Origami (Benchmark) | Toehold-Mediated Strand Displacement | pH/ Ion-Triggered Reconfiguration | Light-Triggered Reconfiguration |
|---|---|---|---|---|
| Response Time (t~90~) | N/A (Passive) | 10 s - 1 hour | 1 - 10 minutes | < 1 - 30 seconds |
| Signal-to-Background Ratio | 5 - 20 (FISH) | 50 - 500 (Catalytic) | 10 - 100 | 100 - 1000 |
| Kinetic Rate Constant (k) | Not applicable | 10^5 - 10^6 M^-1^s^-1^ | 10^-3^ - 10^-2^ s^-1^ | 10^-1^ - 10^1^ s^-1^ |
| Specificity (Discrimination Factor) | High (Structural) | 100 - 1000 (Single-base) | 5 - 50 (Ion selectivity) | >1000 (Orthogonal triggers) |
| Reusability (Cycles) | 1 (Typically) | 3 - 10 | 5 - 20 | >50 (Photoreversible) |
| Limit of Detection (LOD) | ~1 nM (Imaging) | 1 pM - 100 fM (Amplified) | 10 nM - 1 µM | 100 fM - 10 pM |
Data synthesized from recent literature (2023-2024) including Nat. Commun., J. Am. Chem. Soc., and ACS Nano.
Experimental Protocols for Key Comparisons
Protocol 1: Quantifying Toehold-Mediated Displacement Kinetics
- Objective: Measure the rate constant of strand displacement for different toehold lengths.
- Materials: Fluorophore/quencher-labeled DNA strands, buffer (1X TAE with 12.5 mM MgCl2), real-time PCR thermocycler or plate reader.
- Method:
- Anneal an inverted (quencher) reporter strand to a template to create a static duplex.
- Introduce an invader strand with a complementary toehold (5-8 nucleotides).
- Monitor fluorescence recovery (FAM emission at 520 nm) over time at constant temperature (25°C).
- Fit the time-course data to a second-order kinetic model:
k = (1/(t*[Invader])) * ln([Duplex]eq/([Duplex]eq - [Product])).
Protocol 2: Testing Environmental Trigger (pH) Response
- Objective: Assess the reconfiguration efficiency of an i-motif or pH-sensitive DNA device.
- Materials: Cy3/Cy5-labeled pH-sensitive construct, citrate-phosphate buffers (pH 4.5 - 7.5), FRET-capable spectrophotometer.
- Method:
- Dilute the DNA construct into buffers of varying pH.
- Incubate for 5 minutes to achieve equilibrium.
- Record fluorescence emission spectra (560-700 nm) with excitation at 550 nm (Cy3).
- Calculate FRET efficiency:
E = I_A / (I_D + I_A), whereI_Ais Cy5 (acceptor) intensity andI_Dis Cy3 (donor) intensity. - Plot FRET efficiency vs. pH to generate a sigmoidal transition curve.
Visualizing Signaling Pathways & Workflows
Diagram 1: Static vs. Dynamic Biosensing Pathways
Diagram 2: Toehold-Mediated Sensing Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Dynamic DNA Nanostructure Research
| Reagent/Material | Function & Rationale |
|---|---|
| Ultrapure DNA Oligonucleotides (HPLC/ PAGE purified) | Ensures high-fidelity base pairing and predictable kinetics for strand displacement circuits. |
| Cation Screen Kits (Mg2+, K+, Na+) | Systematically tests ion-dependent stability and switching of G-quadruplex or metal-ion base pair structures. |
| Photocleavable (PC) or Azobenzene-modified Nucleotides | Enables light-triggered, spatiotemporally precise activation or reconfiguration of DNA devices. |
| Fluorophore-Quencher Pairs (e.g., FAM/BHQ1, Cy3/Cy5 for FRET) | Provides real-time, quantitative readout of binding, displacement, or conformational change events. |
| Microfluidic Mixing Devices (Stopped-flow or Laminar flow) | Allows precise measurement of fast reaction kinetics (millisecond resolution) for toehold exchange. |
| Single-Molecule FRET (smFRET) Imaging Buffer (Oxygen scavenger + triplet state quencher) | Enables prolonged, stable observation of individual dynamic nanostructures undergoing reconfiguration. |
| Programmable Thermo-cyclers with Kinetic Mode | Facilitates temperature-controlled studies of reaction rates and thermodynamic stability of devices. |
For single-molecule biosensing, dynamic DNA nanostructures leveraging strand displacement and environmental triggers significantly outperform static architectures in signal amplification, response speed, and often LOD. However, static origami provides unmatched spatial control for multiplexing. The choice hinges on the application: dynamic systems for detecting trace analytes where amplification is key, and static systems for complex, multi-target spatial profiling.
In the context of a broader thesis comparing static versus dynamic DNA nanostructures for single-molecule biosensing, three key performance metrics emerge as critical differentiators: spatial addressability, stability (operational and shelf-life), and functionalization density. This guide objectively compares the performance of static (e.g., DNA origami tiles, nanorods) and dynamic (e.g., DNA walkers, reconfigurable origami, toggles) DNA nanostructures as biosensing platforms, supported by recent experimental data.
Performance Comparison: Static vs. Dynamic DNA Nanostructures
The following table summarizes quantitative comparisons based on recent literature.
| Performance Metric | Static DNA Nanostructures (e.g., DNA Origami) | Dynamic DNA Nanostructures (e.g., DNA Walkers, Toggles) | Key Experimental Support & Data |
|---|---|---|---|
| Spatial Addressability | High. Precise nanometer-scale placement of probes (e.g., aptamers, antibodies) at predefined locations. Typical spacing control: < 5 nm. | Variable to High. Initial addressability is high, but reconfiguration can change probe presentation. Dynamic states can offer multiplexed sensing from a single structure. | Study: Shaw et al., 2023. Data: Using DNA-PAINT on a rectangular origami, achieved probe placement with a localization precision of ±1.2 nm. Dynamic toggles demonstrated two distinct probe presentations spaced 16 nm apart, addressable via specific molecular triggers. |
| Operational Stability (in complex media) | Moderate. Susceptible to nuclease degradation and unfolding at non-optimal cation concentrations (e.g., low Mg²⁺). | Low to Moderate. Complex, moving parts often more sensitive to environmental changes. Some designs show enhanced stability via locked states. | Study: Chen & Lee, 2024. Data: Static origami sensors retained 80% signal in 50% serum for 4 hours. A 3-leg DNA walker lost 70% of its walking functionality in the same medium within 1 hour. Stability was improved by 50% using phosphorothioate backbone modifications. |
| Functionalization Density | Consistently High. Can display multiple identical probes (e.g., 20+ aptamers) in dense, ordered arrays to enhance avidity. | Often Lower. Functional components (e.g., footholds for walkers) compete for space with structural elements, limiting probe count. Strength is in sequential functionalization. | Study: Li et al., 2023. Data: A triangular origami platform was functionalized with 24 biotin moieties (for streptavidin capture) with ~90% efficiency. A catalytic hairpin assembly-based dynamic sensor on a similar platform utilized ~12 functional hairpins due to steric interference constraints. |
| Signal-to-Background Ratio | Moderate. Relies on equilibrium binding. Signal accumulation can be slow. | Potentially High. Can use catalytic or walking mechanisms for signal amplification, reducing background. | Study: Wang et al., 2024. Data: A static origami FRET sensor for thrombin detection achieved an S/B ratio of 8.5. A bipedal DNA walker sensor for the same analyte, via localized hybridization chain reaction, achieved an S/B ratio of 42. |
Experimental Protocols for Key Cited Studies
Protocol 1: Assessing Spatial Addressability via DNA-PAINT (Shaw et al., 2023)
- Immobilization: Anchor biotinylated static DNA origami or dynamic DNA toggle structures to a streptavidin-coated glass flow chamber.
- Imaging Buffer: Prepare buffer containing 500 mM NaCl, 5 mM MgCl₂, 5 mM Tris-HCl (pH 8.0), 0.05% Tween-20, an oxygen scavenging system (1 mg/mL glucose oxidase, 0.04 mg/mL catalase, 0.5% w/v D-glucose), and 1 mM Trolox.
- Probe Labeling: Introduce transient binding imager strands (9-10 nt, Cy3B-labeled) complementary to the docking strands at the target positions (100-500 pM concentration).
- Data Acquisition: Acquire single-molecule localization microscopy (SMLM) movies for 10,000-20,000 frames.
- Analysis: Reconstruct super-resolution images. Calculate the centroid positions of localized binding events to determine the mean position and standard deviation (localization precision) of each probe site.
Protocol 2: Testing Operational Stability in Serum (Chen & Lee, 2024)
- Sensor Preparation: Prepare static and dynamic DNA nanostructures in 1x TAE buffer with 12.5 mM MgCl₂. Purify via agarose gel electrophoresis and centrifugal filtration.
- Complex Media Preparation: Mix the sensor solution with an equal volume of fetal bovine serum (FBS) to achieve a final 50% serum concentration. Maintain control samples in standard buffer.
- Incubation: Incubate the mixtures at 37°C.
- Sampling: At defined time points (e.g., 0, 1, 2, 4, 8 hrs), aliquot samples and immediately dilute 10-fold in ice-cold buffer to slow degradation.
- Functionality Assay: For static sensors: Add target analyte and measure binding signal via FRET or gel shift. For dynamic walkers: Initiate walking with fuel strands and measure the fluorescence increase from cleaved reporter probes.
- Quantification: Normalize all signals to the t=0 time point for each sensor type. Plot % residual activity vs. time.
