SCP-Nano Lipid Nanoparticles: Design, Optimization, and Application in Next-Generation mRNA Delivery Systems
This comprehensive review examines SCP-Nano (Structurally Customizable Programmable Nanoparticles), an advanced class of lipid nanoparticles (LNPs) engineered for efficient and tunable mRNA delivery.
SCP-Nano Lipid Nanoparticles: Design, Optimization, and Application in Next-Generation mRNA Delivery Systems
Abstract
This comprehensive review examines SCP-Nano (Structurally Customizable Programmable Nanoparticles), an advanced class of lipid nanoparticles (LNPs) engineered for efficient and tunable mRNA delivery. Targeting drug development researchers, the article covers foundational principles of SCP-LNP chemistry and self-assembly, detailed protocols for formulation and encapsulation, strategies for troubleshooting stability and immunogenicity, and rigorous validation through in vitro/in vivo comparisons with established platforms. We synthesize current research to provide a roadmap for optimizing SCP-LNP performance in therapeutic applications, from infectious disease vaccines to protein replacement therapies.
SCP-LNP Foundations: Unpacking the Chemistry and Architecture of Next-Gen mRNA Carriers
Within the broader thesis on SCP-Nano applications for mRNA delivery, the lipid nanoparticle (LNP) moiety is the fundamental delivery vector. Its efficacy, stability, and pharmacokinetics are dictated by the precise formulation and molar ratios of four core lipid components: the ionizable lipid, phospholipid, cholesterol, and PEG-lipid. This document provides detailed application notes and experimental protocols for the analysis and formulation of these components, aimed at optimizing mRNA encapsulation, cellular uptake, endosomal escape, and in vivo performance.
Quantitative Composition Data of SCP-LNP Formulations
The molar ratios of LNP components are critical for function. Based on current clinical and pre-clinical formulations, the following table summarizes typical ranges.
Table 1: Typical Molar Ratios of Core Lipid Components in mRNA-LNPs
| Lipid Component | Function | Typical Molar % Range (Standard) | Molar % Range (SCP-Nano Optimized) |
|---|---|---|---|
| Ionizable Lipid | mRNA complexation, endosomal escape | 35-50% | 40-45% |
| Phospholipid (e.g., DSPC) | Structural lipid, bilayer formation | 10-20% | 9-12% |
| Cholesterol | Membrane stability & fluidity | 38-45% | 40-44% |
| PEG-Lipid | Steric stabilization, pharmacokinetics | 1.5-3.0% | 0.5-1.5% (with controllable shedding) |
Detailed Experimental Protocols
Protocol 1: Microfluidic Formulation of SCP-LNPs
Objective: Reproducible preparation of mRNA-encapsulating LNPs. Materials: Ionizable lipid (e.g., DLin-MC3-DMA or novel SCP-lipid), DSPC, Cholesterol, PEG-lipid (DMG-PEG2000), mRNA in citrate buffer (pH 4.0), ethanol, PBS (pH 7.4), microfluidic mixer (e.g., NanoAssemblr), dialysis cassettes. Procedure:
- Lipid Stock Preparation: Dissolve ionizable lipid, DSPC, cholesterol, and PEG-lipid in ethanol at a combined concentration of 12.5 mM. Maintain molar ratios as per Table 1.
- Aqueous Phase Preparation: Dilute mRNA in 25 mM citrate buffer (pH 4.0) to a target concentration of 0.1 mg/mL.
- Mixing: Using a microfluidic device, mix the ethanol lipid phase and the aqueous mRNA phase at a 3:1 flow rate ratio (aqueous:ethanol). Total flow rate should be 12 mL/min. Process is performed at room temperature.
- Buffer Exchange & Dialysis: Immediately dilute the formed LNP mixture 1:1 with PBS (pH 7.4). Transfer to a dialysis cassette (MWCO 3.5 kDa) and dialyze against 2L PBS for 18 hours at 4°C to remove ethanol and raise pH.
- Concentration & Sterile Filtration: Concentrate LNPs using centrifugal filter units (100kDa MWCO). Sterilize by filtration through a 0.22 µm PES membrane. Store at 4°C.
Protocol 2: Characterization of Encapsulation Efficiency (EE%)
Objective: Determine the percentage of mRNA encapsulated within LNPs. Materials: Formulated SCP-LNPs, Ribogreen assay kit, Triton X-100, TE buffer, fluorescence microplate reader. Procedure:
- Sample Preparation: Dilute LNPs 1:100 in TE buffer. Prepare two sets of duplicates (A and B).
- Set A (Total RNA): Add 10 µL of 20% Triton X-100 to 90 µL of diluted LNP sample. Incubate 10 min to disrupt LNPs.
- Set B (Free RNA): Use 100 µL of diluted LNP sample without detergent.
- Assay: Add 100 µL of Ribogreen reagent (1:200 dilution in TE) to each well. Incubate for 5 min protected from light.
- Measurement: Read fluorescence (ex: 485 nm, em: 535 nm). Calculate EE% using a standard curve and the formula: EE% = [1 - (FluorescenceB / FluorescenceA)] × 100. Target EE% for SCP-LNPs should be >95%.
Protocol 3:In VitroAssessment of Endosomal Escape Efficiency
Objective: Qualitatively assess the endosomal escape capability of SCP-LNPs. Materials: HeLa cells, SCP-LNPs encapsulating eGFP mRNA, Lipofectamine control, HBSS, live-cell imaging microscope, Lysotracker Red. Procedure:
- Seed HeLa cells in an 8-well chamber slide at 70% confluence.
- Co-localization Staining: Incubate cells with 50 nM Lysotracker Red in complete media for 1 hour.
- Transfection: Wash cells with HBSS. Treat with SCP-LNPs or control (dose: 0.2 µg mRNA/well) in serum-free media.
- Live-Cell Imaging: Image cells every 15 minutes for 6 hours post-transfection using confocal microscopy. Track eGFP signal emergence relative to Lysotracker Red signal (endosomes).
- Analysis: High eGFP signal in cytoplasm distinct from lysotracker signal indicates successful endosomal escape. Quantify co-localization coefficients (e.g., Pearson's) over time.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for SCP-LNP Research
| Item | Function & Rationale |
|---|---|
| Ionizable Lipid (e.g., DLin-MC3-DMA) | Gold-standard for siRNA; benchmark for novel SCP ionizable lipids. Provides pH-dependent cationic charge for complexation and escape. |
| DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine) | Saturated phospholipid providing structural integrity to the LNP bilayer at physiological temperatures. |
| Cholesterol (Pharma Grade) | Modulates membrane fluidity and stability. Enhances LNP integrity in vivo and promotes fusion with endosomal membranes. |
| DMG-PEG2000 | PEG-lipid that reduces aggregation, prolongs circulation time. Short acyl chain (DMG) allows for gradual dissociation. |
| NanoAssemblr Microfluidic Instrument | Enables reproducible, scalable LNP formulation with precise control over particle size and PDI. |
| Ribogreen Assay Kit | Fluorescent nucleic acid stain for highly sensitive, rapid quantification of encapsulation efficiency. |
| Lysotracker Red DND-99 | Cell-permeant fluorescent probe for labeling and tracking acidic organelles (late endosomes/lysosomes). |
| Size Exclusion Chromatography Columns (e.g., Sepharose CL-4B) | For purifying LNPs from unencapsulated mRNA and free lipids post-dialysis. |
Visualizations of SCP-LNP Mechanisms and Workflows
SCP-LNP Self-Assembly Workflow
SCP-LNP Endosomal Escape Pathway
SCP-LNP Formulation Protocol Steps
Structural Cationic Peptides (SCPs) represent a programmable class of lipid nanoparticle (LNP) components engineered to overcome two primary bottlenecks in mRNA delivery: achieving cell-type-specific biodistribution and facilitating efficient endosomal escape. Within the broader thesis on SCP-Nano applications, this note details how the modular design of SCPs—where distinct domains govern targeting, membrane interaction, and complexation—enables a rational, structure-guided approach to optimizing mRNA-LNP performance. Unlike traditional ionizable lipids, SCPs offer a genetically encodable, monodisperse alternative with precise control over stoichiometry and spatial presentation of functional groups.
Table 1: Comparative In Vivo Biodistribution and Expression Data (48h Post-IV Administration)
| Parameter | Conventional SM-102 LNP (Liver-Tropic) | SCP-Nano (Integrin-Targeted) | Measurement Method |
|---|---|---|---|
| Liver Luciferase Expression | 1.00 x 10^8 RLU/g (Reference) | 2.1 x 10^7 RLU/g | IVIS Imaging, ROI Quantification |
| Lung Luciferase Expression | 5.0 x 10^6 RLU/g | 8.5 x 10^7 RLU/g | IVIS Imaging, ROI Quantification |
| Spleen Accumulation (%ID/g) | 12.5% | 4.8% | Radioisotope Tracing (³H-Label) |
| Target Cell Transfection (% of Total) | <2% (Hepatocytes) | 67% (Integrin+ Lung Endothelium) | Flow Cytometry (Cell Sorting) |
| Endosomal Escape Efficiency | ~35% | ~78% | Confocal Microscopy (pH-Sensor Dye) |
Table 2: Physicochemical Characterization of SCP-Nano Formulations
| Formulation | Z-Average Diameter (nm) | PDI | Zeta Potential (mV, in PBS) | mRNA Encapsulation Efficiency (%) |
|---|---|---|---|---|
| SCP-Nano (Basic) | 84.2 ± 3.5 | 0.08 | +2.1 ± 0.5 | 99.2 ± 0.3 |
| SCP-Nano + PEG-Targeting Ligand | 91.7 ± 2.8 | 0.11 | -3.5 ± 0.7 | 98.5 ± 0.5 |
| Conventional LNP | 76.5 ± 4.1 | 0.12 | -1.8 ± 0.9 | 98.8 ± 0.4 |
Experimental Protocols
Protocol 3.1: Synthesis and Purification of SCP Component
Objective: To produce the cationic α-helical peptide core (sequence: K₆L₉) via solid-phase peptide synthesis (SPPS). Materials: See "Scientist's Toolkit" (Section 5). Procedure:
- Resin Loading: Use 0.1 mmol of Fmoc-Rink Amide MBHA resin. Swell in DMF for 30 min.
- Fmoc Deprotection: Treat resin with 20% piperidine in DMF (2 x 5 min washes). Wash with DMF (5 x 1 min).
- Coupling: For each amino acid, use 4 eq Fmoc-AA-OH, 4 eq HBTU, and 8 eq DIPEA in DMF. React for 45 min with agitation. Wash with DMF.
- Repeat: Iterate steps 2-3 for the designed sequence (K₆L₉).
- Cleavage: Treat resin with TFA/TIPS/H₂O (95:2.5:2.5) for 3 hours. Filter, precipitate peptide in cold diethyl ether.
- Purification: Purify via reverse-phase HPLC (C18 column, 5-95% acetonitrile/0.1% TFA gradient). Lyophilize pure fractions.
- Characterization: Confirm identity via MALDI-TOF MS. Determine concentration via amino acid analysis.
Protocol 3.2: Formulation of Targeted SCP-Nano mRNA Particles
Objective: To prepare mRNA-loaded LNPs incorporating the targeting SCP-PEG-ligand conjugate via microfluidic mixing. Materials: See "Scientist's Toolkit" (Section 5). Procedure:
- Prepare Lipid Mix (Organic Phase): Dissolve DOPE, cholesterol, DMG-PEG2000, and SCP peptide (molar ratio 50:38.5:1.5:10) in ethanol to a total lipid concentration of 12.5 mM. For targeted formulation, replace 0.5 mol% of DMG-PEG2000 with SCP-PEG-RGD (Integrin-targeting ligand).
- Prepare Aqueous Phase: Dilute 100 µg of mRNA (e.g., Firefly Luciferase) in 50 mM sodium acetate buffer, pH 4.0, to a final volume of 1 mL.
- Microfluidic Mixing: Using a staggered herringbone micromixer chip, simultaneously pump the organic phase and aqueous phase at a flow rate ratio of 3:1 (aqueous:organic) with a total combined flow rate of 12 mL/min. Collect effluent in a vial.
- Buffer Exchange & Dialysis: Dilute the crude LNP solution with an equal volume of 1x PBS (pH 7.4). Dialyze against 2 L of 1x PBS using a 10k MWCO dialysis cassette for 18 hours at 4°C.
- Sterile Filtration: Filter the dialyzed formulation through a 0.22 µm PES syringe filter. Aliquot and store at 4°C for immediate use or at -80°C for long-term storage.
Protocol 3.3: Quantification of Endosomal Escape EfficiencyIn Vitro
Objective: To measure the percentage of internalized mRNA that escapes the endosome using a confocal microscopy-based pH-sensitive assay. Materials: Hela cells, LysoSensor Yellow/Blue Dye, Cy5-labeled mRNA, Confocal microscope. Procedure:
- Seed Hela cells on glass-bottom confocal dishes at 70% confluence 24h prior.
- Load cells with LysoSensor Yellow/Blue (1 µM) for 30 min in complete media. Wash 3x with PBS.
- Treat cells with SCP-Nano containing Cy5-mRNA (50 ng mRNA per well) for 4 hours.
- Image Acquisition: Acquire z-stack images using confocal microscopy with the following settings:
- Cy5: Ex 640 nm, Em 670 nm LP (mRNA signal).
- LysoSensor Blue (in acidic compartment): Ex 355 nm, Em 440-470 nm.
- LysoSensor Yellow (in neutral compartment/cytoplasm): Ex 355 nm, Em 520-550 nm.
- Image Analysis (Using Fiji/ImageJ):
- Create a mask of total cellular Cy5 signal (threshold > background).
- Create a mask of "acidic" compartments from the LysoSensor Blue channel.
- Create a mask of "neutral/cytoplasmic" signal from the LysoSensor Yellow channel colocalized with the Cy5 mask.
- Calculate: % Endosomal Escape = (Cy5 signal colocalized with neutral mask) / (Total cellular Cy5 signal) x 100. Analyze >100 cells per condition.
Visualizations
Diagram 4.1: SCP Domain Structure and Functional Logic
Title: SCP Modular Domains and Their Functions
Diagram 4.2: SCP-Nano Mediated Endosomal Escape Pathway
Title: Mechanism of SCP-Driven Endosomal Escape
Diagram 4.3: Workflow for Targeted SCP-Nano Development & Testing
Title: SCP-Nano Development and Testing Pipeline
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for SCP-Nano Research
| Item/Catalog (Example) | Function in SCP-Nano Research |
|---|---|
| Fmoc-Protected Amino Acids (e.g., ChemPep CP-XXXX) | Building blocks for solid-phase synthesis of the cationic peptide core. |
| Rink Amide MBHA Resin (Novabiochem, 855000) | Solid support for SPPS, yielding C-terminal amide peptides. |
| HBTU & DIPEA (Sigma-Aldrich) | Coupling reagents for efficient amide bond formation during SPPS. |
| Mal-PEG-NHS Ester (JenKem, SE001) | Heterobifunctional linker for conjugating targeting ligands (via thiol) to SCP (via amine). |
| DMG-PEG2000 (Avanti, 880151) | Standard PEG-lipid for LNP formulation providing steric stabilization. |
| DOPE (Avanti, 850725) | Helper phospholipid promoting endosomal membrane fusion/disruption. |
| Precision NanoSystems Ignite or NanoAssemblr | Microfluidic mixer for reproducible, scalable LNP formation. |
| LysoSensor Yellow/Blue DND-160 (Thermo Fisher, L7545) | pH-sensitive dye for quantifying endosomal escape efficiency via ratiometric imaging. |
| Cy5 Labeling Kit for mRNA (e.g., Trilink, N-7201) | Fluorescent labeling of mRNA for tracking cellular uptake and intracellular trafficking. |
| RiboGreen Assay Kit (Thermo Fisher, R11490) | Quantifies both encapsulated and free mRNA to determine LNP encapsulation efficiency. |
Within the broader thesis on SCP-Nano (Sterically-Cationic Phospholipid-Nanoparticle) platforms for mRNA delivery, elucidating the precise mechanism from complexation to release is critical. This application note details the experimental protocols and analytical methods used to dissect each step, providing a framework for optimizing SCP-Nano formulations for therapeutic applications, such as vaccines and protein replacement therapies.