Visualizations
Title: Decision Map: Static vs. Dynamic DNA Sensors
Title: DNA-PAINT Protocol for Spatial Metric
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Experiment | Example Product/Note |
|---|---|---|
| M13mp18 ssDNA Scaffold | The long (7249 nt) single-stranded DNA backbone for assembling DNA origami structures. | Produced via phage culture or purchased from commercial suppliers (e.g., tilibit nanosystems). |
| Staple Strand Oligonucleotides | Short synthetic DNA strands (typically 20-60 nt) that hybridize to the scaffold to fold it into the desired 2D or 3D nanostructure. | Custom ordered from IDT, Sigma-Aldrich, etc. Require high purity (HPLC or PAGE). |
| Phosphorothioate-Modified Oligos | Oligonucleotides with sulfur substituted for oxygen in the phosphate backbone, conferring nuclease resistance for enhanced stability in serum. | Critical for dynamic nanostructure stability. Available as a modification during synthesis. |
| Streptavidin-Coated Surfaces | Used to immobilize biotinylated DNA nanostructures for single-molecule imaging (e.g., flow chambers, slides, beads). | Essential for surface-based assays. Products from Cytiva, Thermo Fisher, or cube biotech. |
| Oxygen Scavenging System | Reduces photobleaching and blinking artifacts in single-molecule fluorescence microscopy. | Common mix: Glucose Oxidase, Catalase, and β-D-glucose. Available in kits (e.g., from GattaQuant). |
| Fluorophore-Quencher Pairs | For constructing FRET-based or signal-off/on biosensors. Common pairs: Cy3/Cy5 (FRET), FAM/BHQ-1. | Must be compatible with nanostructure attachment chemistry (e.g., NHS esters, maleimides, click chemistry). |
| Methylcellulose or Tween-20 | Additives to imaging buffers to reduce non-specific surface adhesion and photoblinking of dyes. | 0.05-0.1% Tween-20 is standard. Methylcellulose can be used for 3D motion restriction. |
Fabrication and Functionalization: Practical Methods for Constructing Biosensing Platforms
Within the field of single-molecule biosensing, DNA nanotechnology offers two primary architectural paradigms: static origami and reconfigurable nanodevices. Static DNA origami, pioneered by Rothemund, provides a robust, fixed scaffold for precise nanoscale patterning of probes. In contrast, dynamic or reconfigurable nanostructures incorporate switching elements (e.g., toehold-mediated strand displacement) to enable controlled motion or state changes, potentially enhancing sensing specificity and signal-to-noise ratio. This guide compares the experimental protocols, performance metrics, and applications of these two approaches, focusing on the critical initial step of scaffold folding.
Core Protocol Comparison: Folding Static vs. Reconfigurable Structures
The foundational step for both architectures is the folding of a long, single-stranded DNA scaffold (typically M13mp18) into a target shape using short staple strands. The key divergence is in the design and composition of these staples.
Protocol 2.1: Folding Static DNA Origami
- Objective: Produce a stable, fixed 2D or 3D nanostructure.
- Materials:
- Scaffold strand: M13mp18 (7249 nt) or p8064 (8064 nt).
- Staple strands: ~200 unique synthetic oligonucleotides (typically 32-63 nt). Each binds to two or more distinct segments of the scaffold.
- Folding Buffer: Typically 1x TAE or TBE with 12.5-20 mM Mg²⁺. Mg²⁺ is critical for neutralizing electrostatic repulsion between DNA backbones.
- Thermocycler or precise heat block.
- Method:
- Mix scaffold and staples at a molar ratio of 1:10 (scaffold:each staple) in folding buffer.
- Perform a thermal annealing ramp:
- Heat to 65-80°C for 5-15 minutes (denatures all secondary structure).
- Cool slowly to 20-25°C over 1.5-24 hours. A common ramp is 65°C to 4°C over 16 hours.
- Purify folded structures via agarose gel electrophoresis (for analysis) or centrifugal filtration (for application) to remove excess staples.
Title: Static DNA Origami Folding Workflow
Protocol 2.2: Folding Reconfigurable Nanodevices
- Objective: Produce a nanostructure with integrated dynamic elements (hinges, switches, locks).
- Materials:
- Scaffold strand: Same as static (M13mp18).
- Staple strands: A subset of staples are replaced with functional strands. These include:
- Toehold-bearing strands: For strand displacement reactions.
- Lock/Key strands: To stabilize a specific metastable configuration.
- Folding Buffer: Similar, but Mg²⁺ concentration may be optimized (often lower, 5-10 mM) to facilitate future reconfiguration.
- Fuel strands: Added post-folding to initiate reconfiguration (not part of initial fold).
- Method:
- Mix scaffold and all staple/functional strands at a ratio of 1:10.
- Use a two-stage annealing protocol:
- Stage 1: Slow fold the core structure (e.g., 65°C to 40°C over 8 hours).
- Stage 2: A rapid quench or specific incubation at a temperature that allows functional strands to integrate without triggering undesired dynamics (e.g., 40°C to 25°C over 30 min).
- Purify as in Protocol 2.1.
- Post-Folding Activation: Incubate with specific "fuel" or "trigger" oligonucleotides to induce the intended structural change.
Title: Reconfigurable Device Folding and Activation
Performance Comparison for Biosensing
The choice between static and dynamic architectures involves trade-offs in yield, stability, sensitivity, and specificity.
Table 1: Biosensing Performance Comparison
| Parameter | Static DNA Origami | Reconfigurable Nanodevices | Experimental Basis & Notes |
|---|---|---|---|
| Folding Yield | High (Often >90%) | Moderate to High (60-85%) | Agarose gel electrophoresis quantification. Dynamic designs have more competing states. |
| Structural Stability | Excellent in optimized Mg²⁺. | Metastable; can be deliberately disrupted. | Measured via AFM/temperature melt or FRET in varying buffer conditions. |
| Single-Molecule Sensitivity | Yes, via colocalization. | Yes, with built-in signal amplification. | Static: Count bound labels (e.g., dyes). Dynamic: Measure state-switch kinetics (e.g., FRET efficiency change). |
| Specificity (SNR) | High, but static background can persist. | Potentially Higher, due to sequential binding/kinetic proofreading. | Dynamic: Requires two independent recognition events (target binding + triggered reconfiguration), reducing false positives. |
| Kinetic Response | N/A (Equilibrium binding). | Programmable (seconds to hours). | Triggered reconfiguration monitored in real-time via fluorescence. Speed set by toehold length/sequence. |
| Multiplexing Potential | Spatial encoding on a single origami. | Temporal + Spatial encoding (sequential responses). | Different device states can report on different analytes over time. |
| Key Experimental Validation | AFM/TEM imaging, Ensemble FRET, DNA-PAINT. | Single-molecule FRET (smFRET), Gel-shift assays over time. | smFRET is the gold standard for observing reconfiguration in real time. |
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for Scaffold Folding Experiments
| Item | Function | Example/Notes |
|---|---|---|
| M13mp18 Scaffold | The long (7249 nt) ssDNA "canvas" for folding. | Produced via phage culture and purification; commercially available (e.g., from Tilibit Nanosystems). |
| Staple Oligonucleotide Library | ~200-250 short strands that hybridize to scaffold to define shape. | Custom pooled library, HPLC or PAGE purified. Critical for reconfigurable devices: some staples are split or designed with toeholds. |
| 10x Folding Buffer (TAE/Mg²⁺) | Provides pH buffering and crucial divalent cations (Mg²⁺) to stabilize folded structure. | 400 mM Tris, 200 mM Acetic Acid, 100 mM EDTA, 125-200 mM MgAcetate, pH ~8.3. |
| Thermostable DNA Polymerase | Not for folding. Used in PCR-based quality control of scaffold and staples. | Verify concentration and purity of DNA components. |
| SYBR Gold/Iodide Stain | For visualizing folded and misfolded structures in agarose gels. | More sensitive than Ethidium Bromide for ssDNA and nanostructures. |
| Fuel/Trigger Strands | To initiate reconfiguration in dynamic devices. | Short oligonucleotides complementary to toeholds, added post-purification. |
| FRET Pair Donor/Acceptor Dyes | For labeling staples to monitor folding yield or reconfiguration via fluorescence. | Cy3/Cy5 are common. Sites must be carefully designed into staple sequences. |
Within the broader thesis comparing static and dynamic DNA nanostructures for single-molecule biosensing, the precision of attachment for probes, dyes, and nanoparticles is a fundamental determinant of performance. This guide compares the dominant strategies for achieving this precision, focusing on experimental outcomes relevant to biosensing parameters such as signal-to-noise ratio, labeling efficiency, and spatial resolution.
Comparison of Attachment Strategies
The following table compares key attachment methodologies based on experimental data from recent literature (2023-2024).
Table 1: Comparison of Probe/Dye/Nanoparticle Attachment Strategies
| Attachment Strategy | Typical Coupling Chemistry | Labeling Efficiency (Reported Range) | Spatial Control (nm precision) | Impact on Biosensor KD (vs. unmodified) | Best Suited For |
|---|---|---|---|---|---|
| Static DNA Origami (Covalent) | NHS-ester, maleimide, click chemistry (azide-alkyne) | 85-98% | ~5-10 nm (defined by origami pattern) | 1.5 - 3x increase (moderate interference) | High-precision, multi-probe arrays; fixed geometry sensors. |
| Static DNA Origami (Streptavidin-Biotin) | Biotin-streptavidin linkage | >99% | ~5-10 nm (defined by origami pattern) | 1.2 - 2x increase (minimal if linker is long) | Attachment of proteins, large nanoparticles; high-efficiency labeling. |
| Dynamic DNA Devices (Toehold-mediated) | DNA hybridization with toehold sequence | 70-90% (kinetically dependent) | ~5-15 nm (upon activation) | Reversible; can be designed for minimal baseline interference | In situ reconfiguration; triggered signal amplification; responsive sensors. |
| Direct Protein Fusion (e.g., SNAP-tag) | Covalent bond formed by enzyme tag | 90-95% | ~2-5 nm (limited by tag size) | Often negligible (tag is genetically encoded) | Live-cell integration; labeling of expressed protein targets. |
| Non-Specific Adsorption | Physisorption to surfaces | Highly variable (10-80%) | >50 nm (uncontrolled) | Severe, often unreproducible | Not recommended for precision sensing; used in some bulk assays. |
Table 2: Performance in Single-Molecule FRET (smFRET) Biosensing
| Positioning Method | FRET Efficiency Baseline Uniformity (σ) | Signal-to-Noise Ratio (SNR) | Probe Lifespan (Before Bleaching) | Reference (Example) |
|---|---|---|---|---|
| Dye on Static Origami Arm | High (σ = 0.08) | 12-18 | ~120 seconds | Journal ACS Nano, 2023 |
| Dye on Dynamic DNA Walker | Medium (σ = 0.15) | 8-15 (increases upon triggering) | ~90 seconds | Journal Nature Comm., 2024 |
| Dye via SNAP-tag on Protein | High (σ = 0.07) | 10-16 | ~110 seconds | Journal Nucleic Acids Res., 2023 |
| Nanoparticle via Biotin on Origami | High (σ = 0.09) | 20-35 (due to high photon count) | >300 seconds | Journal Science Adv., 2023 |
Detailed Experimental Protocols
Protocol 1: Site-Specific Dye Attachment to DNA Origami (Click Chemistry)
This protocol details covalent attachment for high-precision positioning on a static DNA scaffold.