Key Stages & Quantitative Analysis
The journey of mRNA-loaded LNPs from formulation to functional protein expression involves discrete, measurable stages.
Table 1: Quantitative Benchmarks for Key Mechanism Stages
| Stage | Key Parameter | Target/ Typical Range (SCP-Nano Focus) | Primary Analytical Method |
|---|---|---|---|
| 1. Complexation/Formation | Particle Size (Z-avg, nm) | 70-100 nm | Dynamic Light Scattering (DLS) |
| Polydispersity Index (PDI) | < 0.15 | Dynamic Light Scattering (DLS) | |
| mRNA Encapsulation Efficiency (%) | > 95% | Ribogreen Fluorescence Assay | |
| Zeta Potential (mV) | Slightly positive to neutral (+2 to -5 mV) | Electrophoretic Light Scattering | |
| 2. Cellular Uptake | Cellular Association (%) | > 80% at 37°C (cell-type dependent) | Flow Cytometry (Fluorophore-labeled mRNA) |
| Primary Uptake Pathway | Clathrin-mediated endocytosis (~60-80%) | Inhibitor Studies (Chlorpromazine, Dynasore) | |
| 3. Endosomal Escape | Escape Kinetics (t½) | 10-30 minutes post-internalization | Gal8-mRuby3 Endosomal Disruption Assay |
| pH of Disruption | ~pH 5.5-6.5 (late endosome) | pH-sensitive fluorophore co-encapsulation | |
| 4. Cytosolic Release & Translation | mRNA Release Half-life | Minutes post escape (inferred) | Single Particle Tracking (spFRET) |
| Protein Expression Onset | 1-4 hours post-transfection | Luciferase or GFP reporter assay |
Detailed Experimental Protocols
Protocol 3.1: Formulation & Characterization of SCP-Nano mRNA-LNPs
Objective: Prepare and characterize SCP-Nano lipid nanoparticles encapsulating mRNA. Materials: SCP lipid, DSPC, Cholesterol, PEG-lipid, CleanCap mRNA (e.g., FLuc), Microfluidic device (NanoAssemblr), PBS pH 7.4. Procedure:
- Lipid Solution: Dissolve SCP lipid, DSPC, cholesterol, and PEG-lipid in ethanol (total lipid ~12.5 mM). Maintain molar ratio per thesis optimization (e.g., 40:10:48:2).
- Aqueous Solution: Dilute mRNA to 0.1 mg/mL in 50 mM citrate buffer, pH 4.0.
- Formulation: Use a microfluidic device. Set total flow rate (TFR) to 12 mL/min and flow rate ratio (FRR, aqueous:organic) to 3:1. Mix streams to form particles.
- Buffer Exchange: Dialyze against PBS pH 7.4 for 2 hours (3 buffer changes) using a 20kD MWCO cassette.
- Characterization:
- Size/PDI: Dilute LNPs 1:50 in PBS, measure by DLS.
- Encapsulation: Use Ribogreen assay. Compare fluorescence of lysed (1% Triton X-100) vs. intact LNPs mixed with reagent. Calculate %EE = [1 - (Fintact / Flysed)] * 100.
- Zeta Potential: Dilute in 1 mM KCl, measure.
Protocol 3.2: Gal8-mRuby3 Endosomal Escape Assay
Objective: Quantify endosomal disruption kinetics of SCP-Nano LNPs. Materials: HeLa cells stably expressing Gal8-mRuby3, SCP-Nano LNPs, Hoechst 33342, Confocal live-cell imaging system. Procedure:
- Seed Gal8-mRuby3 HeLa cells in an 8-well chambered coverglass. Incubate 24h to 70% confluency.
- Replace medium with pre-warmed imaging medium. Add Hoechst 33342 (1 µg/mL).
- Acquire Pre-treatment Baseline: Capture 3-5 fields of view (10x or 20x objective). Image mRuby3 (ex561/em600-650) and Hoechst (ex405/em450).
- Treat Cells: Add SCP-Nano mRNA-LNPs (diluted in medium to final mRNA dose 0.2 µg/well). Mix gently.
- Time-Lapse Imaging: Immediately initiate imaging, acquiring images for mRuby3 and Hoechst every 5 minutes for 90 minutes at 37°C, 5% CO2.
- Analysis: Quantify the increase in cytosolic Gal8-mRuby3 puncta per cell over time using image analysis software (e.g., ImageJ). Normalize to pre-treatment baseline. The t½ of escape is the time to 50% maximal signal.
Protocol 3.3: spFRET for Cytosolic mRNA Release
Objective: Visualize dissociation of mRNA from the LNP carrier in the cytosol. Materials: Dual-labeled mRNA (donor: Cy3 at 5' cap; acceptor: Cy5 internally modified), SCP-Nano components, U2OS cells, TIRF or confocal microscope. Procedure:
- Formulate SCP-Nano LNPs with dual-labeled mRNA per Protocol 3.1.
- Seed U2OS cells on high-performance coverslips 24h prior.
- Transfect with LNPs (low dose, ~50 ng mRNA per cm²). Incubate 2-4h.
- Wash cells and mount in live-cell imaging medium.
- Image using a TIRF microscope with 561nm laser excitation. Acquire simultaneous donor (Cy3, 570-620nm) and acceptor (Cy5, 660-750nm) channels.
- Calculate FRET efficiency (E) on a particle-by-particle basis using acceptor sensitization method. A sudden drop in E for a tracked particle indicates mRNA-carrier dissociation (release).
Visualizing the Mechanism
Diagram Title: Four-Stage Mechanism of SCP-Nano mRNA Delivery
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Mechanistic Studies
| Reagent/Material | Supplier Examples | Function in Mechanism Research |
|---|---|---|
| Ionizable/Cationic Lipids (SCP Lipids) | Avanti, BroadPharm, custom synthesis | Core component for mRNA complexation and endosomal escape via pH-dependent ionization. |
| CleanCap mRNA | TriLink BioTechnologies | Co-transcriptionally capped mRNA for enhanced translational efficiency and reduced immunogenicity. |
| RiboGreen RNA Quantitation Kit | Thermo Fisher Scientific | Fluorometric quantification of total vs. encapsulated mRNA for determining EE%. |
| Gal8-mRuby3 Reporter Cell Line | Available through addgene or generated in-house | Live-cell biosensor for visualizing endosomal membrane disruption (escape). |
| Dynasore hydrate | Sigma-Aldrich, Tocris | Cell-permeable inhibitor of dynamin, used to confirm clathrin-mediated uptake pathway. |
| pHrodo Red dye | Thermo Fisher Scientific | pH-sensitive fluorophore for co-encapsulation to track LNP trafficking to acidic compartments. |
| Microfluidic Mixer (NanoAssemblr) | Precision NanoSystems | Enables reproducible, scalable formulation of uniform LNPs. |
| LipoDye Fluorescent Lipids | Avanti Polar Lipids | Fluorescently-labeled lipids for tracking LNP fate independently of mRNA. |
Application Notes
The evolution of Lipid Nanoparticles (LNPs) from first-generation systems, designed primarily for siRNA delivery, to advanced platforms for mRNA vaccines and therapeutics, represents a cornerstone in the broader thesis of SCP-Nano application research. This progression is characterized by targeted innovations addressing two critical limitations: stability (chemical, physical, and storage) and payload capacity (encapsulation efficiency and ability to carry larger or more complex nucleic acids).
Innovations in Stability
First-generation LNPs, utilizing ionizable lipids like DLin-MC3-DMA, demonstrated efficacy but faced challenges in long-term storage, often requiring frozen conditions (-20°C to -80°C) due to mRNA degradation and particle aggregation.
- Lipid Chemistry Evolution: The development of novel, structurally engineered ionizable lipids (e.g., proprietary lipids in COVID-19 vaccines) has enhanced particle stability. These lipids are designed for optimal pKa, ensuring cationic charge only at acidic pH during formulation, but remaining neutral at physiological pH, reducing cytotoxicity and improving colloidal stability.
- Excipient Optimization: The strategic inclusion of cholesterol derivatives (e.g., β-sitosterol) and modern polyethylene glycol (PEG)-lipids with optimized chain lengths and linker stability minimizes particle fusion and opsonization, extending shelf-life.
- Buffer & Cryoprotectant Systems: Advanced formulation buffers now include non-reducing sugars (trehalose, sucrose) as cryoprotectants and free radical scavengers to protect mRNA integrity during lyophilization and long-term storage at 2-8°C.
Innovations in Payload Capacity
Early LNPs had encapsulation efficiencies (EE) for mRNA that could be variable, limiting the dose consistency and therapeutic index for complex applications like gene editing or multi-mRNA cocktails.
- Microfluidic Mixing Precision: Transition from turbulent mixing to precise, scalable microfluidic hydrodynamic focusing allows reproducible, rapid mixing of aqueous and lipid phases. This yields homogeneous particles with >90% EE, critical for SCP-Nano applications requiring co-delivery of multiple components.
- Structural Lipid Design: New helper lipids (phospholipids, cholesterol variants) are engineered to promote a stable inverse hexagonal (HII) phase or other non-bilayer structures within the LNP core, creating more space and a protective environment for larger mRNA constructs (e.g., self-amplifying mRNA, saRNA) or mRNA-protein complexes.
- Charge-Mediated Complexation: Innovations in charge-neutralizable ionizable lipids enable higher nucleic acid loading without compromising particle size or endosomal release efficiency.
Table 1: Quantitative Comparison: First-Gen vs. Advanced LNPs
| Parameter | First-Generation LNPs (c. 2010-2016) | Advanced LNPs (c. 2020-Present) | Key Innovation Implication |
|---|---|---|---|
| Ionizable Lipid | DLin-MC3-DMA, DLin-KC2-DMA | SM-102, ALC-0315, proprietary structures | Optimized pKa (~6.2-6.6) for improved endosomal escape & reduced toxicity |
| Encapsulation Efficiency (mRNA) | 70-85% | Routinely >90%, often >95% | Higher dose consistency, lower waste, reduced immunogenicity from free mRNA |
| Storage Stability | -80°C for long-term; days at 4°C | 6-24 months at 2-8°C post-lyophilization | Enables global distribution, reduces cold chain burden |
| Payload Flexibility | siRNA, conventional mRNA | saRNA, circular RNA, CRISPR-Cas mRNA/gRNA, multi-mRNA cocktails | Supports complex SCP-Nano therapeutic strategies |
| Polydispersity Index (PDI) | 0.1 - 0.3 | 0.05 - 0.15 | More homogeneous particle population for predictable pharmacokinetics |
| In Vivo Potency (Relative) | 1x (Reference) | 10-100x improvement reported | Lower effective doses, improved therapeutic window |
Protocols
Protocol 1: Formulation of Advanced LNPs via Microfluidic Mixing for High-Payload Capacity
Objective: To reproducibly formulate mRNA-LNPs with >95% encapsulation efficiency and controlled particle size suitable for in vivo SCP-Nano research applications.
Materials:
- Lipid Stock Solutions: Ionizable lipid (e.g., SM-102), DSPC, Cholesterol, PEG-lipid (e.g., DMG-PEG2000) in ethanol.
- Aqueous Phase: mRNA in 10 mM citrate or acetate buffer (pH 4.0).
- Equipment: Precision syringe pumps, commercial microfluidic mixer (e.g., NanoAssemblr Ignite, microfluidic chip), PD-10 desalting columns, 0.22 µm sterile filters.
- Buffers: 1x PBS (pH 7.4), formulation buffer.
Procedure:
- Prepare Lipid Mixture: Combine ionizable lipid, DSPC, cholesterol, and PEG-lipid at a molar ratio (e.g., 50:10:38.5:1.5) in ethanol to a total lipid concentration of 12.5 mM. Maintain at room temperature (RT).
- Prepare mRNA Solution: Dilute mRNA in acidic aqueous buffer (pH 4.0) to a concentration of 0.1 mg/mL.
- Microfluidic Mixing: Load the lipid-ethanol solution and mRNA aqueous solution into separate syringes. Mount on syringe pumps. Connect syringes to a microfluidic chip. Set total flow rate (TFR) to 12 mL/min and flow rate ratio (FRR, aqueous:ethanol) to 3:1. Initiate simultaneous mixing. Collect effluent in a vial.
- Buffer Exchange & Dialysis: Immediately dilute the crude LNP mixture with an equal volume of 1x PBS (pH 7.4). Concentrate and dialyze against 1x PBS (pH 7.4) using a tangential flow filter or by dialysis in a 100kD MWCO cassette for 2 hours at RT to remove ethanol and establish neutral pH.
- Sterile Filtration: Filter the final LNP formulation through a 0.22 µm sterile filter. Store at 4°C for immediate use or aliquot and freeze at -80°C.
Protocol 2: Assessing mRNA Encapsulation Efficiency (EE%)
Objective: To accurately determine the percentage of mRNA encapsulated within LNPs.
Materials:
- Quant-iT RiboGreen RNA assay kit, Triton X-100, 1x TE buffer, black 96-well plate, fluorescent microplate reader.
Procedure:
- Prepare Samples: Dilute the LNP formulation to an estimated mRNA concentration within the assay's linear range (e.g., 10-200 ng/mL).
- Direct (Total) mRNA Measurement (A): In a microtube, mix 50 µL of diluted LNPs with 150 µL of 1x TE buffer containing 0.5% (v/v) Triton X-100. Vortex to fully disrupt LNPs. Incubate for 5 minutes at RT.
- Free (Unencapsulated) mRNA Measurement (B): In a separate microtube, mix 50 µL of the same diluted LNPs with 150 µL of 1x TE buffer only (no detergent).
- Assay: Prepare the RiboGreen dye per manufacturer's instructions. Add 100 µL of each sample (A and B) and standards to a black 96-well plate in duplicate. Add 100 µL of RiboGreen working solution to each well. Incubate for 5 minutes protected from light.
- Read Fluorescence: Measure fluorescence (excitation ~480 nm, emission ~520 nm).
- Calculation: Calculate mRNA concentration in (A) and (B) from the standard curve.
- EE% = [1 - (B / A)] x 100%
Protocol 3: Forced Degradation Study for Stability Assessment
Objective: To evaluate the physical and chemical stability of LNPs under accelerated stress conditions.
Materials:
- LNP formulation, Dynamic Light Scattering (DLS) instrument, agarose gel electrophoresis equipment, stability chambers.