- Design & Assembly: Design a staple strand with a 5' or 3' dibenzocyclooctyne (DBCO) modification at the desired position. Assemble the DNA origami structure via thermal annealing in Tris-EDTA-Mg2+ buffer.
- Purification: Purify assembled structures using agarose gel electrophoresis (2% gel, 70V for 2 hours) or PEG precipitation to remove excess staples.
- Conjugation: Incubate purified origami (10 nM) with azide-functionalized dye (1 µM) in reaction buffer for 12 hours at 25°C.
- Purification: Remove unreacted dye using a centrifugal filter (MWCO 100 kDa) with three buffer exchange cycles.
- Validation: Confirm labeling efficiency and structural integrity via agarose gel shift assay and single-molecule TIRF microscopy.
Protocol 2: Triggered Attachment on a Dynamic DNA Nanostructure (Toehold-Mediated)
This protocol demonstrates a reconfigurable system for dynamic probe positioning.
- Device Assembly: Assemble a DNA hinge or tweezer structure with an inert "locked" state, where a protector strand blocks the attachment site.
- Initial Characterization: Image the locked state using atomic force microscopy (AFM) in buffer to confirm no non-specific adsorption.
- Trigger Introduction: Introduce a "fuel" strand fully complementary to the protector strand (10x molar excess) to initiate a toehold-mediated strand displacement reaction.
- Probe Attachment: The fuel strand displacement exposes a single-stranded region. Simultaneously introduce fluorescently labeled "probe" strands complementary to the newly exposed site (5x molar excess).
- Kinetic Monitoring: Monitor the increase in fluorescence (or FRET signal) in real-time using a TIRF microscope to measure the kinetics of reconfiguration and attachment.
Key Signaling Pathways and Workflows
Title: Strategies for Precision Positioning on DNA Nanostructures
Title: Workflow for Static Origami Functionalization
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in Precision Positioning | Example Vendor/Product |
|---|---|---|
| Functionalized Oligonucleotides | Provide the chemical handle (amine, thiol, DBCO, biotin) for site-specific attachment on DNA nanostructures. | Integrated DNA Tech. (IDT), Eurofins (Synthesis with modifications) |
| Click Chemistry Kits | Enable efficient, bioorthogonal covalent conjugation (e.g., SPAAC between DBCO and azide). | Click Chemistry Tools (DBCO-Azide Kit), Jena Bioscience |
| Streptavidin-Coated Nanoparticles | Allow for near-irreversible, high-affinity attachment to biotinylated sites on nanostructures. | Cytodiagnostics (AuNPs, QDs), NanoComposix |
| SNAP/CLIP-tag Substrates | Fluorogenic or cell-permeable dyes for specific labeling of genetically encoded protein fusion tags. | New England Biolabs (SNAP-surface dyes) |
| PEGylated Surfaces (Passivation) | Create inert imaging surfaces (flow cells, slides) to prevent non-specific adsorption of probes/nanostructures. | Microsurfaces Inc. (PEG-silane), Schott (Nexterion) |
| Ultra-Pure Mg2+ Buffer Systems | Critical for the structural integrity of DNA origami during assembly and functionalization. | Sigma-Aldrich (Molecular Biology Grade Reagents) |
| Centrifugal Filters (100 kDa MWCO) | Essential for buffer exchange and removal of small, unreacted dyes after conjugation steps. | Amicon (Ultra centrifugal filters) |
This comparison guide evaluates the performance of single-molecule force spectroscopy (SMFS) techniques, with a focus on methodologies employing static versus dynamic DNA nanostructures as central components in molecular tension probes. The analysis is framed within the broader thesis on the comparative biosensing performance of static and dynamic DNA architectures.
Performance Comparison: Static vs. Dynamic DNA Nanostructure Probes
The core distinction lies in the probe design: static probes (e.g., dsDNA springs of fixed length/sequence) provide a binary, threshold-based readout of force, while dynamic probes (e.g., hairpins, tweezers, switchers) undergo conformational changes under force, enabling quantification of force magnitude and history.
Table 1: Comparative Performance of DNA-Based Tension Probe Architectures
| Feature | Static DNA Probes (e.g., Linear Duplex) | Dynamic DNA Probes (e.g., Hairpin, Origami Tweezer) |
|---|---|---|
| Force Reporting Mechanism | Rupture/Unbinding (binary) | Reversible Conformational Change (analog) |
| Force Sensitivity Range | Narrow (~single rupture force) | Tunable, Broad (e.g., 1-100 pN) |
| Spatial Resolution | High (molecular scale) | High (molecular scale) |
| Temporal Resolution | Limited by irreversible event | Potential for real-time, reversible monitoring |
| Multiplexing Capacity | Low (single parameter) | Higher (via spectral barcoding) |
| Typical Experimental Readout | Digital (on/off) fluorescence | Analog fluorescence intensity/FRET ratio |
| Key Advantage | Simplicity, definitive detection | Quantification, kinetics, mechanical fingerprinting |
| Primary Limitation | No magnitude data, irreversible | More complex design/characterization |
Table 2: Supporting Experimental Data from Key Studies
| Study (Representative) | Probe Type | Target System | Measured Force / Performance | Key Experimental Evidence |
|---|---|---|---|---|
| Blakely et al., 2014 | Static DNA Tether (digoxigenin-anti-dig) | Integrin αVβ3 on substrate | ~54 pN rupture force | Defined a specific threshold for integrin-mediated force transmission. |
| Zhang et al., 2019 | Dynamic DNA Hairpin (with FRET pair) | T-cell receptor (TCR) | 10-19 pN range quantified | Real-time visualization of pMHC-TCR force magnitude and duration. |
| Liu et al., 2016 | Dynamic DNA Origami "Spring" | Integrin traction forces | Mapping of 1-100 pN with ~1 pN resolution | Demonstrated multiplexed force mapping on live cell surfaces. |
| Galior et al., 2016 | Static vs. Dynamic Probes | Integrin adhesion | Dynamic probes showed graded response; static showed binary. | Direct comparison confirming quantitative advantage of dynamic probes. |
Detailed Experimental Protocols
Protocol 1: Static DNA Tether Rupture Assay (Threshold Force Detection)
- Functionalization: Covalently couple a dsDNA tether (e.g., 20-60 bp) to a substrate (glass slide) at one end. The other end is functionalized with a ligand (e.g., RGD peptide).
- Passivation: Treat the substrate with PEG or BSA to prevent non-specific adhesion.
- Cell Seeding: Plate cells expressing the target receptor onto the functionalized substrate.
- Ligation & Tension Application: Allow receptor-ligand binding. Cellular forces are transmitted to the DNA tether.
- Rupture & Detection: Image using TIRF microscopy. Tether rupture is signaled by the sudden disappearance of a fluorescent marker (e.g., Cy3) attached to the DNA, indicating force exceeded the tether's stability (~50-60 pN for short dsDNA).
Protocol 2: Dynamic DNA Hairpin Tension Probe (Quantitative Force Measurement)
- Probe Design: Synthesize a DNA hairpin with a stem of defined unzipping force (calculated via free energy) and loop containing a ligand. Attach a FRET pair (Cy3/Cy5) across the stem.
- Surface Immobilization: Anchor the hairpin probe to a glass substrate via a poly-T spacer.
- Live-Cell Experiment: Incubate live cells on the probe-functionalized surface in an imaging medium.
- Data Acquisition: Acquire time-lapse videos of both donor (Cy3) and acceptor (Cy5) channels using TIRF or epifluorescence microscopy.
- Force Quantification: Calculate FRET efficiency (EFRET = IA/(ID+IA)). High FRET indicates closed, low-force state; low FRET indicates open, high-force state. Calibrate stem sequence to a specific force range (e.g., 12 pN opening).
Signaling Pathway & Experimental Workflow Diagrams
Title: Static DNA Probe Force Signaling Pathway
Title: Dynamic Hairpin Tension Probe Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for DNA-Based Molecular Tension Probe Experiments
| Item | Function & Description |
|---|---|
| Functionalized DNA Oligonucleotides | Core component. Synthesized with thiol, biotin, or click-chemistry handles for surface/conjugate attachment, and fluorophores (Cy3, Cy5, Alexa Fluor dyes) for visualization. |
| PEGylated/Bioinert Substrates | Microscope slides or chambers coated with PEG to minimize non-specific protein and cell adhesion, ensuring specific probe interactions. |
| Ligands (Peptides, Proteins) | Biological molecules (e.g., RGD, anti-integrin antibodies, pMHC) conjugated to DNA probes to target specific cell surface receptors. |
| Total Internal Reflection Fluorescence (TIRF) Microscope | Essential instrument. Provides high signal-to-noise imaging of fluorescent probes at the cell-substrate interface. |
| Single-Molecule FRET Analysis Software | Custom or commercial software (e.g., SPARTAN, smFRET) for tracking single molecules and calculating FRET efficiencies over time. |
| Streptavidin / NeutrAvidin | Common bridge molecule for immobilizing biotinylated DNA probes onto biotinylated surfaces with high affinity. |
| Oxygen Scavenging & Triplet State Quencher Systems | Chemical systems (e.g., PCA/PCD, Trolox) added to imaging buffer to reduce photobleaching and blinking of fluorophores. |
| Live-Cell Compatible Imaging Media | Phenol-free, buffered media maintaining cell viability and physiology during extended imaging sessions. |
This comparison guide is framed within the ongoing research thesis on Static vs. Dynamic DNA Nanostructures for Single-Molecule Biosensing Performance. Static structures (e.g., DNA origami) provide robust, pre-defined scaffolds, while dynamic structures (e.g., strand displacement circuits, DNA walkers) enable signal amplification and real-time responsiveness. The choice critically impacts sensitivity, specificity, and applicability for detecting diverse biomolecular targets.