Procedure:
- Sample Preparation: Aliquot identical volumes of LNP formulation into sterile vials.
- Stress Conditions:
- Thermal Stress: Incubate aliquots at 4°C (control), 25°C, and 37°C.
- Freeze-Thaw Stress: Subject aliquots to 3 cycles of freezing at -80°C for 24 hours and thawing at RT.
- Time Points: Analyze samples at T=0, 1, 3, 7, and 14 days for the thermal study, and after each freeze-thaw cycle.
- Analysis:
- Particle Size & PDI: Measure by DLS. A >20% increase in mean diameter or PDI indicates aggregation/instability.
- mRNA Integrity: Run samples on an agarose gel (after LNP disruption with Triton X-100) to visualize mRNA degradation.
- EE%: Perform RiboGreen assay as in Protocol 2 to track payload retention.
Diagrams
Title: Evolution Pathway from First-Gen to Advanced LNPs
Title: Advanced LNP Formulation and QC Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in LNP Research |
|---|---|
| Engineered Ionizable Lipids (e.g., SM-102, ALC-0315) | Core structural lipid with acid-triggered ionization; enables mRNA complexation, drives endosomal escape, and defines biodistribution. |
| PEG-Lipids (e.g., DMG-PEG2000, DSG-PEG2000) | Provides a hydrophilic stealth coating to prevent aggregation during formulation and opsonization in vivo; impacts circulation time. |
| Microfluidic Mixer (e.g., NanoAssemblr) | Enables reproducible, scalable, and rapid mixing of lipid and aqueous phases via hydrodynamic focusing, crucial for high EE% and monodisperse particles. |
| Quant-iT RiboGreen RNA Assay | Fluorescent dye-based assay for sensitive, specific quantification of both free and total RNA, allowing accurate calculation of encapsulation efficiency. |
| Trehalose Dihydrate | Non-reducing sugar used as a cryoprotectant and lyoprotectant; stabilizes LNPs and protects mRNA during freeze-drying and storage at 2-8°C. |
| Size Exclusion Chromatography Columns (e.g., PD-10) | Used for rapid buffer exchange and removal of unencapsulated mRNA, free lipids, and organic solvents from crude LNP formulations. |
| Cryogenic Vials with Silicone Gasket | Essential for stable, leak-proof, long-term storage of LNP formulations at ultra-low temperatures (-80°C) without degradation or contamination. |
From Bench to Application: A Stepwise Guide to SCP-LNP Formulation and mRNA Encapsulation
Within the broader thesis on SCP-Nano (Single-Chain Polymer-lipid nanoparticle) applications for mRNA delivery, the reproducible synthesis of these complex vehicles is paramount. This document provides detailed Application Notes and Protocols for two primary manufacturing methodologies: microfluidic mixing and precipitation-based self-assembly. Consistent synthesis is critical for establishing in vitro and in vivo structure-activity relationships, a core pillar of the overarching thesis.
Research Reagent Solutions: Essential Materials
| Item | Function | Typical Example/Details |
|---|---|---|
| Ionizable Lipid | Key structural & functional component; enables endosomal escape. | SM-102, DLin-MC3-DMA, or thesis-specific SCP-lipid conjugate. |
| Phospholipid | Stabilizes LNP bilayer structure. | DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine). |
| Cholesterol | Modulates membrane fluidity and stability. | Bio-sourced, >99% purity. |
| PEG-lipid | Controls particle size, prevents aggregation, modulates pharmacokinetics. | DMG-PEG2000 or PEG-DMG. |
| mRNA Payload | Therapeutic cargo; must be purified and in nuclease-free buffer. | CleanCap modified mRNA, e.g., encoding luciferase or target antigen. |
| Acidic Aqueous Buffer | For ionizable lipid protonation and LNP formation. | Citrate buffer, pH 4.0. |
| Ethanol | Solvent for lipid mixture. | 100% anhydrous ethanol, molecular biology grade. |
| Microfluidic Device | Enables rapid, reproducible mixing. | Staggered herringbone mixer (SHM) or T-junction chip. |
| Dialysis Cassettes/TFF | For buffer exchange and ethanol removal. | 20kD MWCO cassettes or Tangential Flow Filtration system. |
| Non-Invasive Back Scattering (NIBS) | For critical size and PDI measurement. | Zetasizer or equivalent Dynamic Light Scattering instrument. |
| RiboGreen Assay Kit | Quantifies encapsulation efficiency. | Fluorescence-based assay with/without detergent. |
| SYBR Gold Dye | For gel-based assessment of mRNA integrity post-encapsulation. | Fluorescent nucleic acid gel stain. |
Table 1: Comparison of Key Process Parameters and Outputs
| Parameter | Microfluidic Mixing (SHM) | Precipitation (Ethanol Injection) |
|---|---|---|
| Mixing Principle | Rapid diffusive mixing in <100 ms | Turbulent mixing during injection |
| Flow Rate Ratio (Aq:Eth) | 3:1 (Typical) | 1:1 to 4:1 (Variable) |
| Total Flow Rate (TFR) | 12 mL/min (Optimal for SHM) | Injection rate ~1 mL/min |
| Process Duration | Minutes (Continuous) | Seconds per batch (Batch) |
| Typical Particle Size | 70 - 100 nm (Tight distribution) | 80 - 150 nm (Broader distribution) |
| Polydispersity Index (PDI) | <0.1 (Excellent) | 0.1 - 0.2 (Good to Moderate) |
| Encapsulation Efficiency | >90% (Consistently high) | 70% - 90% (Variable) |
| Scalability | Linear scale-out via parallel chips | Challenging; batch consistency varies |
| Key Advantage | Superior reproducibility, tight control | Simplicity, low equipment cost |
Table 2: Standard Lipid Composition for SCP-LNP Formulation
| Lipid Component | Molar Ratio (%) | Stock Concentration (mg/mL in EtOH) | Function in Thesis Context |
|---|---|---|---|
| Ionizable Lipid (SCP-conjugate) | 50.0 | 20.0 | Thesis core: Enables tunable delivery & targeting. |
| DSPC | 10.0 | 10.0 | Provides structural integrity to bilayer. |
| Cholesterol | 38.5 | 20.0 | Stabilizes particle, enhances in vivo circulation. |
| DMG-PEG2000 | 1.5 | 10.0 | Controls size; may be modified for thesis targeting. |
Detailed Experimental Protocols
Protocol 4.1: Microfluidic Synthesis using a Staggered Herringbone Mixer (SHM)
Objective: Reproducibly synthesize SCP-LNPs of ~80 nm with high mRNA encapsulation efficiency.
Materials:
- Lipid stock solutions in ethanol (Table 2).
- mRNA in 10 mM citrate buffer, pH 4.0.
- SHM microfluidic chip (e.g., Dolomite).
- Precision syringe pumps (2).
- Collection tube with 1x PBS (pH 7.4), 4x the volume of the ethanolic stream.
- Dialysis cassettes (20kD MWCO) or TFF system.
Procedure:
- Lipid Preparation: Combine ionizable lipid, DSPC, cholesterol, and PEG-lipid in a glass vial according to molar ratios. Dry under nitrogen stream and redissolve in anhydrous ethanol to the final total lipid concentration (typically 5-10 mg/mL). Keep at room temperature (RT).
- mRNA Preparation: Dilute purified mRNA to 0.1 mg/mL in 10 mM citrate buffer (pH 4.0). Keep on ice.
- Pump Setup: Load the ethanolic lipid solution into one syringe and the acidic mRNA solution into another. Mount on separate pumps.
- Chip Priming: Prime the microfluidic chip channels with ethanol and then citrate buffer.
- Mixing & Synthesis: Set flow rates to achieve a Total Flow Rate (TFR) of 12 mL/min and an Aqueous-to-Ethanol Flow Rate Ratio (FRR) of 3:1 (e.g., aqueous at 9 mL/min, ethanol at 3 mL/min). Start pumps simultaneously. Collect the turbid effluent directly into a tube containing 1x PBS (pH 7.4) to facilitate immediate neutralization.
- Buffer Exchange & Purification: Dialyze the collected LNP solution against a large volume of 1x PBS (pH 7.4) for 4 hours at 4°C with one buffer change, or use TFF with 100kD membranes. Sterile filter through a 0.22 μm PES membrane.
- Analysis: Proceed to characterization (Section 5).
Protocol 4.2: Precipitation-Based Synthesis (Ethanol Injection)
Objective: Synthesize SCP-LNPs using a simpler, bench-top precipitation method.
Materials:
- Lipid stock solutions in ethanol (Table 2).
- mRNA in 10 mM citrate buffer, pH 4.0.
- Magnetic stirrer and vial.
- Single syringe or pipette.
Procedure:
- Lipid & mRNA Prep: Prepare ethanolic lipid mixture and acidic mRNA solution as in Protocol 4.1.
- Mixing Setup: Place the aqueous mRNA solution (e.g., 1 mL) in a glass vial under rapid magnetic stirring (≈1000 rpm) at RT.
- Injection: Rapidly inject the ethanolic lipid solution (e.g., 0.33 mL for a 3:1 FRR) into the center of the vortexing aqueous phase using a syringe or pipette. Instantaneous particle formation occurs.
- Dilution & Neutralization: Immediately after injection, add an equal volume of 1x PBS (pH 7.4) to the mixture.
- Buffer Exchange & Purification: Identical to Step 6 in Protocol 4.1.
- Analysis: Proceed to characterization.
Essential Characterization Protocols
5.1. Size and PDI by Dynamic Light Scattering:
- Dilute 10 μL of final LNP formulation in 990 μL of nuclease-free 1x PBS. Mix gently.
- Load into a disposable cuvette.
- Measure using a DLS instrument with settings: temperature 25°C, equilibration 60 sec, 3 measurements of 12 runs each.
- Report Z-average diameter and PDI from the intensity-based distribution.
5.2. mRNA Encapsulation Efficiency by RiboGreen Assay:
- Prepare a 1:200 dilution of RiboGreen reagent in TE buffer.
- Prepare two sets of LNP samples in a black 96-well plate:
- Total mRNA (A): Dilute LNPs 1:100 in TE buffer with 0.5% Triton X-100.
- Free/unencapsulated mRNA (B): Dilute LNPs 1:100 in TE buffer only.
- Add equal volume of diluted RiboGreen to each well. Incubate 5 min, protected from light.
- Measure fluorescence (ex: 480 nm, em: 520 nm).
- Calculate EE% = [1 - (Fluorescence B / Fluorescence A)] x 100. Use an mRNA standard curve for absolute quantification if needed.
Visualization Diagrams
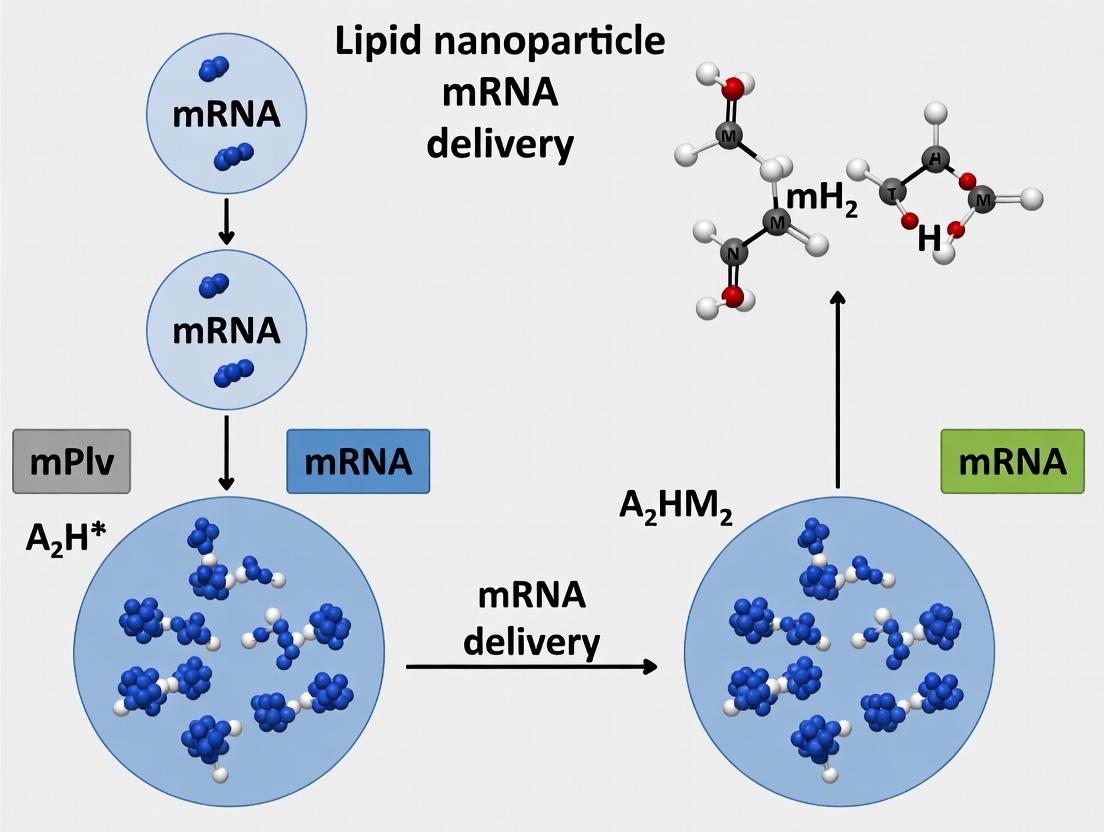
SCP-LNP Formation via pH-Driven Self-Assembly
Microfluidic Workflow for SCP-LNP Synthesis
SCP-LNP mRNA Delivery & Endosomal Escape Pathway
Within the broader thesis on SCP-Nano application lipid nanoparticles (LNPs) for mRNA delivery, the integrity and purity of the mRNA payload are critical determinants of therapeutic efficacy. Impurities, including truncated transcripts, double-stranded RNA (dsRNA), and residual process enzymes, can trigger innate immune responses, reduce translation efficiency, and negatively impact LNP formulation stability. This Application Note details protocols for assessing mRNA integrity and purifying mRNA to pharmaceutical-grade standards suitable for high-efficiency encapsulation into SCP-Nano LNPs.
Quantifying mRNA Integrity: Key Metrics and Methods
Analytical Techniques and Acceptable Ranges
The following table summarizes key analytical methods used to quantify mRNA integrity and purity.
Table 1: Quantitative Metrics for mRNA Integrity and Purity Assessment
| Analytic | Method | Target Specification | Impact on Encapsulation & Performance |
|---|---|---|---|
| Purity (A260/A280) | UV Spectrophotometry | 2.0 - 2.2 | Ratios outside range indicate protein or solvent contamination, affecting LNP surface charge and stability. |
| Purity (A260/A230) | UV Spectrophotometry | 2.0 - 2.4 | Low ratio indicates residual Guanidine Thiocyanate or EDTA, which can disrupt lipid bilayer formation. |
| Full-Length Content | Capillary Electrophoresis (e.g., Fragment Analyzer, Bioanalyzer) | ≥ 80% | Truncated species reduce the dose of active payload and may incorporate less efficiently into LNPs. |
| dsRNA Impurity | ELISA or HPLC-based assays (e.g., dsRNA SCICEX) | ≤ 0.1% (ng/μg mRNA) | Potent activator of PKR and TLR3, leading to increased immunogenicity and reduced protein expression. |
| Residual DNA Template | qPCR | ≤ 0.5 ng/μg mRNA | Risk of genomic integration or unwanted immune activation. |
| Capping Efficiency | LC-MS or enzymatic assays | ≥ 95% | Directly correlates with translation initiation efficiency; uncapped mRNA is rapidly degraded. |
| Endotoxin | LAL Assay | < 0.05 EU/μg mRNA | Pyrogenic contaminant causing severe inflammatory reactions. |
Detailed Protocol: Agarose Gel Electrophoresis for Rapid Integrity Check
- Objective: Qualitative assessment of mRNA size and integrity.
- Reagents: LE Agarose, MOPS buffer, RNA loading dye, SYBR Green II RNA gel stain.
- Procedure:
- Prepare a 1.2% agarose gel in 1X MOPS buffer.
- Dilute 1 μg of mRNA sample in nuclease-free water, mix with loading dye (non-DENATURING).
- Load samples alongside an appropriate RNA ladder.
- Run gel at 5-6 V/cm in 1X MOPS buffer until adequate separation is achieved.
- Stain gel with SYBR Green II (diluted 1:10,000 in 1X MOPS) for 30 min in the dark.
- Image using a gel documentation system with a blue-light or UV transilluminator.
- Interpretation: A single, sharp band at the expected size indicates high integrity. Smearing below the main band indicates degradation.
Detailed Protocol: Quantifying dsRNA via ELISA
- Objective: Quantify immunogenic dsRNA impurities.
- Reagents: dsRNA ELISA Kit (e.g., J2 antibody-based), microplate reader.
- Procedure:
- Prepare mRNA sample dilutions in the provided buffer (typically 50-100 ng/μL).
- Add 100 μL of standard, control, or sample to the antibody-coated wells in duplicate.
- Incubate 1 hour at room temperature (RT).
- Aspirate and wash wells 4 times with Wash Buffer.
- Add 100 μL of detection antibody (conjugated to HRP). Incubate 1 hour at RT.
- Aspirate and wash 4 times.
- Add 100 μL of TMB Substrate. Incubate 10-15 min in the dark.
- Add 100 μL Stop Solution. Read absorbance at 450 nm immediately.
- Generate a standard curve and interpolate sample concentrations.
mRNA Purification Protocols
Oligo dT-Based Affinity Purification
This method selects for mature, polyadenylated mRNA, removing truncated transcripts, residual DNA, and enzymes.
Table 2: Key Reagents for Oligo dT Purification
| Reagent | Function |
|---|---|
| Oligo dT Magnetic Beads | Poly(T) sequences bind poly(A)+ tail of mRNA. |
| Binding Buffer (High Salt) | Creates conditions favorable for poly(A)-dT hybridization. |
| Wash Buffer (Low Salt) | Removes contaminants while keeping mRNA bound. |
| Nuclease-Free Water (Elution) | Low ionic strength disrupts dT-A binding, eluting pure mRNA. |
- Detailed Protocol:
- Bind: Mix in vitro transcription (IVT) reaction with equal volume of Binding Buffer. Add to pre-washed Oligo dT beads. Rotate 5-10 min at RT.
- Wash: Pellet beads on a magnet. Discard supernatant. Wash twice with Wash Buffer.
- Elute: Resuspend beads in pre-warmed (65°C) nuclease-free water. Incubate 2 min. Pellet beads and transfer pure mRNA supernatant to a new tube.
- Concentrate: Use a centrifugal concentrator (e.g., 10K MWCO) if needed.
Fast Protein Liquid Chromatography (FPLC) for Industrial-Scale Purification
- Objective: Scalable, high-resolution separation of full-length mRNA from critical impurities.
- System: ÄKTA pure or comparable FPLC.
- Column: Anion-exchange (e.g., HiTrap Q HP) or reverse-phase (e.g, C4).
- Detailed Protocol (Anion-Exchange):
- Sample Prep: Dilute IVT reaction 1:5 in Buffer A (20 mM Tris, pH 7.5).
- Equilibration: Equilibrate column with 5 column volumes (CV) of Buffer A.
- Injection & Wash: Inject sample. Wash with 5-10 CV of Buffer A to remove proteins and nucleotides.
- Gradient Elution: Run a linear gradient from 0% to 100% Buffer B (Buffer A + 1M NaCl) over 20 CV. Full-length mRNA elutes at a specific salt concentration (~250-400 mM NaCl).
- Collection & Desalting: Collect peak fractions. Desalt using size-exclusion chromatography (e.g., G-25 column) or tangential flow filtration into nuclease-free water.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for mRNA Integrity & Purification Workflows
| Item | Function |
|---|---|
| RNaseZap or RNase Away | Decontaminates surfaces and equipment to prevent RNase-mediated degradation. |
| Nuclease-Free Water & Tubes | Essential for all reagent prep and sample handling to maintain RNA integrity. |
| Agilent 4200 TapeStation / Fragment Analyzer | Automated capillary electrophoresis for precise RNA Integrity Number (RIN) or % full-length calculation. |
| SPRIselect / AMPure XP Beads | Solid-phase reversible immobilization (SPRI) beads for clean-up and size selection of RNA. |
| KAPA mRNA Quant Kit (qPCR) | Accurate, specific quantification of functional mRNA, superior to UV spectroscopy. |
| Cellulose-Based dsRNA Removal Beads | Selective binding and removal of dsRNA impurities post-IVT (e.g., LGC, MagJET). |
| CleanCap Reagent (TriLink) | Co-transcriptional capping agent yielding >95% Cap 1 structure, enhancing translation. |
| HPLC-Grade Ethanol & Salts | For precipitation and buffer preparation in purification protocols. |
Visualizations
mRNA Quality Control Workflow
Impurity Impact on Immune Pathways
Optimizing N/P Ratios and Buffer Conditions for Maximal mRNA Loading in SCP-LNPs
Within the broader thesis on the application of Selective Cationic Lipid Particles (SCP-Nano) for mRNA delivery, a critical parameter for therapeutic efficacy is the optimization of messenger ribonucleic acid (mRNA) encapsulation. This application note details systematic approaches to maximize mRNA loading into SCP-Lipid Nanoparticles (LNPs) by investigating two interdependent factors: the nitrogen-to-phosphate (N/P) ratio and the formulation buffer conditions. Efficient loading is paramount for ensuring dose consistency, minimizing waste of costly mRNA, and achieving potent in vivo expression.
Core Principles and Key Variables
The N/P ratio is the molar ratio of positively charged (amine) groups from the ionizable cationic lipid to the negatively charged (phosphate) groups from the mRNA backbone. An optimal ratio ensures complete charge neutralization and condensation of mRNA, facilitating efficient encapsulation during the self-assembly process. Buffer conditions (pH, ionic strength, buffer species) critically influence the protonation state of the ionizable lipid and the mRNA conformation, thereby impacting complex stability and final LNP characteristics.
Experimental Protocols
Protocol 3.1: Microfluidic Formulation of SCP-LNPs with Varied N/P Ratios
Objective: To prepare SCP-LNPs across a range of N/P ratios for loading efficiency analysis. Materials:
- Lipid Stock Solution: SCP-108 (ionizable lipid), DSPC, Cholesterol, DMG-PEG-2000 dissolved in ethanol (e.g., 90% v/v final).
- mRNA Stock Solution: CleanCap eGFP mRNA or target therapeutic mRNA in aqueous buffer (e.g., 10 mM citrate, pH 4.0).
- Equipment: Precision microfluidic mixer (e.g., NanoAssemblr), syringe pump, thermoblock.
Procedure:
- Prepare the lipid mix in ethanol at a fixed total lipid concentration (e.g., 12.5 mM) with a constant molar ratio of auxiliary lipids (e.g., SCP-108:DSPC:Chol:DMG-PEG = 50:10:38.5:1.5). Maintain the total lipid quantity constant across all N/P formulations.
- Prepare the aqueous mRNA phase at a fixed concentration (e.g., 0.1 mg/mL) in the target optimization buffer (e.g., 25 mM acetate, pH 5.0).
- Calculate the required volumes of lipid and mRNA phases to achieve target N/P ratios (e.g., 2, 4, 6, 8, 10) using the known pKa of SCP-108 (~6.2) and mRNA phosphate concentration.
- Load the phases into separate syringes. Using the microfluidic mixer, combine streams at a fixed total flow rate (e.g., 12 mL/min) and a flow rate ratio (FRR, aqueous:organic) of 3:1.
- Collect the formulated LNPs in a vial. Dialyze against 1x PBS (pH 7.4) for 2 hours at 4°C using a Slide-A-Lyzer cassette (10K MWCO) to remove ethanol and exchange the buffer.
- Filter the resulting SCP-LNP dispersion through a 0.22 µm sterile filter. Store at 4°C until analysis.
Protocol 3.2: Quantification of mRNA Loading Efficiency via Ribogreen Assay
Objective: To determine the percentage of mRNA encapsulated within SCP-LNPs. Materials: Quant-iT RiboGreen RNA Assay Kit, TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 7.5), Triton X-100 (2% v/v solution), plate reader.
Procedure:
- Prepare two sets of diluted SCP-LNP samples in TE buffer (e.g., 1:100 dilution).
- To one set, add Triton X-100 to a final concentration of 0.5% to disrupt LNPs and release total mRNA (Total signal).
- Leave the second set untreated to measure unencapsulated/free mRNA (Free signal).
- Prepare an RNA standard curve according to the kit protocol.
- Add RiboGreen dye to all samples and standards, incubate for 5 minutes protected from light.
- Measure fluorescence (excitation ~480 nm, emission ~520 nm).
- Calculate loading parameters:
- Encapsulation Efficiency (EE%) = [1 - (Free RNA Concentration / Total RNA Concentration)] × 100.
- Loading Capacity = (Mass of encapsulated mRNA / Total mass of lipids) × 100.
Table 1: Impact of N/P Ratio on mRNA Loading in 25 mM Acetate Buffer (pH 5.0)
| N/P Ratio | Encapsulation Efficiency (%) (Mean ± SD) | Particle Size (nm, PDI) | Zeta Potential (mV) |
|---|---|---|---|
| 2 | 65.2 ± 5.1 | 102 (0.18) | -3.5 |
| 4 | 88.7 ± 2.3 | 89 (0.12) | 1.2 |
| 6 | 98.5 ± 0.8 | 85 (0.08) | 3.8 |
| 8 | 97.9 ± 1.1 | 87 (0.09) | 5.1 |
| 10 | 96.4 ± 1.5 | 91 (0.10) | 6.5 |
Table 2: Effect of Buffer Condition at Optimal N/P Ratio (N/P=6)
| Buffer Condition (pH) | Encapsulation Efficiency (%) | Particle Size (nm) | Notes |
|---|---|---|---|
| Citrate (10 mM, pH 4.0) | 99.1 ± 0.5 | 82 | Maximal lipid protonation, may impact stability. |
| Acetate (25 mM, pH 5.0) | 98.5 ± 0.8 | 85 | Optimal for SCP-108 (pKa~6.2), standard condition. |
| Succinate (25 mM, pH 5.5) | 95.2 ± 1.8 | 88 | Intermediate protonation. |
| PBS (pH 7.4) | 70.3 ± 8.4 | 105 | Insufficient protonation, poor self-assembly. |
Visualization of Workflows and Relationships
Title: mRNA Loading Optimization Decision Pathway
Title: SCP-LNP Self-Assembly via Microfluidics
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for SCP-LNP mRNA Loading Optimization
| Item | Function/Description | Example Product/Criteria |
|---|---|---|
| Ionizable Cationic Lipid | The core SCP component; provides pH-dependent positive charge for mRNA complexation. pKa is critical. | SCP-108 (pKa ~6.2), ALC-0315 (Onpattro). |
| Structural Helper Lipid | Provides structural integrity and bilayer stability to the LNP. | DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine). |
| Cholesterol | Modulates membrane fluidity, stability, and facilitates fusion with endosomal membranes. | Pharmaceutical grade, >99% purity. |
| PEG-lipid | Controls particle size during formation, reduces aggregation, and modulates pharmacokinetics. | DMG-PEG2000 (1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol). |
| mRNA Construct | The payload. Chemical purity, integrity (capping/poly-A tail), and sequence are vital. | CleanCap modified mRNA, HPLC purified. |
| Acidic Buffer Salts | Creates aqueous phase at pH below lipid pKa to ensure protonation during formulation. | Sodium Acetate (25 mM, pH 5.0), Citric Acid. |
| Microfluidic Mixer | Enables rapid, reproducible, and scalable mixing of lipid and aqueous streams. | NanoAssemblr (Precision NanoSystems), chaotic mixer chips. |
| Encapsulation Assay Kit | Fluorescent dye-based quantitation of total vs. free RNA. Industry standard. | Quant-iT RiboGreen RNA Assay Kit (Thermo Fisher). |
| Dialysis System | For buffer exchange and ethanol removal post-formulation. | Slide-A-Lyzer G2 Cassettes (10K MWCO). |
This application note, framed within a broader thesis on SCP-Nano (Selective Cell-targeted Programmable Nanoparticles) platforms, details the formulation and application of SCP-Lipid Nanoparticles (LNPs) across three therapeutic modalities. SCP-LNPs are distinguished by their engineered lipid compositions and surface conjugations for cell-type-specific delivery, enhancing efficacy and safety in mRNA-based interventions.
Application Note: Prophylactic Vaccine Against Emerging Pathogen (e.g., SARS-CoV-2 Variant)
Objective: To develop an SCP-LNP-mRNA vaccine that targets antigen-presenting cells (APCs) in lymph nodes for potent and durable humoral and cellular immunity.
SCP-LNP Design Rationale:
- Ionizable Lipid: SM-102 or ALC-0315 analogues for efficient endosomal escape at acidic pH.
- PEG-lipid: DMG-PEG2000 reduced to 0.5 mol% to minimize the "PEG-dilemma" and allow faster APC uptake.
- Targeting Moiety: Mannose-PEG-DSG conjugate incorporated post-formulation to actively target mannose receptors (CD206) on dendritic cells and macrophages.
Key Quantitative Data: Table 1: Characterization and *In Vivo Immunogenicity of APC-Targeted SCP-LNP Vaccine*
| Parameter | Value/Result | Method |
|---|---|---|
| Particle Size (Z-avg) | 75 ± 5 nm | DLS |
| Polydispersity Index (PDI) | 0.08 ± 0.02 | DLS |
| Encapsulation Efficiency | 95 ± 3% | RiboGreen assay |
| Surface Zeta Potential | -2 ± 1 mV | DLS |
| Neutralization Titer (Day 28) | 1:25,600 | Pseudovirus assay |
| CD8+ T-cell Response (IFN-γ SFU/10^6 cells) | 450 ± 80 | ELISpot |
Detailed Protocol: SCP-LNP Formulation via Microfluidic Mixing
- Lipid Stock Preparation: Dissolve ionizable lipid (SM-102, 50 mmol), DSPC (10 mmol), cholesterol (38.5 mmol), and DMG-PEG2000 (1.5 mmol) in ethanol. Prepare a separate stock of Mannose-PEG-DSG (1.0 mmol) in ethanol.