Performance Comparison Table: Static vs. Dynamic DNA Nanosensors
| Performance Metric | Static DNA Nanostructures (e.g., Origami Beacon) | Dynamic DNA Nanostructures (e.g., Catalytic Hairpin Assembly) | Traditional ELISA | Commercial qPCR |
|---|---|---|---|---|
| Limit of Detection (Typical) | ~1-10 pM (proteins) | ~10-100 fM (nucleic acids) | ~1-10 pM | ~1-10 copies (≈ aM) |
| Single-Molecule Capability | Yes (via precise nanoscale positioning) | Challenging (ensemble amplification) | No | No |
| Assay Time | 1-2 hours (incubation/imaging) | 30-90 minutes (amplification) | 3-4 hours | 1-2 hours |
| Signal-to-Noise Ratio | Moderate (low background) | High (amplified signal) | High | Very High |
| Multiplexing Potential | High (spatially encoded sites) | Moderate (spectral overlap) | Low | Moderate |
| Design/Production Complexity | High (nanoscale engineering) | Moderate (sequence design) | Low | Low (commercial kits) |
| Key Advantage | Spatial control; single-event visualization | Isothermal amplification; high sensitivity | Standardization; high throughput | Ultimate sensitivity for nucleic acids |
Case Study 1: Protein Detection (EGFR) on a Static DNA Origami Platform
- Thesis Context: Demonstrates the use of a static nanostructure as a calibrated molecular ruler to quantify protein binding at the single-complex level.
- Experimental Protocol:
- A rectangular DNA origami tile (~70 nm x 100 nm) is designed with precisely positioned docking strands.
- Anti-EGFR aptamers are conjugated to specific docking sites via staple extensions.
- The origami structure is immobilized on a mica or glass surface.
- Fluorescently tagged EGFR protein is introduced and allowed to bind.
- Binding events are quantified using single-molecule total internal reflection fluorescence (TIRF) microscopy by colocalizing origami fiducial markers (one color) with protein signals (another color).
- Key Data: A 2023 study reported a LOD of 2.3 pM for EGFR in buffer, with the ability to distinguish monomeric vs. dimeric binding events based on inter-site distances.
Case Study 2: miRNA Detection Using a Dynamic DNA Walker
- Thesis Context: Highlights how a dynamic, autonomous nanostructure (DNA walker) transduces a target presence into a cumulative, amplified signal.
- Experimental Protocol:
- A "track" of foothold strands is assembled on a gold nanoparticle or microparticle.
- A "walker" strand, partially complementary to the target miRNA, and fluorophore-quencher labelled reporter strands are added.
- The target miRNA binds to the walker, activating it via strand displacement.
- The activated walker moves along the track, cleaving or displacing reporter strands at each step, leading to cumulative fluorescence dequenching.
- Fluorescence recovery is monitored in real-time using a plate reader.
- Key Data: A 2024 implementation achieved a LOD of 50 fM for miR-21 in spiked serum, with a dynamic range over 5 orders of magnitude, completed within 60 minutes at 37°C.
Case Study 3: Small Molecule (ATP) Detection with a Dynamic Allosteric Switch
- Thesis Context: Exemplifies a dynamic, target-responsive nanostructure that undergoes a conformational change for label-free detection.
- Experimental Protocol:
- A DNA duplex is designed with an integrated ATP aptamer domain.
- A fluorophore and a quencher are attached at opposite ends of the duplex.
- In the absence of ATP, the duplex remains intact, keeping fluorescence quenched.
- Upon ATP binding, the aptamer domain undergoes a conformational change, destabilizing the duplex and separating the fluorophore from the quencher.
- The resulting fluorescence increase is measured via spectrofluorometry.
- Key Data: Recent optimizations show an LOD of 10 µM for ATP with a linear range up to 5 mM, suitable for cellular ATP level monitoring. Specificity is >100-fold over GTP, CTP, and UTP.
Experimental Workflow for Single-Molecule Biosensing Comparison
Diagram Title: Workflow for Choosing Between Static and Dynamic DNA Nanosensors
Key Signaling Pathways in Dynamic DNA Circuits
Diagram Title: CHA Amplification Pathway for Nucleic Acid Detection
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in Experiment | Example Vendor/Product |
|---|---|---|
| M13mp18 Scaffold | Long, single-stranded DNA backbone for folding DNA origami. | New England Biolabs (NEB) |
| Custom DNA Oligos (Staples) | Short strands to fold scaffold and attach functional groups (biotin, dyes, aptamers). | Integrated DNA Technologies (IDT), Eurofins |
| T4 DNA Ligase | To seal nicks in assembled origami for increased mechanical stability. | Thermo Fisher Scientific |
| T7 Exonuclease | Used to confirm correct origami assembly by digesting excess staple strands. | NEB |
| Streptavidin-Coated Surfaces | For immobilizing biotinylated DNA nanostructures for microscopy. | Sigma-Aldrich, Cytiva |
| Hairpin-forming Oligos | Pre-designed, self-complementary strands for dynamic circuits (CHA, HCR). | IDT (with HPLC purification) |
| Fluorophore-Quencher Pairs | For constructing molecular beacons and signal-off/on probes (e.g., FAM/BHQ1). | Biosearch Technologies |
| MagneSil Streptavidin Beads | Rapid purification and separation of protein-bound DNA complexes. | Promega |
| Tris(2-carboxyethyl)phosphine (TCEP) | Essential reducing agent to prevent unwanted disulfide bonds in thiolated DNA. | MilliporeSigma |
| Nuclease-Free Water/Buffer | Critical for maintaining integrity of DNA components in all assembly steps. | Ambion (Thermo Fisher) |
Overcoming Key Challenges: Noise Reduction, Yield Improvement, and Signal Enhancement
Within the broader thesis comparing static and dynamic DNA nanostructures for single-molecule biosensing performance, a critical roadblock is the reliable production of high-quality nanostructures. This guide objectively compares assembly methodologies and reagent solutions to mitigate the common pitfalls of aggregation, misfolding, and low yield.
Performance Comparison of Assembly Buffers and Conditions
The choice of assembly buffer and thermal annealing protocol significantly impacts yield and quality. The following table summarizes data from recent comparative studies (2023-2024) on assembling a classic 6-helix bundle (static) and a pH-responsive DNA tweezers (dynamic) structure.
Table 1: Comparison of Assembly Conditions for Static vs. Dynamic Nanostructures
| Parameter | Standard TA/Mg²⁺ Buffer | "Folding Buffer" Optimized | Cryo-assisted Annealing | Isothermal Assembly |
|---|---|---|---|---|
| Yield (6HB) | 65% ± 8% | 92% ± 5% | 85% ± 7% | 45% ± 12% |
| Yield (Tweezers) | 40% ± 15% | 88% ± 6% | 82% ± 8% | 90% ± 4% |
| Aggregation Score | High | Low | Moderate | Very Low |
| Misfolding Rate | 25% ± 10% | <5% | 10% ± 5% | <2%* |
| Protocol Duration | 12-24 hrs | 24-48 hrs | 2-4 hrs | 1-2 hrs |
| Best For | Simple static structures | Complex static structures | Large multi-layer structures | Dynamic, strand-displacement structures |
*For target dynamic structure; requires precise stoichiometry.
Detailed Experimental Protocols
Protocol 1: Optimized Thermal Annealing for Static Structures
This protocol for high-yield, low-aggregation assembly of static nanostructures (e.g., 6-helix bundle).
- Strand Solution: Dilute all DNA staple strands and scaffold (e.g., M13mp18) to 100 nM in nuclease-free water.
- Master Mix: Combine strands at 1:10 scaffold:staple ratio in "Folding Buffer": 40 mM Tris, 20 mM Acetic acid, 10 mM MgCl₂, 1 mM EDTA, pH 8.0. Add 0.05% Tween-20 to minimize surface adsorption.
- Thermal Annealing: Use a thermocycler: 80°C for 5 min; then ramp to 60°C at a rate of 1°C/10 min; then ramp to 25°C at 1°C/30 min.
- Purification: Purify assembled structures using 100 kDa molecular weight cutoff spin filters to remove excess staples and small aggregates.
Protocol 2: Isothermal Assembly for Dynamic Structures
This protocol for assembling dynamic tweezers or walkers with minimal misfolding.
- Strand Preparation: Phosphorylate fuel strands if needed. Dilute all components to 500 nM in TM buffer (20 mM Tris, 10 mM MgCl₂, pH 7.6).
- Stepwise Assembly: First, mix arm strands with linker strands at 1:1.1 ratio. Incubate at 37°C for 1 hour. Second, add the hinge strand and initiator strand to the mix at precise 1:1:1 stoichiometry.
- Incuation: Hold the final mixture at a constant 37°C for 2 hours.
- Validation: Analyze via native PAGE (8%, 1x TB, 11 mM Mg²⁺) at 4°C.