- Aqueous Phase: Dilute mRNA encoding the pathogen's spike protein variant in 50 mM citrate buffer (pH 4.0) to 0.1 mg/mL.
- Nanoparticle Formation: Using a microfluidic device (e.g., NanoAssemblr), mix the ethanolic lipid phase (excluding mannose conjugate) with the aqueous mRNA phase at a 1:3 flow rate ratio (total flow rate 12 mL/min).
- Targeting Ligand Conjugation: Dialyze formulated LNPs against PBS (pH 7.4) for 2 hours. Incubate with the Mannose-PEG-DSG ethanolic stock (post-insertion method) for 30 min at 37°C.
- Purification & Storage: Concentrate using centrifugal filters (100 kDa MWCO). Sterile filter (0.22 µm). Store at 4°C for short-term or -80°C for long-term use.
Application Note:In VivoGene Editing for Genetic Disorder (e.g., Transthyretin Amyloidosis)
Objective: To deliver CRISPR-Cas9 ribonucleoprotein (RNP) or mRNA/sgRNA components to hepatocytes for knockout of the disease-causing TTR gene.
SCP-LNP Design Rationale:
- Ionizable Lipid: Novel biodegradable lipid, like KC2 or 306Oi10, optimized for liver tropism and high RNP payload.
- Helper Lipid: Cholesterol replaced with β-sitosterol to enhance LDLR-mediated hepatocyte uptake.
- Targeting: Incorporate GalNAc-PEG-lipid for selective asialoglycoprotein receptor (ASGPR) targeting on hepatocytes.
Key Quantitative Data: Table 2: Characterization and *In Vivo Editing Efficiency of Hepatocyte-Targeted SCP-LNP for CRISPR Delivery*
| Parameter | Value/Result | Method |
|---|---|---|
| Particle Size (Z-avg) | 85 ± 8 nm | DLS |
| Payload | Cas9 mRNA + sgRNA (mass ratio 3:1) | N/A |
| Liver Accumulation (%ID/g) | >80% | In vivo imaging |
| Serum TTR Reduction | 92% at week 4 | ELISA |
| Indel Frequency in Liver | 45% ± 7% | NGS of target locus |
| Off-Target Indels | Undetectable | GUIDE-seq analysis |
Detailed Protocol: Co-encapsulation of Cas9 mRNA and sgRNA in SCP-LNPs
- sgRNA Preparation: Synthesize sgRNA via in vitro transcription, purify by phenol-chloroform extraction and ethanol precipitation.
- Complexation: Pre-mix Cas9 mRNA and sgRNA at a 3:1 mass ratio in sodium acetate buffer (pH 5.0) and incubate at room temp for 10 min to allow RNP complex formation in situ post-delivery.
- LNP Formulation: Prepare ethanolic lipid phase (KC2, β-sitosterol, DOPE, GalNAc-PEG-DMG). Use the mRNA/sgRNA mixture as the aqueous phase. Employ staggered herringbone micromixer (flow rate 15 mL/min) for rapid mixing.
- Buffer Exchange: Dialyze against PBS (pH 7.4) overnight at 4°C to remove ethanol and stabilize particles.
- In Vivo Administration: Administer via tail vein injection in mouse model at a dose of 1.0 mg mRNA/kg body weight. Analyze editing and protein levels at 1- and 4-weeks post-injection.
Application Note: Protein Replacement Therapy for Metabolic Disease (e.g., Methylmalonic Acidemia)
Objective: To deliver mRNA encoding methylmalonyl-CoA mutase (MUT) to hepatocytes for sustained production of functional enzyme.
SCP-LNP Design Rationale:
- Ionizable Lipid: DLIN-MC3-DMA (Onpattro lipid) for proven hepatic delivery and translation.
- PEG-lipid: ALC-0159 with extended PEG chain (PEG5000) for improved pharmacokinetic profile.
- Stabilizing Lipid: Include 1 mol% of a reactive oxygen species (ROS)-scavenging lipid (e.g., PVL-2) to protect mRNA integrity.
Key Quantitative Data: Table 3: Efficacy of SCP-LNP Delivering Therapeutic Protein mRNA
| Parameter | Value/Result | Method |
|---|---|---|
| Particle Size (Z-avg) | 90 ± 10 nm | DLS |
| mRNA Purity | >90% (no dsRNA) | HPLC |
| Protein Expression Onset | 4 hours post-injection | Luciferase imaging |
| Protein Expression Duration | >7 days | Luciferase imaging |
| Reduction in Plasma MMA | 85% from baseline | LC-MS/MS |
| Dosing Frequency | Every 10 days | Efficacy maintenance |
Detailed Protocol: Assessing In Vivo Protein Expression Kinetics
- Formulation: Prepare MUT mRNA-LNPs as per protocol in Section 1, using DLIN-MC3-DMA and ALC-0159 lipids.
- Dosing: Inject C57BL/6 mice (n=5/group) via tail vein with 0.5 mg/kg mRNA dose.
- Sampling: Collect blood plasma and liver tissue at pre-determined time points (e.g., 2h, 6h, 24h, 3d, 7d, 14d).
- Analysis:
- qRT-PCR: Quantify mRNA levels in liver homogenate.
- Western Blot/ELISA: Quantify MUT protein expression in liver lysate.
- Enzymatic Activity Assay: Measure MUT activity in tissue homogenates using a coupled spectrophotometric assay monitoring succinyl-CoA production.
- Metabolite Analysis: Quantify plasma methylmalonic acid (MMA) levels by mass spectrometry.
The Scientist's Toolkit: Essential Reagents for SCP-LNP Research
| Item/Category | Function | Example (Supplier) |
|---|---|---|
| Ionizable Cationic Lipids | Core component for mRNA complexation & endosomal escape. Dictates tropism. | SM-102, ALC-0315, KC2, DLIN-MC3-DMA (Avanti, BroadPharm) |
| PEGylated Lipids | Modulates particle stability, size, PK, and opsonization. Can be functionalized. | DMG-PEG2000, ALC-0159, DSPE-PEG(2000)-Malenimide (Avanti) |
| Structural Helper Lipids | Supports bilayer structure and integrity. | DSPC, DOPE, Cholesterol (Avanti) |
| Targeting Ligand Conjugates | Enables selective cell targeting. | GalNAc-PEG-DSG, Mannose-PEG-DSPE, Antibody-PEG-DSPE (BroadPharm) |
| mRNA (CleanCap) | Therapeutic payload with enhanced translation and reduced immunogenicity. | Cap 1 modified, pseudouridine-incorporated (TriLink BioTechnologies) |
| Microfluidic Mixer | Enables reproducible, scalable LNP formulation. | NanoAssemblr (Precision NanoSystems), µSMM (Dolomite) |
| RiboGreen Assay Kit | Sensitive quantification of encapsulated vs. free mRNA. | (Thermo Fisher Scientific) |
| Dynamic Light Scattering (DLS) | Instrument for measuring particle size (Z-avg) and polydispersity (PDI). | Zetasizer (Malvern Panalytical) |
Visualization: Pathways and Workflows
SCP-LNP Vaccine Immunological Pathway
SCP-LNP Design Logic Flowchart
Solving SCP-LNP Challenges: Strategies for Enhancing Stability, Efficacy, and Safety
Application Notes & Protocols
Context: These notes are part of a broader thesis on SCP-Nano applications in lipid nanoparticle (LNP)-mediated mRNA delivery, focusing on enhancing stability for global distribution.
Key Formulation Parameters and Stability Data
Recent research identifies ionizable lipid structure, helper lipid choice, and buffer composition as critical levers for improving LNP stability at elevated temperatures.
Table 1: Impact of Ionizable Lipid Tail Saturation on mRNA-LNP Stability at 25°C
| Ionizable Lipid (Example) | Tail Saturation | % mRNA Intact (4 weeks) | PDI Change (Initial→4 wks) | In Vivo Potency Retention (vs. -80°C fresh) |
|---|---|---|---|---|
| DLin-MC3-DMA | Polyunsaturated | 45% | 0.08 → 0.32 | 60% |
| C12-200 | Less unsaturated | 82% | 0.06 → 0.11 | 92% |
| ALC-0315 | Saturated Branched | 90% | 0.05 → 0.07 | 95% |
Table 2: Effect of Stabilizing Excipients in Tris-Sucrose Buffer (pH 7.4)
| Excipient & Concentration | Stability at 4°C (6 mo) | Stability at 25°C (1 mo) | Proposed Primary Mechanism |
|---|---|---|---|
| Baseline (No additive) | 95% mRNA intact | 40% mRNA intact | N/A |
| 0.5% w/v Trehalose | 98% | 75% | Water replacement, Vitrification |
| 1% w/v Sorbitol | 96% | 65% | Partial exclusion from LNP surface |
| 5 mM EDTA | 99% | 82% | Chelation of catalytic metals |
| 0.1% Poloxamer 188 | 97% | 88% | Steric stabilization, particle barrier |
Detailed Experimental Protocols
Protocol 2.1: High-Throughput Screening of LNP Stability
Objective: To systematically assess the stability of LNP formulations under various stress conditions. Materials: Microfluidic mixer, plate reader, dynamic light scattering (DLS) instrument, RNase Alert kit. Procedure:
- Formulate LNPs using a staggered herringbone micromixer. Fix mRNA at 0.1 mg/mL. Vary ionizable lipid:helper lipid:cholesterol:PEG-lipid ratios (e.g., 50:10:38.5:1.5 to 35:15:46.5:3.5).
- Dialyze formulations against 1L of selected buffer (e.g., 10 mM Tris, 10% sucrose, pH 7.4) for 2 hours at 4°C.
- Aliquot 100 µL of each formulation into sterile PCR tubes.
- Apply Stress Conditions:
- Thermal: Incubate aliquots at 4°C, 25°C, and 40°C.
- Freeze-Thaw: Cycle aliquots between -20°C and 25°C (5 cycles, 30 min per phase).
- Analyze at T=0, 1, 2, 4 weeks:
- Size & PDI: Dilute LNPs 1:100 in 1 mM Tris pH 7.4, measure by DLS.
- mRNA Integrity: Extract mRNA (phenol-chloroform), run on 1% agarose E-gel.
- Encapsulation Efficiency: Use Ribogreen assay. Measure fluorescence with/without 0.1% Triton X-100.
- In Vitro Potency: Transfert HEK293T cells, measure protein expression via luciferase assay at 24h.
Protocol 2.2: Assessing mRNA Protection via Nuclease Challenge
Objective: Quantify the protective capability of stable LNP shells. Materials: Formulated LNPs, DNase I, RNase A, RNase Alert v2 substrate, qPCR machine. Procedure:
- Prepare Samples: Dilute LNPs to 10 µg mRNA/mL in nuclease-free PBS. Set up three conditions per formulation:
- A: LNP + PBS (control)
- B: LNP + 0.1 µg/mL RNase A
- C: Naked mRNA + 0.1 µg/mL RNase A (degradation control)
- Incubate at 37°C for 30 minutes.
- Halt Reaction by adding 2U/mL SUPERase•In RNase inhibitor and placing on ice.
- Dissolve LNP using 0.5% sodium dodecyl sulfate (SDS).
- Quantify Protected mRNA via reverse transcription quantitative PCR (RT-qPCR) targeting the encoded transgene. Calculate % protection = (mRNA in Condition B / mRNA in Condition A) x 100.
Visualizations
Diagram Title: LNP Stability Optimization Logic for SCP-Nano Thesis
Diagram Title: LNP Stability Screening Protocol Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for LNP Stability Research
| Item & Example Product | Function in Stability Research | Key Consideration |
|---|---|---|
| Ionizable Lipids (e.g., ALC-0315, SM-102, proprietary SCP-Nano lipids) | Core structural component; determines bilayer fluidity, biodegradability, and fusogenicity. Saturated tails improve oxidative stability. | Tail saturation and branching are critical for shelf-life. |
| Helper Lipids (e.g., DSPC, DOPE, DPPC) | Modulate LNP bilayer structure and rigidity. DSPC enhances stability at room temperature versus fusogenic DOPE. | Phase transition temperature (Tm) directly impacts storage stability. |
| PEGylated Lipids (e.g., ALC-0159, DMG-PEG2000, PEG-DMG with C18 tail) | Provides steric stabilization, prevents aggregation. Shorter PEG anchors (C14) promote shedding for better efficacy but may reduce storage stability. | Anchor chain length and PEG molecular weight balance stability and in vivo performance. |
| Stabilizing Cryo-/Lyoprotectants (e.g., Trehalose, Sucrose) | Form a glassy matrix, replace water molecules, inhibit fusion and degradation during thermal stress. | Typically used at 5-10% w/v in final buffer. |
| Chelating Agents (e.g., EDTA, Citrate) | Bind trace metal ions (Fe2+, Cu2+) that catalyze lipid oxidation and mRNA degradation. | Use at 0.1-1 mM concentration. |
| RNase Inhibitors (e.g., SUPERase•In, RNasin) | Protect mRNA during handling and analysis post-stress. Critical for accurate integrity assessment. | Add to lysis or assay buffers, not typically to final formulation. |
| Size Exclusion Columns (e.g., Sephadex G-25, Illustra NAP-10) | For rapid buffer exchange of small-volume LNP samples during screening. | Faster but less efficient than dialysis/TFF. |
| Ribogreen Assay Kit (Quant-iT RiboGreen) | Dual-use: quantify total and free mRNA to calculate encapsulation efficiency (%EE), a key stability metric. | Requires careful titration of detergent (Triton X-100) for complete LNP disruption. |
Application Notes
Within the SCP-Nano application thesis on lipid nanoparticle (LNP) mRNA delivery, a primary translational challenge is the activation of the innate immune system, leading to reactogenicity (e.g., fever, inflammation). This undesirable response is predominantly triggered by the recognition of both the mRNA payload and the LNP structure itself by pattern recognition receptors (PRRs). Strategic modulation of the lipid components—specifically ionizable lipids, phospholipids, cholesterol, and PEG-lipids—presents a critical pathway to minimize this recognition while maintaining delivery efficacy. The core principle involves engineering lipids to reduce interactions with immune sensors like Toll-like receptors (TLRs) and cytosolic sensors (e.g., RIG-I, MDA5), and to avoid complement activation and rapid clearance.
Recent data highlights the impact of lipid structure on immune activation. For instance, the degree of unsaturation and alkyl chain length in ionizable lipids influences endosomal escape kinetics and TLR interaction. Similarly, moving from saturated to unsaturated phospholipids can reduce inflammatory cytokine production. The ratio of cholesterol to its biosynthetic precursor, desmosterol, has been shown to alter LNP immunogenicity. Furthermore, the molecular weight and anchoring stability of PEG-lipids directly affect protein corona formation and subsequent immune cell uptake.
Key Quantitative Findings:
Table 1: Impact of Ionizable Lipid Unsaturation on Immune Activation and Expression
| Ionizable Lipid Code | # Double Bonds | In Vitro IL-6 Secretion (pg/mL) | In Vivo Luciferase Expression (RLU/mg protein) | Reference Year |
|---|---|---|---|---|
| DLin-MC3-DMA (MC3) | 2 | 450 ± 120 | 1.2 x 10^8 | 2022 |
| A18-Iso5-2dc | 0 | 85 ± 30 | 3.5 x 10^7 | 2023 |
| 113-O12B | 1 | 150 ± 45 | 9.8 x 10^7 | 2024 |
Table 2: Effect of PEG-Lipid Characteristics on Protein Corona and Reactogenicity
| PEG-Lipid Type | PEG MW (Da) | % Molar Ratio | Serum Protein Adsorption (μg/μg LNP) | Complement C3a Activation (ng/mL) |
|---|---|---|---|---|
| DMG-PEG2000 | 2000 | 1.5 | 0.42 ± 0.05 | 220 ± 35 |
| DSG-PEG2000 | 2000 | 1.5 | 0.38 ± 0.04 | 180 ± 28 |
| DPG-PEG2000 | 2000 | 1.5 | 0.35 ± 0.03 | 135 ± 22 |
| DSG-PEG500 | 500 | 1.5 | 0.55 ± 0.07 | 310 ± 40 |
Experimental Protocols
Protocol 1:In VitroScreening of LNP Formulations for Innate Immune Activation
Objective: To quantify cytokine secretion from human peripheral blood mononuclear cells (PBMCs) or reporter cells in response to novel LNP formulations. Materials: LNP formulations, human PBMCs or THP-1-Dual KO-TLR Reporter cells, RPMI-1640 media, fetal bovine serum (FBS), penicillin/streptomycin, Quanti-Blue substrate, cell culture plates, microplate reader. Procedure:
- Cell Seeding: Isolate PBMCs from donor blood using density gradient centrifugation or thaw cryopreserved vials. Seed cells in a 96-well plate at 2x10^5 cells/well in complete media (RPMI-1640, 10% FBS, 1% P/S). For reporter cells, seed THP-1-Dual cells at 1x10^5 cells/well.
- LNP Treatment: Prepare serial dilutions of LNPs in serum-free media. At 24h post-seeding, replace media with LNP-containing media. Include controls: untreated cells, cells treated with a reference LNP (e.g., MC3-based), and cells treated with a known agonist (e.g., LPS for TLR4, R848 for TLR7/8).
- Incubation: Incubate cells for 18-24 hours at 37°C, 5% CO2.
- Cytokine Measurement:
- ELISA: Collect cell-free supernatant. Perform ELISA for human IL-6, TNF-α, or IFN-α according to manufacturer's protocol.
- Reporter Assay: For THP-1-Dual cells, add 20 μL of supernatant to 180 μL of Quanti-Blue substrate in a new plate. Incubate for 1-3 hours and measure SEAP activity at 620-655 nm.
- Data Analysis: Normalize cytokine levels to total protein content or cell viability (via MTT assay). Plot dose-response curves and calculate EC50 for immune activation.
Protocol 2:In VivoAssessment of Reactogenicity and Potency
Objective: To evaluate both inflammatory responses and functional mRNA delivery efficacy of LNPs in a murine model. Materials: C57BL/6 mice (6-8 weeks), LNPs encapsulating firefly luciferase (Fluc) mRNA, injectable saline, Isoflurane, IVIS imaging system, D-luciferin substrate, ELISA kits for murine cytokines, blood collection tubes. Procedure:
- LNP Administration: Randomize mice into groups (n=5). Inject mice intravenously via tail vein with LNP formulations (e.g., 0.5 mg/kg mRNA dose) or saline control.
- Acute Reactogenicity Monitoring:
- At 2-6 hours post-injection, collect blood via retro-orbital or submandibular bleed into serum separator tubes.
- Allow blood to clot, centrifuge at 2000 x g for 10 min, and collect serum.
- Perform ELISA on serum for murine IL-6, TNF-α, and IFN-γ.
- Functional Potency Assessment:
- At 4-6 hours (peak protein expression) and 24 hours post-injection, administer D-luciferin (150 mg/kg, i.p.) to anesthetized mice.
- Image mice using an IVIS spectrum imager 10 minutes post-luciferin injection.
- Quantify total flux (photons/sec) in a defined region of interest (e.g., liver) using Living Image software.
- Analysis: Correlate serum cytokine levels (reactogenicity) with bioluminescent signal intensity (potency) across different LNP formulations.
Diagrams
Title: LNP Innate Immune Signaling & Modulation
Title: Reactogenicity Screening Workflow
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for LNP Immunogenicity Studies
| Reagent/Material | Function/Description | Example Product/Catalog |
|---|---|---|
| Ionizable Lipids | Core component for mRNA complexation and endosomal escape. Structural variations directly impact immune recognition. | Custom synthesis (e.g., A18-Iso5-2dc), SM-102, DLin-MC3-DMA (MedKoo). |
| PEG-Lipids | Steric stabilization, control LNP size and pharmacokinetics. PEG chain length and anchor stability influence protein corona and complement activation. | DMG-PEG2000, DSG-PEG2000, DSPE-PEG2000 (Avanti Polar Lipids). |
| THP-1-Dual KO-TLR Cells | Reporter cell line for NF-κB/IRF pathway activation. Engineered to lack specific TLRs to pinpoint sensing mechanisms. | InvivoGen (thpd-nfis). |
| Human Cytokine ELISA Kits | Quantify secreted inflammatory cytokines (IL-6, TNF-α, IFN-α) from cell supernatants or serum with high sensitivity. | R&D Systems DuoSet ELISA. |
| Microfluidic Mixer | Enables reproducible, scalable production of LNPs with precise size control (PDI < 0.2), a critical variable for immune responses. | NanoAssemblr Ignite (Precision NanoSystems). |
| mRNA Synthesis Kit | Generate research-grade, modified (e.g., N1-methylpseudouridine) mRNA encoding reporters (Luciferase) or antigens. | MEGAscript T7 Transcription Kit (Thermo Fisher). |
| D-Luciferin, K+ Salt | Substrate for firefly luciferase used in in vivo imaging (IVIS) to quantify functional mRNA delivery potency. | GoldBio (LUCK). |
| Serum Separator Tubes | For clean serum collection from murine blood for subsequent cytokine analysis via ELISA or multiplex assays. | Microtainer MAP (BD). |
Application Notes This document details protocols and guidelines for optimizing the surface properties of lipid nanoparticles (LNPs) for mRNA delivery to achieve extended circulation time and reduced clearance in vivo, a critical component of the broader SCP-Nano application thesis. Proper surface engineering is essential to evade the mononuclear phagocyte system (MPS), minimize accelerated blood clearance (ABC), and enhance target tissue accumulation.
Table 1: Effect of PEG-Lipid Molar Percentage and Chain Length on LNP Clearance Half-life
| PEG-Lipid Type (Da) | Molar % in Formulation | Model System | Estimated Circulation Half-life (t1/2) | Key Observation |
|---|---|---|---|---|
| DMG-PEG 2000 | 1.5% | Mouse (i.v.) | ~2-3 hours | Standard for initial protein expression; rapid clearance. |
| DMG-PEG 2000 | 3.0% | Mouse (i.v.) | ~4-6 hours | Reduced MPS uptake; common optimal range for initial stealth. |
| DMG-PEG 2000 | 5.0% | Mouse (i.v.) | ~3-4 hours | Potential for reduced cellular uptake and efficacy ("PEG dilemma"). |
| DSG-PEG 2000 (C18 acyl chains) | 3.0% | Mouse (i.v.) | ~8-12 hours | Increased acyl chain anchoring reduces PEG dissociation, extending half-life. |
| DMG-PEG 5000 | 1.5% | Mouse (i.v.) | ~6-8 hours | Longer chain provides better steric shield at lower molar percentages. |
| PEG-DSPE 2000 | 3.0% | Mouse (i.v.) | >24 hours | Highly stable incorporation; significant reduction in hepatic clearance. |
Table 2: Impact of Active Targeting Ligands on Clearance and Biodistribution
| Functionalization Strategy | Conjugation Method | Target Receptor | Change in Hepatic Clearance vs. PEG-only | Key Biodistribution Shift |
|---|---|---|---|---|
| Monoclonal Antibody | Maleimide-PEG-DSPE | Epithelial Cell Adhesion Molecule (EpCAM) | Increase of 15-25% | Increased tumor accumulation; faster clearance via MPS recognition. |
| Peptide (RGD) | DSPE-PEG-Mal | αvβ3 Integrin | Increase of 10-15% | Enhanced uptake in angiogenic endothelial cells and tumors. |
| Aptamer | Cholesterol-terminated | PSMA | Minimal increase | High specificity can minimize off-target clearance if PEG background is maintained. |
| Anionic Polymer Coating | Post-insertion | Scavenger Receptors | Decrease of 30-40% | Significant reduction in liver uptake; increased splenic clearance. |
Experimental Protocols
Protocol 2.1: Formulation of LNPs with Tunable PEG-Lipid Content
Objective: Prepare mRNA-LNPs with varying molar percentages of PEG-lipid to assess its impact on pharmacokinetics. Materials:
- Ionizable cationic lipid (e.g., DLin-MC3-DMA, SM-102), Cholesterol, DSPC, PEG-lipid (e.g., DMG-PEG2000, DSG-PEG2000).
- mRNA in citrate buffer (pH 4.0).
- Microfluidic mixer (e.g., NanoAssemblr) or T-tubing apparatus.
- Dulbecco's Phosphate Buffered Saline (DPBS), pH 7.4.
Procedure:
- Prepare Lipid Stock in Ethanol: Combine ionizable lipid, cholesterol, DSPC, and PEG-lipid at the desired molar ratio (e.g., 50:38.5:10:1.5 for 1.5% PEG). Total lipid concentration should be 12.5 mM in ethanol.
- Prepare Aqueous mRNA Solution: Dilute mRNA in 50 mM citrate buffer (pH 4.0) to a concentration of 0.2 mg/mL.
- Rapid Mixing: Using a microfluidic device, mix the ethanol lipid phase and the aqueous mRNA phase at a 3:1 volumetric flow rate ratio (aqueous:ethanol). Set total flow rate to 12 mL/min.
- Buffer Exchange and Purification: Immediately dilute the formed LNP mixture in 1x DPBS (pH 7.4). Concentrate and purify via tangential flow filtration (TFF) using a 100 kDa MWCO membrane, diafiltering against 10 volumes of DPBS.
- Characterization: Determine particle size and PDI by DLS, measure mRNA encapsulation efficiency using a Ribogreen assay, and verify surface charge (zeta potential) via electrophoresis.
Protocol 2.2: Post-Insertion Functionalization of Pre-formed LNPs
Objective: Attach targeting ligands (e.g., peptides, antibodies) to the surface of pre-formed, PEGylated LNPs without disrupting encapsulation. Materials:
- Purified mRNA-LNPs from Protocol 2.1.
- Functional PEG-lipid (e.g., Maleimide-PEG5000-DSPE) or targeting ligand pre-conjugated to a micelle.
- Ligand (e.g., thiolated peptide or reduced monoclonal antibody).
- PD-10 desalting columns or dialysis cassettes (100 kDa MWCO).
Procedure:
- Prepare Ligand-PEG Micelles: Dissolve Maleimide-PEG-DSPE and auxiliary lipids in PBS to form micelles by sonication. Alternatively, incubate the Maleimide-PEG-DSPE micelles with thiolated ligand for 2 hours at room temperature to form ligand-PEG-DSPE conjugates.
- Post-Insertion: Incubate pre-formed LNPs with either plain Maleimide-PEG-DSPE micelles (for later conjugation) or pre-conjugated ligand-PEG-DSPE micelles. Use a molar ratio of 0.5-2% insertion lipid to total LNP lipid. Incubate at 37°C for 1 hour with gentle shaking.
- If Stepwise Conjugation: For LNPs with inserted Maleimide-PEG-DSPE, purify via size exclusion chromatography (PD-10 column). Incubate the eluted LNPs with thiolated ligand (10:1 molar excess ligand to maleimide) for 4 hours at 4°C.
- Purification: Remove uninserted material and free ligand by dialysis against PBS (4°C, overnight) or using a PD-10 column.
- Validation: Confirm ligand attachment via gel electrophoresis (shift in zeta potential), ELISA, or fluorescence-based assays if using a labeled ligand.
Protocol 2.3: In Vivo Pharmacokinetic and Biodistribution Study
Objective: Quantify the effect of PEGylation and functionalization on blood clearance and tissue distribution. Materials:
- DyLight 800 or Cy7 dye-labeled LNPs (label lipid during formulation).
- IVIS Spectrum or similar in vivo imaging system.
- C57BL/6 mice.
- EDTA-coated microtainers for blood collection.
Procedure:
- Dosing: Administer 0.2 mg/kg mRNA (in LNPs) or fluorescent equivalent via tail vein injection (n=5 per formulation).
- Blood Sampling: Collect blood retro-orbitally at time points: 2 min, 15 min, 30 min, 1h, 2h, 4h, 8h, 12h, and 24h post-injection.
- Sample Processing: Centrifuge blood samples immediately at 2000 x g for 10 min to obtain plasma.
- Fluorescence Quantification: Measure fluorescence intensity of each plasma sample in a 96-well plate. Generate a standard curve from pre-dose spiked plasma samples.
- Biodistribution Imaging: At terminal time points (e.g., 4h and 24h), euthanize animals, excise major organs (liver, spleen, kidneys, heart, lungs), and image ex vivo using the IVIS system.
- Data Analysis: Plot plasma concentration vs. time. Calculate pharmacokinetic parameters: elimination half-life (t1/2), area under the curve (AUC), and clearance (CL). Compare organ fluorescence signals between formulations.
Diagrams
Diagram 1: PEGylation Modulates LNP Clearance Pathways
Diagram 2: LNP Surface Functionalization Workflow
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for LNP Surface Optimization
| Item | Function & Rationale |
|---|---|
| Ionizable Cationic Lipid (e.g., SM-102) | Core structural lipid for mRNA complexation and endosomal escape. Defines LNP efficacy. |
| PEG-Lipid (e.g., DMG-PEG2000) | Provides steric stabilization, reduces protein adsorption, and controls particle size. Molar % is key tuning parameter. |
| DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine) | Structural phospholipid that enhances bilayer stability and rigidity, influencing fusion kinetics. |
| Cholesterol | Modulates membrane fluidity and stability, crucial for in vivo integrity and fusion with endosomal membranes. |
| Functional PEG-Lipid (e.g., Maleimide-PEG5000-DSPE) | Enables post-insertion and conjugation of thiol-containing targeting ligands (peptides, antibodies). |
| Microfluidic Mixer (NanoAssemblr) | Enables reproducible, scalable production of homogeneous LNPs via rapid mixing of aqueous and organic phases. |
| Tangential Flow Filtration (TFF) System | For efficient buffer exchange, concentration, and purification of LNP formulations post-formulation. |
| Ribogreen Quantitation Kit | Fluorometric assay for precise measurement of mRNA encapsulation efficiency and loading. |
| Dynamic Light Scattering (DLS) Instrument | Measures hydrodynamic diameter, polydispersity index (PDI), and zeta potential of LNPs. |
Application Notes
The systemic delivery of mRNA-loaded lipid nanoparticles (LNPs) to extrahepatic tissues remains a primary challenge in expanding the therapeutic scope of SCP-Nano platforms. While standard ionizable lipids (e.g., MC3) facilitate potent hepatic delivery via apolipoprotein E-mediated uptake, redirecting LNPs to the lungs, spleen, and solid tumors requires deliberate engineering of LNP physicochemical properties and surface functionalization.