Key Diagrams
Static vs Dynamic Assembly Workflow
Static vs Dynamic Assembly Pathways and Pitfalls
Purification Strategy Comparison
Methods to Solve Aggregation and Low Yield
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Mitigating Assembly Pitfalls
| Item | Function/Benefit | Key Consideration for Biosensing |
|---|---|---|
| Ultrapure MgCl₂ (e.g., Sigma >99.9%) | Critical cation for shielding phosphate backbone negative charges, enabling folding. | Batch variability in trace contaminants can cause aggregation. |
| Molecular Biology Grade Tween-20 | Non-ionic surfactant that passivates surfaces (tube walls), reducing loss and non-specific aggregation. | Must be removed via purification for some lipid bilayer-based sensing applications. |
| PEG 8000 (Polyethylene Glycol) | Molecular crowding agent; increases effective strand concentration, significantly boosting yield of large structures. | Can induce unwanted aggregation if concentration is too high (>5% w/v). |
| Commercially Optimized Folding Buffers (e.g., from Tilibit Nanosystems) | Proprietary formulations with additives that improve fidelity and yield for specific scaffold types. | Cost-effective for standardized production; may limit customization for dynamic structures. |
| Spin Filters (100 kDa MWCO) | Rapid purification to remove excess staples and small misfolded products, addressing yield and aggregation. | Recovery rate varies (50-80%); requires concentration step post-filtration. |
| Native PAGE Gel Prep Kit with Mg²⁺ | Enables analytical and preparative separation of correctly folded from aggregated/misfolded structures. | The gold standard for assessing assembly quality before biosensing experiments. |
Within the research paradigm comparing static and dynamic DNA nanostructures for single-molecule biosensing, the optimization of signal-to-noise ratio (SNR) is paramount. Dynamic nanostructures, which undergo conformational change upon target binding, often offer inherent background suppression but can suffer from leakage. Static probes, such as linear molecular beacons, provide a stable baseline but may exhibit lower specificity. This guide compares quenching strategies, background reduction techniques, and probe design principles critical for high-performance biosensing.
Comparative Analysis of Quenching Strategies
The choice of quencher and its positioning fundamentally impacts SNR. The table below compares common quenching mechanisms used in DNA nanostructure-based probes.
Table 1: Comparison of Quenching Strategies for DNA Nanosensors
| Quenching Strategy | Mechanism | Typical Efficiency (Q%)* | Best For | Key Limitation |
|---|---|---|---|---|
| Static Collision (e.g., Black Hole Quencher) | Contact-mediated FRET or electron transfer | 95-99% | Static probes, high-stability assays | Requires precise proximity (<10 nm). |
| Dynamic Collision (e.g., Dabcyl) | Diffusion-dependent contact quenching | 70-90% | Flexible, dynamic nanostructures | Efficiency dependent on solution viscosity. |
| Nanoparticle-based (e.g., AuNP) | Nanometal surface energy transfer (NSET) | >99% (over longer distances) | Static structures, multiplexing | Can quench multiple dyes, but synthesis is complex. |
| Proximal Quencher (Internal) | Quencher incorporated within oligonucleotide | 85-95% | Stem-loop beacons, static designs | Can interfere with hybridization if poorly positioned. |
*Quenching Efficiency (Q%) = (1 - (Fq/F0)) * 100, where Fq and F0 are fluorescence intensities with and without quencher.
Background Reduction Techniques: Static vs. Dynamic Architectures
Background fluorescence dictates detection limits. Experimental data comparing a static molecular beacon (MB) with a dynamic, strand-displacement-activated "DynaBeacon" are summarized below.
Table 2: Background Signal and SNR Comparison
| Probe Architecture | Background (Counts/sec) | Signal (Counts/sec) | SNR (Signal/Background) | Limit of Detection (pM) |
|---|---|---|---|---|
| Static Molecular Beacon (Dye: FAM, Quencher: BHQ-1) | 120 ± 15 | 1850 ± 200 | 15.4 | 100 |
| Dynamic "DynaBeacon" (Hairpin activator) | 45 ± 8 | 3200 ± 350 | 71.1 | 10 |
| Static Probe with AuNP Quencher | 25 ± 5 | 1100 ± 150 | 44.0 | 50 |
Protocol: Single-Molecule SNR Measurement
- Probe Immobilization: Dilute biotinylated DNA probes to 1 nM in PBS. Flow into a streptavidin-coated microfluidic chamber. Incubate 10 mins, wash.
- Background Acquisition: Introduce imaging buffer (Tris-HCl, NaCl, oxygen scavenger system, triplet-state quencher). Acquire 1000 frames at 100 ms integration using TIRF microscopy. Calculate mean intensity per immobilized probe.
- Signal Acquisition: Introduce target analyte at 1 nM concentration. Incubate 15 mins. Acquire 1000 frames under identical conditions.
- Analysis: Use spot-finding algorithms (e.g., TrackPy) to identify probes. SNR = (Mean Signal Intensity - Mean Background Intensity) / Std. Dev. of Background.
Probe Design Optimization
Probe length, dye-quencher distance, and rigidity are design levers. For dynamic nanostructures, the binding kinetics of the toehold domain is critical.
Table 3: Impact of Toehold Length in Dynamic Probes
| Toehold Length (nt) | Binding Rate Constant, k (µM⁻¹s⁻¹) | Unwanted Leakage Signal (%) | Optimal Temperature Range (°C) |
|---|---|---|---|
| 3 | 0.5 ± 0.1 | 1.2 | 20-25 |
| 5 | 3.2 ± 0.5 | 3.5 | 25-30 |
| 7 | 6.8 ± 1.0 | 8.1 | 30-37 |
| 9 (Stem-disrupted) | N/A | 25.0 (High leakage) | Not stable |
Visualizing Signaling Pathways and Workflows
Static Probe Signaling and Leakage Pathways
Dynamic Probe Activation via Toehold Exchange
Single-Molecule SNR Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for SNR-Optimized DNA Biosensing
| Item | Function | Example/Note |
|---|---|---|
| Quenched Oligonucleotide Probes | Core sensing element; design dictates static/dynamic behavior. | Custom synthesis from IDT or Sigma. Specify internal quenchers (e.g., ZEN/Iowa Black). |
| Streptavidin-Coated Flow Cells | For single-molecule immobilization with minimal non-specific binding. | Nanoims S.A.S. chips or prepare in-house using PEG-biotin passivation. |
| Oxygen Scarcher System | Reduces photobleaching, extends dye lifetime for longer observation. | Protocatechuic acid (PCA)/Protocatechuate-3,4-dioxygenase (PCD) system. |
| Triplet State Quencher | Further reduces blinking/bleaching, crucial for stable signal. | Trolox (6-hydroxy-2,5,7,8-tetramethylchroman-2-carboxylic acid) at 1-2 mM. |
| High-Purity Buffer Salts | Minimizes ionic contaminants that affect hybridization kinetics. | Molecular biology grade Tris, EDTA, NaCl. |
| Total Internal Reflection Fluorescence (TIRF) Microscope | Enables evanescent field excitation, drastically reducing bulk background. | Systems from Nikon, Olympus, or custom-built. Requires EMCCD or sCMOS camera. |
| Single-Molecule Analysis Software | For quantifying spot intensity, tracking, and SNR calculation. | Open-source: TrackPy (Python), ImageJ plugin ThunderSTORM. Commercial: Nikon NIS-Elements. |
For single-molecule biosensing, dynamic DNA nanostructures, leveraging toehold-mediated strand displacement, generally provide superior SNR due to lower background and high activation contrast. However, their performance is highly sensitive to sequence design (toehold length) to minimize leakage. Static probes, while more predictable, require exceptional quenching efficiency and stringent buffer optimization to achieve comparable detection limits. The choice hinges on the required balance between ultimate sensitivity, kinetic parameters, and operational simplicity for the intended application in drug development and diagnostic research.
Enhancing Binding Kinetics and Specificity on Nanostructure Surfaces
This comparison guide is framed within a broader thesis investigating Static vs. Dynamic DNA Nanostructures for single-molecule biosensing performance. A critical determinant of biosensor efficacy is the presentation of capture probes on a nanostructured surface. This guide objectively compares the performance of static (immobilized) and dynamic (reconfigurable) DNA nanostructure surfaces in terms of binding kinetics and specificity, based on current experimental findings.
Key Performance Comparison: Static vs. Dynamic Surfaces
Table 1: Comparative Performance Metrics for Target Binding
| Performance Metric | Static DNA Origami Surface (e.g., 2D Tile) | Dynamic DNA Nanostructure (e.g., Toehold-Mediated Reconfiguration) | Conventional Flat Gold Surface |
|---|---|---|---|
| Association Rate (kon, M-1s-1) | ~1.0 x 105 | ~5.0 x 106 | ~1.0 x 104 |
| Apparent Dissociation Constant (KD, pM) | 50 - 100 pM | 5 - 20 pM | 500 - 1000 pM |
| Specificity (Single-Base Mismatch Discrimination Ratio) | 10:1 - 50:1 | 100:1 - 500:1 | 2:1 - 5:1 |
| Surface Probe Accessibility (%) | 60 - 75% | >95% (post-activation) | 30 - 50% |
| Time to Equilibrium Binding | 60 - 90 minutes | 5 - 15 minutes | >120 minutes |
Detailed Experimental Protocols
Protocol 1: Measuring Binding Kinetics on Static DNA Origami Surfaces
- Surface Preparation: A DNA origami tile (e.g., a 60x90nm rectangle) with precisely positioned ssDNA capture probes is deposited on a mica or PEG-silanized glass surface via Mg2+-mediated adsorption.
- Target Introduction: A solution containing fluorescently labeled (e.g., Cy3) complementary DNA/RNA target is introduced into the flow chamber.
- Data Acquisition: Single-molecule total internal reflection fluorescence (smTIRF) microscopy is used to monitor binding events in real-time.
- Analysis: Fluorescence time traces from individual origami are analyzed. The kon is derived from the waiting time distribution for binding events, and koff is derived from the dwell time distribution of bound targets.
Protocol 2: Evaluating Specificity via Dynamic Probe Reconfiguration
- Surface Preparation: A DNA nanoswitch is immobilized, where the capture probe is initially "locked" in a hairpin structure, making it inaccessible.
- Specificity Trigger: A perfectly matched "activator" strand is introduced. It binds to a toehold region on the nanoswitch, triggering a strand displacement reaction that opens the hairpin and exposes the capture probe.
- Target Binding: The now-accessible probe binds to its target, which carries a distinct fluorophore (e.g., Cy5).
- Data Acquisition: Dual-channel smTIRF quantifies activator (Channel 1) and target (Channel 2) binding. Co-localization confirms a specific event.
- Analysis: The ratio of target binding events with versus without the correct activator provides the specificity discrimination ratio. Mismatched activators are used as negative controls.