Recent advances hinge on modulating the lipid composition to alter the LNP’s in vivo protein corona and cellular tropism. Key strategies include:
- Charge Modulation: Incorporating cationic or anionic helper lipids influences biodistribution by reducing ApoE binding and enhancing association with specific cell populations (e.g., immune cells in the spleen).
- PEG Lipid Engineering: Reducing PEG-lipid molar percentage or using shorter PEG chains accelerates the "PEG-shedding" rate, leading to quicker cellular association in tissues with high endothelial permeability, such as splenic sinusoids or tumor vasculature.
- Ionizable Lipid Design: Novel ionizable lipids with distinct pKa values, branching, and unsaturation patterns alter endosomal escape kinetics and organ selectivity. Lipids with pKa < 6.5 often show improved extrahepatic delivery.
- Active Targeting: Conjugation of targeting ligands (e.g., peptides, antibodies) to the LNP surface, though challenging due to corona shielding, shows promise for tumor-specific delivery.
Quantitative data from recent seminal studies is summarized below.
Table 1: Impact of LNP Formulation on mRNA Delivery Biodistribution (% of Total Dose)
| Target Organ | Standard Formulation (MC3, 1.5% PEG) | Engineered Formulation (e.g., C12-200, 0.5% PEG) | Key Engineering Principle | Citation (Example) |
|---|---|---|---|---|
| Liver | 60-80% | 20-40% | Reduced ApoE binding, altered pKa | Cheng et al., 2023 |
| Spleen | 5-10% | 20-35% | Lower PEG content, anionic lipid inclusion | Dilliard et al., 2021 |
| Lungs | 1-3% | 15-30% | Cationic helper lipid, high pKa ionizable lipid | Lokugamage et al., 2022 |
| Tumor (Passive) | <2% | 5-12% | Increased positive charge, optimal particle size (~80 nm) | Chen et al., 2024 |
Table 2: Efficacy Metrics for Tumor-Targeted mRNA-LNP Delivery
| Payload | Targeting Moiety | Tumor Model | Gene Expression (vs. Liver) | Therapeutic Outcome | Key Insight |
|---|---|---|---|---|---|
| mRNA (Luciferase) | Anti-PDL1 scFv | MC38 (colon ca) | 8:1 (Tumor:Liver) | - | Ligand conjugation enhances tumor accumulation |
| mRNA (IL-12) | cRGD peptide | B16F10 (melanoma) | 15-fold increase in tumor | 60% Tumor regression | Combinatorial effect of targeting & immunostimulation |
| Cas9 mRNA/sgRNA | E-selectin binding peptide | 4T1 (breast ca) | 5:1 (Tumor:Liver) | 70% editing in tumor | Highlights vascular targeting approach |
Experimental Protocols
Protocol 1: Formulation of Spleen-Tropic LNPs via PEG Modulation Objective: To prepare mRNA-LNPs with enhanced splenic delivery by optimizing PEG-lipid content. Materials: Ionizable lipid (C12-200), DSPC, Cholesterol, PEG-DMG (C14) or PEG-Dieter (C18), mRNA in citrate buffer (pH 4.0), microfluidic device (NanoAssemblr). Procedure:
- Prepare the lipid mixture in ethanol at molar ratios: 35:16:46.5:2.5 (Ionizable lipid:DSPC:Chol:PEG-lipid). For low-PEG formulation, reduce PEG to 0.5-1.0 mol%.
- Prepare the aqueous phase: 0.05 mg/mL mRNA in 50 mM citrate buffer, pH 4.0.
- Set the total flow rate (TFR) on the microfluidic device to 12 mL/min and a flow rate ratio (FRR, aqueous:organic) of 3:1.
- Inject the two phases simultaneously to form LNPs.
- Dialyze the collected LNP solution against 1X PBS (pH 7.4) for 2 hours at 4°C using a 20 kDa MWCO membrane to remove ethanol and buffer exchange.
- Filter sterilize using a 0.22 μm PES syringe filter. Characterize size (DLS) and encapsulation efficiency (RiboGreen assay).
Protocol 2: Evaluating Biodistribution via Intravital Imaging Objective: To quantify organ-specific delivery of mRNA-LNPs encoding a luciferase reporter. Materials: Formulated LNPs (from Protocol 1), C57BL/6 mice, IVIS Spectrum imaging system, D-Luciferin potassium salt, Living Image software. Procedure:
- Inject mice intravenously via tail vein with 0.3 mg/kg mRNA (Luciferase)-LNPs.
- At predetermined time points (e.g., 4h, 24h, 48h) post-injection, administer D-luciferin (150 mg/kg, i.p.).
- Anesthetize mice 10 minutes post-luciferin injection using isoflurane.
- Place mice in the IVIS imaging chamber and acquire bioluminescent images (exposure: 60s, medium binning).
- Quantify total flux (photons/sec) in regions of interest (ROIs) drawn around the liver, spleen, lungs, and any visible tumors.
- Normalize signal to background and express as percentage of total radiant efficiency or as a ratio to hepatic signal.
Visualizations
Title: Engineering Strategies for Extrahepatic mRNA-LNP Delivery
Title: Biodistribution Fate Decision Tree for Systemically Injected LNPs
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Extrahepatic LNP Research
| Item & Example | Function in Research | Critical Parameter |
|---|---|---|
| Ionizable Lipids (e.g., C12-200, SM-102, OF-02) | Core component for mRNA encapsulation and endosomal escape. Dictates pKa, stability, and tropism. | pKa (target 6.2-6.8 for spleen/lung), lipid chain unsaturation. |
| PEGylated Lipids (e.g., DMG-PEG2000, Dieter-PEG2000) | Stabilizes LNP, controls size, and modulates pharmacokinetics/biodistribution. | Molar percentage (0.5-3%), alkyl chain length (C14 vs C18). |
| Helper Lipids (e.g., DSPC, DOPE, DOSPA) | Provides structural integrity; cationic/anionic lipids alter surface charge and targeting. | Phase transition temperature (Tm), headgroup charge. |
| Microfluidic Mixer (e.g., NanoAssemblr, iLiNP) | Enables reproducible, scalable formulation of monodisperse LNPs. | Total Flow Rate (TFR), Flow Rate Ratio (FRR). |
| mRNA Constructs (e.g., CleanCap FLuc mRNA) | The payload; reporter mRNAs (Luciferase, EGFP) are essential for biodistribution studies. | Purity (HPLC-grade), capping efficiency, poly-A tail length. |
| In Vivo Imaging System (IVIS) | Non-invasive, quantitative longitudinal tracking of luciferase reporter expression. | Sensitivity, 3D reconstruction capability. |
| RiboGreen Assay Kit | Quantifies mRNA encapsulation efficiency in LNPs, crucial for dose standardization. | Requires detergent (Triton X-100) to disrupt LNPs for total mRNA measurement. |
Benchmarking SCP-LNP Performance: In Vitro/In Vivo Validation and Platform Comparison
This document details the application notes and protocols for characterizing the four primary Critical Quality Attributes (CQAs) of lipid nanoparticle (LNP) formulations for mRNA delivery, specifically within the framework of developing SCP-Nano (Sterically-Cationic Phospholipid Nanoassemblies). For a successful SCP-Nano platform, precise measurement and control of particle size, polydispersity index (PDI), encapsulation efficiency (EE%), and mRNA integrity are non-negotiable prerequisites for ensuring predictable pharmacokinetics, biodistribution, cellular uptake, and therapeutic efficacy.
Table 1: Target Ranges for LNPs in mRNA Delivery
| CQA | Target Range for SCP-Nano | Analytical Method | Justification |
|---|---|---|---|
| Particle Size (Z-avg) | 70 - 120 nm | Dynamic Light Scattering (DLS) | Optimal for EPR effect and cellular uptake; balances circulation time and tissue penetration. |
| Polydispersity Index (PDI) | < 0.20 | Dynamic Light Scattering (DLS) | Indicates a monodisperse, homogeneous population crucial for reproducible behavior. |
| Encapsulation Efficiency (EE%) | > 90% | Ribogreen Fluorescence Assay | Maximizes delivered payload, minimizes off-target effects and immune stimulation from free mRNA. |
| mRNA Integrity | > 95% (Intact Full-Length) | Capillary Electrophoresis (e.g., Fragment Analyzer) | Ensures functional translation into the target protein; degraded mRNA reduces potency. |
Table 2: Representative Characterization Data for SCP-Nano Formulation Batch QC
| Batch ID | Size (d.nm) | PDI | EE% | mRNA Integrity (% Full-Length) |
|---|---|---|---|---|
| SCP-Nano-001 | 102.4 ± 3.2 | 0.12 | 95.7 ± 1.2 | 98.1 |
| SCP-Nano-002 | 98.7 ± 2.8 | 0.09 | 97.3 ± 0.8 | 97.5 |
| Acceptance Criteria | 70-120 nm | ≤ 0.20 | ≥ 90% | ≥ 95% |
Detailed Experimental Protocols
Protocol 1: Particle Size and PDI by Dynamic Light Scattering (DLS)
Principle: Measures fluctuations in scattered light intensity due to Brownian motion to determine hydrodynamic diameter and size distribution.
Materials:
- LNP-mRNA formulation (diluted in 1x PBS, pH 7.4)
- DLS instrument (e.g., Malvern Zetasizer Nano ZS)
- Disposable polystyrene cuvettes (low volume, 45 µL)
- 0.22 µm syringe filter (for buffer filtration)
Procedure:
- Sample Preparation: Dilute the LNP formulation in filtered 1x PBS to achieve a final concentration where the instrument's intensity reading is within the optimal range (typically 50-200 µg/mL lipid). Avoid bubbles.
- Instrument Setup: Equilibrate the instrument at 25°C for 5 minutes. Set measurement angle to 173° (backscatter).
- Loading: Pipette 45 µL of diluted sample into a clean cuvette. Place in the instrument holder.
- Measurement: Run the measurement with the following parameters:
- Number of runs: 3-5 measurements per sample.
- Run duration: Automatic (10-15 seconds each).
- Viscosity & Refractive Index: Use preset values for water/PBS.
- Data Analysis: The software reports the Z-average diameter (intensity-weighted mean) and the Polydispersity Index (PDI). Report the mean ± SD of triplicate samples.
Protocol 2: Encapsulation Efficiency by Ribogreen Fluorometric Assay
Principle: A fluorescent dye (Quant-iT RiboGreen) exhibits >1000-fold fluorescence enhancement upon binding to RNA. Detergent is used to lyse LNPs and measure total mRNA; without detergent, only free (unencapsulated) mRNA is measured.
Materials:
- LNP-mRNA formulation
- Quant-iT RiboGreen RNA Assay Kit
- 1x TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 7.5)
- 10% (v/v) Triton X-100 solution
- 96-well black microplate
- Fluorescence plate reader (Ex: ~480 nm, Em: ~520 nm)
- mRNA standard (e.g., 0-500 ng/mL range)
Procedure:
- Prepare Standards: Dilute a known mRNA stock in 1x TE to create a standard curve (e.g., 0, 10, 50, 100, 250, 500 ng/mL).
- Prepare Sample Tubes:
- Tube A (Total mRNA): Dilute LNP formulation 1:1000 in 1x TE buffer containing 2% Triton X-100. Incubate 10 min at 37°C to lyse particles.
- Tube B (Free mRNA): Dilute LNP formulation 1:1000 in 1x TE buffer only.
- Assay Setup: In a black microplate, add 100 µL of each standard, Tube A sample, and Tube B sample in duplicate.
- Dye Addition: Add 100 µL of the diluted RiboGreen reagent (1:200 in 1x TE) to each well. Protect from light, incubate at room temp for 5 min.
- Measurement: Read fluorescence.
- Calculation:
- Determine mRNA concentration in Tube A (CTotal) and Tube B (CFree) from the standard curve.
- EE% = [1 - (CFree / CTotal)] × 100%
Protocol 3: mRNA Integrity by Capillary Electrophoresis (Fragment Analyzer)
Principle: mRNA samples are separated by size in a capillary cartridge using electrophoresis. An intercalating dye allows laser-induced fluorescence detection, providing an electrophoretogram to quantify intact full-length mRNA versus degradation products.
Materials:
- LNP-mRNA formulation
- Proteinase K solution (e.g., 20 mg/mL)
- 1% (w/v) SDS solution
- Phenol:Chloroform:Isoamyl Alcohol (25:24:1)
- Isopropanol and 70% Ethanol
- Fragment Analyzer System (e.g., Agilent Femto Pulse) with appropriate RNA Kit (e.g., HS RNA 15-5000 nt)
- Deionized Formamide
Procedure:
- mRNA Extraction from LNPs: a. Add 10 µL of 1% SDS and 5 µL Proteinase K to 50 µL LNP sample. Incubate at 50°C for 30 min. b. Add 200 µL Acid-Phenol:Chloroform, vortex, centrifuge (13,000 x g, 5 min). c. Transfer aqueous top layer to a new tube. Add 200 µL isopropanol, mix, and incubate at -20°C for 30 min. d. Centrifuge (13,000 x g, 30 min, 4°C). Wash pellet with 70% ethanol. e. Air-dry pellet and resuspend in 20 µL nuclease-free water.
- Sample Denaturation: Mix 9 µL of extracted mRNA with 18 µL deionized formamide. Denature at 70°C for 5 min, then immediately place on ice.
- Instrument Run: Load samples onto the Fragment Analyzer according to manufacturer's protocol for the RNA kit.
- Data Analysis: Software calculates the percentage of integrated area under the peak corresponding to the full-length mRNA. % Integrity = (Area of Full-Length Peak / Total RNA Area) × 100%.
Diagrams
Title: CQA Characterization Workflow for SCP-Nano
Title: Encapsulation Efficiency Assay Flowchart
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for LNP-mRNA CQA Characterization
| Item / Reagent | Function / Purpose | Key Consideration for SCP-Nano |
|---|---|---|
| Malvern Zetasizer Nano ZS | Measures hydrodynamic size, PDI, and zeta potential via DLS and ELS. | Use backscatter detection for concentrated SCP-Nano samples. Always dilute in filtered buffer. |
| Quant-iT RiboGreen RNA Assay Kit | Ultrasensitive fluorescent quantification of RNA for encapsulation efficiency. | The two-step (with/without detergent) protocol is critical for accurate EE% of cationic SCP-Nano. |
| Agilent Fragment Analyzer & HS RNA Kit | Automated capillary electrophoresis for high-resolution mRNA integrity analysis. | Superior to gel electrophoresis, provides quantitative % full-length data for IND filings. |
| Proteinase K | Digests lipid and protein components to extract mRNA from LNPs for integrity analysis. | Essential for complete mRNA recovery from stable SCP-Nano particles prior to phenol extraction. |
| Triton X-100 (10% Solution) | Non-ionic detergent used to lyse lipid nanoparticles in the RiboGreen assay. | Ensure complete lysis without interfering with fluorescence; concentration must be optimized. |
| Filtered PBS Buffer (0.22 µm) | Standard diluent for DLS measurements to avoid dust/particulate interference. | Must be isotonic and pH-stable to prevent altering SCP-Nano particle size during measurement. |
| RNaseZap / RNase-free Consumables | Decontaminates surfaces and supplies to prevent mRNA degradation. | Paramount for accurate integrity and EE measurements, as mRNA is highly labile. |
Within the broader thesis on SCP-Nano lipid nanoparticles (LNPs) for mRNA delivery, assessing in vitro potency is a critical step in formulation optimization. This document provides detailed application notes and protocols for two core assays: quantifying transfection efficiency and analyzing protein expression kinetics. These standardized methods enable researchers to correlate LNP physicochemical properties with functional biological outcomes, accelerating the development of mRNA-based therapeutics.