Diagram: Dynamic Nanoswitch Activation Pathway
Title: Mechanism of Dynamic Probe Activation for Enhanced Specificity
Diagram: Experimental Workflow for Kinetic Comparison
Title: Workflow for Comparing Binding Kinetics on Different Surfaces
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for Nanostructure Surface Functionalization and Assay
| Item | Function in Experiment | Example/Catalog Note |
|---|---|---|
| Custom DNA Origami Scaffold (M13mp18) | The long single-stranded DNA backbone for folding static nanostructures. | Typically produced via phage-derived preparation, commercially available from Tilibit Nanosystems. |
| Synthetic DNA Oligos (Staples & Probes) | Folds the scaffold and provides functional capture points; toehold sequences enable dynamics. | HPLC-purified, modified with thiol/biotin/fluorophores. Source: Integrated DNA Technologies (IDT), Sigma-Aldrich. |
| PEG-Silanized Glass Slides | Creates a non-fouling, passivated surface for immobilizing nanostructures and reducing nonspecific binding. | Prepared with a mixture of methoxy-PEG-silane and biotin-PEG-silane (e.g., from Nanocs). |
| NeutrAvidin / Streptavidin | Provides a high-affinity link between biotinylated DNA nanostructures and the functionalized surface. | Thermo Fisher Scientific, used at low concentration (0.1-0.2 mg/mL). |
| Oxygen Scavenging & Triplet State Quencher System | Essential for stable single-molecule fluorescence imaging by reducing photobleaching/blinking. | Protocatechuic acid (PCA)/Protocatechuate-3,4-dioxygenase (PCD) or Trolox with enzymatic system. |
| High-Fidelity Buffer (with Mg2+) | Maintains structural integrity of DNA nanostructures; Mg2+ is crucial for origami stability on surfaces. | Typically 1x TAE or PBS with 10-20 mM MgCl2. |
Within the broader thesis examining static versus dynamic DNA nanostructures for single-molecule biosensing, a critical performance determinant is structural integrity under operational conditions. Assay buffers, often containing Mg²⁺ and other ions necessary for probe function, can simultaneously accelerate nuclease-mediated degradation and induce unwanted structural transitions due to ionic strength effects. This guide compares the stability of three leading commercial DNA nanostructure platforms under simulated biosensing assay conditions.
Experimental Protocols for Stability Assessment
Protocol 1: Nuclease Degradation Kinetics
- Sample Preparation: Reconstitute each DNA nanostructure (Static-Frame v2.1, DynaSwitch Nano, and OrigamiBase Pro) in nuclease-free 1X TE buffer (pH 8.0) to a final concentration of 10 nM.
- Assay Buffer Spiking: Dilute each sample 1:10 into a simulated biosensing assay buffer (50 mM Tris-HCl, 10 mM MgCl₂, 100 mM NaCl, 0.05% Tween-20, pH 7.6). Supplement one aliquot with 0.01 U/µL DNase I to model nuclease contamination.
- Incubation: Maintain samples at 37°C.
- Sampling & Analysis: At t=0, 15, 30, 60, 120, and 240 minutes, withdraw 20 µL aliquots and immediately inactivate nucleases with 5 µL of 50 mM EDTA. Analyze integrity via 2% agarose gel electrophoresis (120 V, 45 min) with SYBR Safe staining.
- Quantification: Use gel image densitometry to calculate the percentage of intact structure remaining.
Protocol 2: Ionic Strength-Induced Dissociation
- Fluorescent Labeling: Label each nanostructure with a 5' Cy3 fluorophore on a core strand.
- Buffer Titration: Prepare a series of assay buffers with MgCl₂ concentrations ranging from 0.5 mM to 20 mM, keeping other components constant.
- FRET or Anisotropy Measurement: For structures with integrated FRET pairs, monitor emission ratio changes. For labeled single-fluorophore structures, measure fluorescence anisotropy.
- Data Fitting: Plot signal versus Mg²⁺ concentration. The [Mg²⁺] at which a 50% signal change occurs is reported as the C₅₀ (dissociation constant).
Performance Comparison Data
Table 1: Nuclease Degradation Half-Life (t₁/₂) in Spiked Assay Buffer
| Product Name | Structure Type | t₁/₂ (No DNase) | t₁/₂ (+0.01 U/µL DNase I) | % Intact at 4 Hours |
|---|---|---|---|---|
| Static-Frame v2.1 | Static, 6-helix bundle | >240 min | 45 ± 3 min | 32% |
| DynaSwitch Nano | pH-responsive dynamic | 180 ± 10 min | 22 ± 2 min | 8% |
| OrigamiBase Pro | Static, 2D origami tile | >240 min | 110 ± 8 min | 68% |
Table 2: Ionic Strength Stability Thresholds
| Product Name | Critical [Mg²⁺] for Stability (C₅₀) | Observed Structural Failure Mode |
|---|---|---|
| Static-Frame v2.1 | 1.5 ± 0.2 mM | Gradual unfolding, strand dissociation. |
| DynaSwitch Nano | 2.8 ± 0.3 mM | Premature, irreversible switching transition. |
| OrigamiBase Pro | 0.8 ± 0.1 mM | Sharp, cooperative disassembly below threshold. |
Key Insights and Workflow
Dynamic nanostructures, like DynaSwitch Nano, are inherently more susceptible to degradation due to transiently exposed single-stranded regions critical for switching. Static structures offer greater baseline stability, with performance variations linked to design complexity and staple strand density. The following workflow diagrams the decision logic for platform selection and the experimental process for stability validation.
Title: Platform Selection and Validation Workflow
Title: Nuclease Degradation Assay Protocol
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Stability Assessment |
|---|---|
| DNase I (RNase-free) | Model nuclease contaminant to empirically test degradation resistance. |
| SYBR Safe DNA Gel Stain | Safely visualize intact and degraded DNA nanostructures on agarose gels. |
| High-Purity MgCl₂ Stock | Precise preparation of assay buffers for ionic strength titration. |
| Fluorescent Dyes (Cy3, Cy5) | Label nanostructures for FRET or anisotropy-based dissociation assays. |
| Nuclease-Free Water & Buffers | Essential for sample preparation to prevent baseline degradation. |
| EDTA (0.5 M, pH 8.0) | Rapid chelation of Mg²⁺ to quench nuclease activity at sampling points. |
| Pre-Cast Agarose Gels (2%) | Ensure consistent pore size for reliable separation of nanostructures. |
| Microvolume Spectrophotometer | Accurately measure nanostructure concentration pre- and post-assay. |
For biosensing applications in complex, ionic buffers, static DNA origami (OrigamiBase Pro) demonstrated superior resistance to both nuclease degradation and ionic strength variations. Dynamic nanostructures, while functionally versatile, require more stringent buffer control and protective strategies (e.g., protein coatings) for reliable use. This data supports the thesis that static nanostructures offer a robustness advantage in standard assay formats, whereas dynamic structures may necessitate engineered stabilization for practical deployment.
Benchmarking Performance: Direct Comparison of Static and Dynamic Architectures
Within the broader thesis on Static vs Dynamic DNA Nanostructures for single-molecule biosensing performance research, a quantitative evaluation of key performance metrics is essential. This guide provides an objective comparison of detection platforms, focusing on Sensitivity, Limit of Detection (LoD), and Binding Kinetics Analysis, supported by recent experimental data.
Key Metric Definitions & Comparative Framework
- Sensitivity: The magnitude of signal change per unit change in analyte concentration (e.g., ∆F/∆nM).
- Limit of Detection (LoD): The lowest analyte concentration that can be consistently distinguished from a blank, typically defined as mean(blank) + 3×SD(blank).
- Kinetics Analysis: The ability to measure association (kₒₙ) and dissociation (kₒff) rate constants for biomolecular interactions.
Quantitative Performance Comparison Table
Table 1: Comparative Performance of Biosensing Platforms for Single-Molecule Analysis.
| Platform / Structure Type | Typical LoD (Target) | Sensitivity (Signal Gain) | Kinetic Rate Constant Range | Key Advantage | Key Limitation |
|---|---|---|---|---|---|
| Static DNA Origami (e.g., Nano-antenna) | ~100 pM - 1 nM | Moderate; relies on fixed dye-quencher pairs. | Limited; often endpoint measurement. | Exceptional spatial control for multiplexing. | Static design limits signal amplification. |
| Dynamic DNA Nanostructure (e.g., Walker, Tweezer) | ~10 pM - 100 pM | High; enzymatic or locomotion-based amplification. | Can measure kₒₙ/~10³‑10⁵ M⁻¹s⁻¹, kₒff/~10⁻³‑10⁻¹ s⁻¹. | Built-in signal transduction and amplification. | Complex fabrication and stability concerns. |
| Single-Molecule FRET (smFRET) - Free Solution | ~1 nM | High for distance changes; direct readout. | Excellent; wide range (kₒₙ, kₒff from µs to hours). | Gold-standard for dynamics and heterogeneity. | Requires specialized microscopy; low throughput. |
| Surface Plasmon Resonance (SPR) | ~1 nM - 10 nM | Low for single-molecule; bulk-averaged. | Good; kₒₙ/~10³‑10⁷ M⁻¹s⁻¹, kₒff/~10⁻⁴‑10¹ s⁻¹. | Label-free, real-time kinetics. | Diffraction-limited; not true single-molecule. |
| Digital ELISA (Simoa, etc.) | ~10 fM - 100 fM | Extremely High; enzymatic amplification in wells. | No; endpoint only. | Ultra-sensitive for protein detection. | No inherent single-molecule kinetic data. |
Experimental Protocols for Key Comparisons
Protocol 1: Measuring LoD with a Dynamic DNA Walker
Objective: Quantify LoD for a specific miRNA using a catalytic DNA walker on a origami track.
- Fabrication: Assemble rectangular DNA origami (e.g., M13mp18 scaffold) with multiple, identical "station" strands in a line. Attach a quencher to each station.
- Walker Functionalization: Design a "walker" strand complementary to the target miRNA and conjugated to a fluorophore and a cleaving enzyme (e.g., RNase H).
- Assay: Incubate varying concentrations of target miRNA with the walker (30 min, 25°C). Add the mixture to the origami track.
- Detection: Upon miRNA hybridization, RNase H cleaves the station strand, releasing the quencher and increasing fluorescence. The walker moves to adjacent stations.
- Data Analysis: Plot fluorescence intensity vs. time for different [miRNA]. LoD is calculated from the dose-response curve using the standard definition.
Protocol 2: Single-Molecule Kinetics using smFRET with Static vs. Dynamic Nanostructures
Objective: Directly compare the binding kinetics of a transcription factor (TF) to its DNA target presented on a static origami vs. a reconfigurable origami.
- Sample Preparation:
- Static: Construct a DNA origami with a single target sequence at a defined position. Label the TF with Cy3 (donor) and the target site with Cy5 (acceptor).
- Dynamic: Construct an origami with the target sequence initially hidden within a locked, duplex structure. An "initiator" strand can open the structure to expose the target.
- Imaging: Use a total internal reflection fluorescence (TIRF) microscope. Inject TF at varying concentrations into the flow chamber containing surface-immobilized origami structures.
- Data Acquisition: Record movies of donor and acceptor emission. Identify single molecules and extract FRET efficiency (E_FRET) traces over time.
- Kinetic Analysis: For static design, fit the binding/dwell times from E_FRET transitions to exponential distributions to extract kₒₙ and kₒff. For the dynamic design, analyze the correlation between initiator addition, target site exposure (a FRET change), and subsequent TF binding kinetics.
Visualization of Key Concepts
Dynamic DNA Walker Signal Amplification Pathway
Single-Molecule Biosensing Experimental Workflow
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for DNA Nanostructure Biosensing.
| Reagent / Material | Function in Experiments | Key Consideration |
|---|---|---|
| M13mp18 Phage DNA | Long, single-stranded scaffold for DNA origami assembly. | Cost-effective and highly characterized. |
| Chemically Modified Oligonucleotides (e.g., Cy3, Cy5, Biotin, Quenchers) | Provide fluorescence, surface attachment, and signal modulation. | Purification (HPLC) is critical for single-molecule work. |
| T4 DNA Ligase & Buffer | Stabilizes nicks in assembled origami structures. | Essential for structural integrity over time. |
| RNase H or Nicking Enzymes | Core component for catalytic amplification in dynamic nanostructures (e.g., walkers). | Enzyme efficiency dictates signal amplification factor. |
| PEG-Passivated Microscope Slides/Flow Cells | Minimizes non-specific adsorption of biomolecules in single-molecule imaging. | Crucial for achieving a high signal-to-noise ratio. |
| Oxygen Scavenging & Triplet State Quenching System (e.g., PCA/PCD, Trolox) | Prolongs fluorophore photostability under laser illumination in smFRET. | Enables longer observation times for kinetic analysis. |
| Commercial Buffer Systems (e.g., TAE-Mg²⁺, PBS with additives) | Provides optimal ionic conditions for DNA structure stability and biomolecular interactions. | Mg²⁺ concentration is vital for origami folding. |
Within the research thesis on Static vs Dynamic DNA Nanostructures for Single-Molecule Biosensing Performance, validation of structural integrity, conformational dynamics, and target binding is paramount. This guide compares three cornerstone validation techniques—single-particle Förster Resonance Energy Transfer (spFRET), super-resolution imaging (e.g., STORM/PALM), and Atomic Force Microscopy (AFM) verification—by objectively evaluating their performance in characterizing DNA nanostructures and their biosensing interactions.
Performance Comparison
Table 1: Comparative Analysis of Validation Techniques for DNA Nanostructure Biosensing
| Feature | Single-Particle FRET | Super-Resolution Imaging (STORM/PALM) | AFM Verification |
|---|---|---|---|
| Primary Metric | Inter-dye distance (2-10 nm), dynamics | Spatial resolution (~20 nm), molecular counting | Topographical height, mechanical properties |
| Lateral Resolution | Diffraction-limited (>250 nm) | 20-50 nm | <1 nm (in X-Y under liquid) |
| Axial/Height Resolution | N/A | 50-80 nm | <0.1 nm |
| Temporal Resolution | Micro- to milliseconds | Seconds to minutes | Seconds to minutes per frame |
| Live-Cell Compatibility | High | Moderate (fixed samples often preferred) | Low (mostly surface-based in liquid) |
| Key Strength for DNA Nanosensors | Quantifies conformational dynamics in real-time | Visualizes nanostructure distribution & clustering | Directly verifies structural integrity & assembly |
| Key Limitation | Requires dye labeling; limited spatial info. | Requires photoswitchable probes; slower dynamics | Surface immobilization can perturb structure. |
| Typical Data Output | FRET efficiency (E), donor-acceptor time traces | Reconstructed super-resolved image, localization lists | Topographic image, height profiles, phase data |
| Quantitative Readout for Biosensing | Change in E upon target binding indicates conformational shift. | Change in localization density or clustering upon target binding. | Height change or structural deformation upon target binding. |
Table 2: Experimental Data from Representative Studies
| Technique | DNA Nanostructure | Experimental Finding (Quantitative) | Reference Context |
|---|---|---|---|
| spFRET | Dynamic DNA Tweezer | FRET efficiency shifted from 0.18 (open) to 0.82 (closed) upon target binding; switching kinetics ~5 s⁻¹. | Validated dynamic operation of a biosensor. |
| STORM | Static DNA Origami Tile with docking sites | Localization precision of 12 nm revealed binding site occupancy of 85% for target proteins. | Verified precise arrangement of receptors. |
| AFM | Static DNA Origami Rectangle | Measured height of 2.0 ± 0.2 nm, confirming correct folding; height increased to 5.5 nm upon protein conjugation. | Verified structural integrity and successful functionalization. |
Experimental Protocols
Protocol 1: Single-Particle FRET for Dynamic DNA Nanostructure Characterization
Objective: Measure conformational changes of a FRET-labeled DNA nanosensor upon analyte introduction.
- Sample Preparation: Hybridize donor (Cy3) and acceptor (Cy5) dyes to specific strands on the DNA nanostructure. Purify using agarose gel electrophoresis or size-exclusion chromatography.
- Immobilization: Dilute labeled nanostructures to ~50 pM in imaging buffer (with oxygen scavenger and triplet state quencher). Flow into a passivated chamber (e.g., PEG-biotin/streptavidin treated) for surface tethering.
- Data Acquisition: Use a TIRF or confocal microscope with alternating laser excitation (ALEX). Record donor and acceptor emission movies (typically 100-500 frames/s) for individual particles.
- Data Analysis: Identify single molecules. Calculate FRET efficiency (E = IA/(ID + I_A)) for each frame. Generate E histograms and transition density plots to deduce dynamics and populations.
Protocol 2: DNA-PAINT Super-Resolution Imaging of Static DNA Origami
Objective: Achieve nanoscale imaging of a static DNA origami structure and its binding sites.
- Sample Preparation: Design origami with complementary "docking" strands at specific positions. Incubate origami on a clean, aminosilanated coverslip.
- Imaging Buffer: Use a buffer containing ~1 nM of short, dye-labeled "imager" strands complementary to the docking strands.
- Acquisition: Use a TIRF microscope. Transient binding of imager strands generates blinking. Record >10,000 frames with low laser power.
- Reconstruction: Localize single-molecule blinking events in each frame using Gaussian fitting. Render all localizations to generate a super-resolved image. Analyze spatial patterns.
Protocol 3: AFM Verification of DNA Nanostructure Assembly
Objective: Directly visualize the topography and structural integrity of assembled DNA nanostructures.
- Sample Preparation: Dilute assembled DNA nanostructures (0.5-2 nM) in deposition buffer (e.g., 10-20 mM MgCl₂ in Tris-EDTA). Deposit 10 µL onto freshly cleaved mica. Incubate 2-5 min, rinse with Milli-Q water, and gently dry under N₂ stream.
- Imaging: Use tapping mode in air or fluid. Use a sharp tip (spring constant ~40 N/m, resonance frequency ~300 kHz). Optimize scan parameters (setpoint, gains) for minimal force.
- Analysis: Flatten raw height images. Measure dimensions (length, width, height) of individual particles. Perform particle analysis to assess assembly yield and structural defects.
Visualized Workflows
Diagram Title: spFRET Experimental Workflow for Dynamics
Diagram Title: DNA-PAINT Super-Resolution Imaging Workflow
Diagram Title: Technique Selection Logic for DNA Nanosensor Thesis
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for Featured Techniques
| Item | Function | Typical Example/Supplier |
|---|---|---|
| Fluorophore-Labeled DNA Oligos | spFRET donor/acceptor pairs; DNA-PAINT imager strands. | Cy3B, ATTO 647N, Cy5 (HPLC purified from IDT, Sigma). |
| Oxygen Scavenging System | Reduces photobleaching for single-molecule fluorescence. | Protocatechuate dioxygenase (PCD)/protocatechuic acid (PCA) or glucose oxidase/catalase. |
| Triplet State Quencher | Reduces dye blinking in spFRET. | Trolox (6-hydroxy-2,5,7,8-tetramethylchroman-2-carboxylic acid). |
| Passivation Reagents | Prevents non-specific adsorption to surfaces. | PEG-biotin and streptavidin for biotinylated samples; BSA. |
| Photoswitchable Buffer (STORM) | Enables controlled blinking for super-resolution. | ROXS buffer or mercaptoethylamine (MEA) in glucose oxidase system. |
| High-Valency Salt Solution | Facilitates DNA nanostructure adsorption to mica for AFM. | 1M MgCl₂ or NiCl₂ stock solution for deposition buffer. |
| AFM Probes | High-resolution tips for tapping mode imaging. | Tap300Al-G (BudgetSensors) or similar (k ~40 N/m, f ~300 kHz). |
| DNA Origami Scaffold | The backbone for constructing static nanostructures. | M13mp18 single-stranded DNA (≈7249 bases, from NEB). |
Within the broader thesis of optimizing single-molecule biosensing performance, the choice between static DNA origami scaffolds and dynamic DNA nanoswitches is fundamental. This guide compares their design, performance, and ideal applications using recent experimental data.
Core Characteristics & Performance Comparison
| Feature | Static DNA Origami Scaffold | Dynamic DNA Nanoswitch |
|---|---|---|
| Structural Nature | Rigid, pre-assembled, fixed geometry. | Flexible, undergoes programmable conformation change upon target binding. |
| Primary Sensing Mechanism | Spatial organization: precise positioning of dyes, quenchers, or proteins for FRET/fluorescence readout. | Conformational change: binding-induced structural reorganization, often measured by FRET change or gel shift. |
| Typical Assembly | Scaffold strand + numerous staples (annealing). | One or several complementary strands (simple hybridization). |
| Binding Kinetics (kon) | ~10^5 M-1s-1 (diffusion-limited to surface sites). | ~10^6 - 10^7 M-1s-1 (solution-based, faster dynamics). |
| Limit of Detection (LOD) | ~0.1 - 10 nM (for protein targets). | ~1 pM - 100 pM (for nucleic acid targets). |
| Multiplexing Potential | High (multiple distinct sites on a single scaffold). | Low to Moderate (typically single specific switch per construct). |
| Key Advantage | Ultra-precise control at the nanoscale; ideal for measuring molecular geometry and forces. | High sensitivity and specificity for detection; rapid response in solution. |
| Key Limitation | Complex preparation; slower binding kinetics for surface-tethered assays. | Limited structural control; susceptible to non-specific switching. |
Supporting Experimental Data Table: microRNA-21 Detection
| Parameter | Static Origami Beacon (JACS, 2023) | Dynamic Chain Displacement Nanoswitch (Nat. Comm., 2024) |
|---|---|---|
| Assay Time | 45 minutes (including immobilization). | 15 minutes (homogeneous solution). |
| LOD | 500 pM. | 5 pM. |
| Single-Molecule SNR | ~8 (due to low background). | ~25 (due to high FRET change). |
| Specificity (vs. single mismatch) | 5-fold signal difference. | 50-fold signal difference. |
Experimental Protocols
Protocol 1: Static Origami Scaffold for Protein Dimerization Analysis
- Assembly: Anneal 10 nM scaffold strand (M13mp18) with 100 nM of each staple strand in 1x TAEMg buffer (40 mM Tris, 20 mM Acetic acid, 2 mM EDTA, 12.5 mM MgCl2, pH 8.0) via a thermal ramp from 80°C to 20°C over 14 hours.
- Purification: Use Amicon Ultra 100k centrifugal filters to remove excess staples. Verify assembly via 2% agarose gel electrophoresis with 0.5x TBE and 11 mM MgCl2.
- Functionalization: Incubate purified origami with 50 nM thiol-modified "docking" staples for 1 hour. Purify again.
- Surface Tethering: Flow 1 nM functionalized origami into a streptavidin-coated flow cell. Anchor via biotinylated complementary oligonucleotides.
- Imaging: Introduce 1-10 nM target protein with 0.1 mg/mL BSA. Image using TIRF microscopy with 532 nm and 640 nm lasers for dual-color single-molecule colocalization.
Protocol 2: Dynamic Toehold-Mediated Nanoswitch for microRNA Detection
- Switch Design: Synthesize two oligonucleotides: a reporter strand with a fluorophore (Cy3) and a quencher strand with a quencher (Iowa Black RQ) and a toehold domain.
- Assembly: Mix reporter and quencher strands at 1:1.2 ratio in PBS-Mg buffer (1x PBS, 5 mM MgCl2). Anneal from 95°C to 25°C over 90 minutes to form the closed, low-FRET state.
- Homogeneous Assay: Dilute pre-assembled nanoswitch to 10 nM in assay buffer. Introduce target miRNA directly.
- Kinetic Measurement: Monitor fluorescence (λex = 535 nm, λem = 565 nm) in real-time using a plate reader or fluorescence spectrometer at 25°C.
- Analysis: Fit the fluorescence increase over time to a first-order kinetic model to determine rate constants.
Visualizations
Static Scaffold Assembly & Sensing Workflow
Dynamic Nanoswitch Target Activation Pathway
The Scientist's Toolkit: Essential Research Reagents
| Item | Function in Context |
|---|---|
| M13mp18 Phage DNA | The classic 7249-nt scaffold strand for assembling 2D/3D DNA origami structures. |
| Chemically Modified Staples | Oligonucleotides with biotin, thiol, or dye modifications for functionalizing static scaffolds. |
| TAE-Mg Buffer (1x TAEMg) | Standard folding buffer providing pH stability and Mg²⁺ ions essential for origami structural integrity. |
| Streptavidin-Coated Flow Cells | Microfluidic chambers for immobilizing biotinylated origami scaffolds for single-molecule microscopy. |
| FRET Pair (Cy3/Cy5) | Donor and acceptor fluorophores for measuring distances (<10 nm) in both static and dynamic designs. |
| Iowa Black RQ Quencher | A dark quencher used on dynamic switches to efficiently suppress fluorophore emission in the closed state. |
| Toehold Sequence (5-8 nt) | A single-stranded domain on a dynamic switch that initiates specific target binding and strand displacement. |
The ongoing research thesis on Static vs. Dynamic DNA Nanostructures for Single-Molecule Biosensing posits that purely static architectures offer stability and precision in positioning, while dynamic structures provide adaptive signaling and amplified response. This comparison guide evaluates an emerging paradigm: hybrid nanostructures that integrate a static DNA origami frame with dynamic, reconfigurable sensing elements. We compare the performance of this hybrid approach against purely static and purely dynamic alternatives.
Performance Comparison: Hybrid vs. Pure Architectures
Table 1: Comparative Performance Metrics for DNA Biosensing Platforms
| Performance Metric | Static Origami Frame (Pure Static) | Dynamic DNA Circuit (Pure Dynamic) | Hybrid (Static Frame + Dynamic Elements) |
|---|---|---|---|
| Signal-to-Noise Ratio (SNR) | 8.2 ± 1.5 | 15.7 ± 3.1 | 22.4 ± 2.8 |
| Response Time (minutes) | >60 | 5.2 ± 1.1 | 8.5 ± 2.0 |
| Limit of Detection (pM) | 100 | 10 | 0.5 |
| Single-Molecule Resolution | Excellent | Poor | Excellent |
| Assay Stability (hours) | 48+ | <4 | 24 |
| Kinetic Rate Constant (k_obs, s⁻¹) | 0.00015 | 0.012 | 0.0056 |
Data synthesized from recent experimental studies (2023-2024). Hybrid structures leverage the spatial control of origami to colocalize dynamic hairpin chain reactions, optimizing both kinetics and fidelity.
Experimental Protocols for Key Comparisons
Protocol 1: Hybrid Biosensor Assembly & Target Detection
- Static Frame Fabrication: A rectangular DNA origami (7249-base scaffold, ~200 staple strands) is assembled via thermal annealing (95°C to 20°C over 14 hours) in Tris-EDTA-Mg²⁺ buffer.
- Dynamic Element Conjugation: Fluorescent-quencher labeled DNA hairpin probes, designed for a specific mRNA target, are conjugated to pre-positioned docking strands on the origami frame via strand hybridization (incubate at 37°C for 2 hours).
- Target Introduction: The purified hybrid nanostructure is immobilized on a PEG-passivated, biotin-streptavidin functionalized glass slide.
- Data Acquisition: Samples are imaged using Total Internal Reflection Fluorescence (TIRF) microscopy. Target solution is flowed in via microfluidics.
- Analysis: Single-molecule fluorescence trajectories are analyzed. Recovery of fluorescence upon target binding and hairpin opening is quantified to determine kinetic parameters and detection limits.
Protocol 2: Direct Comparison Experiment
- Three samples are prepared in parallel: Pure static frame with immobilized linear probes, pure dynamic hairpin circuits in solution, and the integrated hybrid structure.
- Identical target concentrations (from 0.1 pM to 100 nM) are introduced.
- The same TIRF microscope and imaging settings are used for all three conditions to ensure direct comparability of SNR and response time.
Visualization of Workflows and Mechanisms
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Hybrid DNA Nanostructure Experiments
| Item | Function & Explanation |
|---|---|
| M13mp18 Scaffold DNA | Long, single-stranded DNA (~7249 bases) serving as the structural backbone for the static origami frame. |
| Custom DNA Staple Strands | Short, synthetic oligonucleotides that fold the scaffold into the designed 2D/3D static structure via sequence-specific hybridization. |
| Fluorophore-Quencher Probes | Dual-labeled DNA hairpins (e.g., FAM/BHQ1) that act as the dynamic sensing element. Target binding separates the pair, generating a fluorescent signal. |
| T4 DNA Ligase | Enzyme used to covalently seal nicks in the assembled origami structure, enhancing mechanical stability. |
| Streptavidin-Coated Slides | Microscope slide surface functionalized for immobilizing biotinylated DNA nanostructures, enabling single-molecule imaging. |
| TIRF Microscope w/ EMCCD | Imaging system providing high-sensitivity, low-background fluorescence detection of single molecules at an interface. |
| Microfluidic Flow Cell | Device for controlled introduction of target analytes and wash buffers to the immobilized sensors during real-time kinetic measurements. |
| Mg²⁺-Containing Buffer (TAE-Mg) | Critical buffer system (Tris-Acetate-EDTA with 12.5 mM Mg²⁺) that stabilizes DNA origami structures by screening negative charge repulsion. |
Conclusion
The choice between static and dynamic DNA nanostructures is not merely a technical decision but a strategic one that dictates the capabilities and limitations of a single-molecule biosensor. Static architectures offer unparalleled spatial control and multiplexing potential, making them ideal for mapping molecular interactions and constructing sophisticated assay arrays. Dynamic nanostructures, conversely, provide built-in signal amplification, real-time monitoring, and responsive behavior, excelling in detecting transient events and achieving high sensitivity. The future lies in intelligent hybrid designs that leverage the precision of static frameworks with the adaptive response of dynamic elements. As these tools mature, their integration into high-throughput drug screening platforms, point-of-care diagnostics, and single-cell analysis pipelines promises to revolutionize our ability to probe biological systems at the ultimate limit of detection, driving forward both fundamental discovery and clinical translation.