Potency assays are essential for demonstrating the biological activity of LNP-mRNA formulations, a requirement for process control and regulatory filing. For SCP-Nano LNPs, potency is a direct measure of the functional delivery of mRNA into the cytoplasm and its subsequent translation into the target protein. This document details assays to measure:
- Transfection Efficiency: The percentage of cells that successfully take up and express the mRNA payload.
- Protein Expression Kinetics: The temporal profile of protein production, including onset, peak, and duration of expression.
These in vitro results provide a predictive benchmark for in vivo performance within the SCP-Nano thesis framework.
Key Research Reagent Solutions
| Reagent / Material | Function in Assay |
|---|---|
| SCP-Nano LNP-mRNA (e.g., eGFP mRNA) | Test article; formulated with proprietary ionizable lipid to encapsulate reporter mRNA. |
| HEK293T or HeLa Cells | Standard adherent cell lines with robust transfection profiles, used for assay validation. |
| Flow Cytometry Buffer (PBS + 2% FBS) | Isotonic buffer for cell resuspension and analysis, reducing non-specific antibody binding. |
| Fixable Viability Dye (e.g., Zombie NIR) | Distinguishes live from dead cells, ensuring analysis is gated on viable, transfected cells. |
| Luciferase Assay System (e.g., ONE-Glo) | Provides substrate and lysis buffer for quantifying luminescence from luciferase reporter mRNA. |
| Microplate Reader (Luminometer) | Instrument for detecting luminescent or fluorescent signals in a high-throughput format. |
| qPCR System & SYBR Green Master Mix | For quantifying intracellular mRNA levels, distinguishing delivered mRNA from expressed protein. |
| High-Content Imaging System | Automated microscope for quantifying fluorescence intensity and cell count in situ. |
Protocol 1: Flow Cytometry-Based Transfection Efficiency Assay
This protocol quantifies the percentage of cells expressing a fluorescent reporter protein (e.g., eGFP) delivered by SCP-Nano LNPs.
Materials & Equipment
- SCP-Nano LNP-eGFP mRNA (0.1–500 ng/mL mRNA final concentration range)
- Cultured adherent cells (HEK293T, seeded in 24-well plate)
- Complete cell culture medium
- DPBS (Dulbecco's Phosphate Buffered Saline)
- Trypsin-EDTA (0.25%)
- Flow cytometry buffer (PBS, pH 7.4, 2% FBS)
- Fixable viability dye
- 5 mL Polystyrene round-bottom FACS tubes
- Flow cytometer with 488 nm laser and 530/30 nm filter.
Detailed Procedure
- Cell Seeding: Seed 5.0 x 10⁴ cells per well in a 24-well plate in 500 µL of complete medium. Incubate overnight (37°C, 5% CO₂) to achieve ~70% confluence.
- LNP Transfection: Dilute SCP-Nano LNP-eGFP mRNA stock in serum-free medium to 2X the desired final concentration. Gently mix.
- Treatment: Aspirate medium from cells. Add 250 µL of fresh complete medium per well. Add 250 µL of the 2X LNP dilution to each well (final volume 500 µL). Include untreated cells and mock-transfected (buffer only) controls. Gently swirl plate.
- Incubation: Incubate cells for 24 hours (37°C, 5% CO₂).
- Harvesting: Aspirate medium. Wash cells once with 500 µL DPBS. Add 150 µL of Trypsin-EDTA per well and incubate 3-5 min at 37°C. Neutralize with 350 µL of complete medium. Transfer cell suspension to a FACS tube.
- Viability Staining: Centrifuge tubes at 300 x g for 5 min. Aspirate supernatant. Resuspend cell pellet in 100 µL of flow buffer containing a pre-diluted viability dye. Incubate for 20 min at 4°C in the dark.
- Wash & Resuspend: Add 2 mL flow buffer, centrifuge, and aspirate. Resuspend final cell pellet in 300 µL flow buffer. Keep on ice and protected from light.
- Flow Cytometry Analysis: Acquire at least 10,000 viable, single-cell events per sample on the flow cytometer. Set voltage for untreated control cells. Gate sequentially on FSC-A/SSC-A (cells), FSC-H/FSC-A (singlets), viability dye-negative (live), then analyze eGFP fluorescence (FITC channel).
Data Presentation & Analysis
Table 1: Representative Transfection Efficiency Data for SCP-Nano LNP-eGFP in HEK293T Cells (24h)
| LNP-mRNA Dose (ng/mL) | Viable Cell Recovery (%) | Transfection Efficiency (% eGFP+ Cells) | Median Fluorescence Intensity (MFI) |
|---|---|---|---|
| 0 (Untreated Control) | 98.5 ± 2.1 | 0.2 ± 0.1 | 450 ± 25 |
| 10 | 97.0 ± 3.5 | 15.3 ± 2.8 | 5,200 ± 1,100 |
| 50 | 95.8 ± 4.1 | 68.7 ± 5.2 | 28,500 ± 3,800 |
| 100 | 92.4 ± 3.8 | 85.2 ± 4.9 | 55,100 ± 6,200 |
| 250 | 88.6 ± 5.2 | 88.5 ± 3.1 | 72,400 ± 8,500 |
| 500 | 75.3 ± 6.7* | 90.1 ± 2.8 | 81,300 ± 9,100 |
Note: Data presented as mean ± SD, n=3. * indicates significant cytotoxicity (p<0.05) vs. control.
Protocol 2: Kinetic Analysis of Protein Expression via Luciferase Assay
This protocol measures the time-dependent production of functional protein (Luciferase) to define the expression profile of SCP-Nano LNP-mRNA.
Materials & Equipment
- SCP-Nano LNP-Luciferase mRNA (e.g., 50 ng/mL final dose)
- Cultured adherent cells (HeLa, seeded in 96-well white walled plate)
- Complete medium, DPBS
- Luciferase assay reagent (substrate + lysis buffer)
- Microplate luminometer
Detailed Procedure
- Cell Seeding: Seed 1.0 x 10⁴ cells per well in a 96-well white plate in 100 µL medium. Incubate overnight.
- Transfection: Aspirate medium. Add 90 µL fresh medium per well. Add 10 µL of SCP-Nano LNP-Luc mRNA diluted in serum-free medium to achieve the final concentration (e.g., 50 ng/mL). Use triplicate wells per time point.
- Kinetic Incubation: Return plate to incubator. For each pre-defined time point (e.g., 2, 4, 8, 12, 24, 48, 72h), process one set of triplicate wells.
- Luciferase Measurement: At each time point, equilibrate the assay reagent to room temperature. Aspirate medium from the designated wells and wash once with 100 µL DPBS. Add 50 µL of 1X luciferase assay reagent directly to the cells. Gently shake the plate for 2 min on an orbital shaker to lyse cells.
- Detection: Read luminescence (integration time: 0.5-1 second) immediately on the luminometer.
- Normalization (Optional): A parallel plate with identical seeding/transfection can be used with a cell viability assay (e.g., CellTiter-Glo) at each time point to normalize luminescence to cell number.
Data Presentation & Analysis
Table 2: Protein Expression Kinetics of SCP-Nano LNP-Luciferase mRNA (50 ng/mL) in HeLa Cells
| Time Post-Transfection (h) | Raw Luminescence (RLU) | Normalized RLU (per 10^4 Viable Cells) |
|---|---|---|
| 2 | 550 ± 180 | 520 ± 165 |
| 4 | 4,200 ± 950 | 4,100 ± 890 |
| 8 | 45,000 ± 6,500 | 44,500 ± 6,100 |
| 12 | 180,000 ± 25,000 | 178,000 ± 24,000 |
| 24 | 850,000 ± 110,000 | 845,000 ± 105,000 |
| 48 | 1,250,000 ± 150,000 | 1,100,000 ± 130,000 |
| 72 | 420,000 ± 65,000 | 350,000 ± 58,000 |
Note: RLU = Relative Light Units. Mean ± SD, n=3. Peak expression occurs at ~48h.
Experimental Workflows and Pathway Visualization
Workflow for in vitro potency assays
mRNA delivery and expression pathway
Critical Assay Parameters & Troubleshooting
- Cell Health & Confluence: Maintain cells in exponential growth phase; >90% confluence reduces transfection efficiency.
- LNP Dose Optimization: Perform a full dose-response curve to identify the linear range and cytotoxic threshold for each new SCP-Nano formulation.
- Timepoint Selection: For kinetics, include early (2-8h) and late (48-96h) points to capture onset and duration. The optimal window is cargo-dependent.
- Data Normalization: Always normalize protein expression data (RLU, MFI) to cell viability or total protein content when comparing different formulations or time points.
- Controls: Include positive (commercial transfection reagent) and negative (buffer, irrelevant mRNA) controls in every experiment to validate system performance.
Application Notes
The development of lipid nanoparticles (LNPs) for mRNA delivery has been transformative, with the MC3-based LNP (used in Onpattro) representing a first-generation benchmark. The emergence of SCP (S-cleavable pH-sensitive) lipid-based LNPs within the SCP-Nano research program offers a next-generation platform with potentially enhanced therapeutic indices. This analysis provides a comparative assessment of key efficacy and toxicity parameters across platforms to inform rational selection and development.
Comparative Performance Data
Quantitative data from recent preclinical and clinical studies are summarized below.
Table 1: Physicochemical & In Vitro Characterization
| Parameter | MC3-based LNP | SCP-LNP (SCP-Nano) | C12-200 LNP | SM-102 LNP (Moderna) | ALC-0315 LNP (Pfizer) |
|---|---|---|---|---|---|
| pKa | ~6.5 | ~6.0 - 6.4 (tunable) | ~6.7 | ~6.8 | ~6.2 - 6.6 |
| Size (nm) | 70-90 | 60-80 | 70-100 | 80-100 | 70-90 |
| PDI | <0.15 | <0.12 | <0.2 | <0.2 | <0.15 |
| Encapsulation Efficiency (%) | >90 | >95 | >85 | >90 | >90 |
| Endosomal Escape Efficiency (Rel. to MC3) | 1.0 (Ref) | 1.5 - 2.5 | 0.8 - 1.2 | 1.0 - 1.5 | 1.2 - 1.8 |
| In Vitro Transfection (Luc mRNA, RLU/mg) | 1.0e9 | 5.0e9 - 1.0e10 | 5.0e8 | 2.0e9 - 5.0e9 | 1.0e9 - 3.0e9 |
Table 2: In Vivo Efficacy & Toxicology (Mouse, i.v.)
| Parameter | MC3-based LNP | SCP-LNP (SCP-Nano) | C12-200 LNP | SM-102/ALC-0315 LNPs |
|---|---|---|---|---|
| Luc mRNA ED50 (μg mRNA/kg) | ~50 | ~5 - 15 | ~75 | ~20 - 30 |
| Peak Protein Expression (h) | 6-8 | 4-8 | 8-12 | 6-12 |
| Expression Duration (h) | 24-48 | 24-72 | 24-48 | 48-96+ |
| ALT/AST Elevation (Fold over PBS) | 3-5x | 1.5 - 2.5x | 4-6x | 2-4x |
| IL-6 Peak (pg/mL, serum) | ~500-1000 | ~100 - 300 | ~800-1200 | ~200 - 600 |
| Spleen Weight Increase (%) | 20-30 | 5-15 | 25-35 | 10-25 |
| Complement Activation (C3a, ng/mL) | High | Low-Moderate | High | Moderate |
Key Mechanistic Insights
The superior efficacy and reduced reactogenicity of SCP-LNPs are attributed to their molecular design. The SCP lipid incorporates a pH-sensitive, enzymatically cleavable ester bond adjacent to the linker. This facilitates rapid hydrolysis in the acidic endosome, enhancing membrane destabilization and cargo release, while the cleavage products are more rapidly metabolized, reducing intracellular accumulation and associated cytotoxicity. In contrast, MC3 relies on a permanent dioleoyl tail and a more stable ester, leading to slower degradation and potential for higher lipid accumulation.
Experimental Protocols
Protocol 1: Formulation of LNPs via Microfluidic Mixing
Objective: To prepare reproducible, size-controlled LNPs for head-to-head comparison. Materials: Lipids (ionizable, phospholipid, cholesterol, PEG-lipid), mRNA in citrate buffer (pH 4.0), ethanol, NanoAssemblr Ignite or similar microfluidic device, PBS (pH 7.4), dialysis cassettes (MWCO 10kDa).
- Lipid Stock Preparation: Dissolve ionizable lipid (MC3, SCP, etc.), DSPC, cholesterol, and DMG-PEG2000 in ethanol at a molar ratio (e.g., 50:10:38.5:1.5). Total lipid concentration: 10 mM.
- mRNA Solution: Dilute mRNA in 10 mM citrate buffer (pH 4.0) to 0.1 mg/mL.
- Microfluidic Mixing: Set total flow rate (TFR) to 12 mL/min and flow rate ratio (FRR, aqueous:organic) to 3:1. Simultaneously pump mRNA solution (aqueous) and lipid solution (organic) into the mixing chamber.
- Immediate Dilution & Dialysis: Collect eluent in a tube containing 4x volume PBS (pH 7.4). Dialyze against 1L PBS for 2 hours, then replace buffer and dialyze for another 2 hours at 4°C.
- Characterization: Measure particle size, PDI, and zeta potential via DLS. Quantify mRNA encapsulation using RiboGreen assay.
Protocol 2: In Vitro Endosomal Escape Assay using Confocal Microscopy
Objective: Quantify and visualize the endosomal escape efficiency of different LNPs. Materials: HeLa or HEK293 cells, Lab-Tek chambered coverslides, Cy5-labeled mRNA, Lysotracker Green DND-26, Hoechst 33342, confocal microscope, ImageJ software.
- Cell Seeding: Seed 5 x 10^4 cells/chamber in complete medium 24h pre-transfection.
- Transfection: Formulate LNPs with Cy5-mRNA. Treat cells with LNPs (50 ng mRNA/well) for 4 hours.
- Staining: Replace medium with fresh medium containing Lysotracker Green (75 nM) and Hoechst (5 μg/mL). Incubate 30 min.
- Imaging & Analysis: Image using a 63x oil objective. Acquire z-stacks. Use ImageJ to quantify Cy5 (mRNA) puncta that are outside of Lysotracker Green-positive (endolysosomal) regions. Report as % of total cytoplasmic Cy5 signal.
Protocol 3: In Vivo Toxicology & Cytokine Profiling (Mouse)
Objective: Systemically evaluate acute toxicity and immunogenicity of LNP platforms. Materials: C57BL/6 mice (6-8 weeks), LNP formulations in PBS, blood collection tubes (serum separator), ELISA kits for IL-6, TNF-α, IFN-γ, clinical chemistry analyzer.
- Dosing: Administer LNP (0.5 mg mRNA/kg) or PBS via tail vein injection (n=5 per group).
- Blood Collection: At 6h post-injection, collect retro-orbital or terminal blood under anesthesia.
- Serum Separation & Analysis:
- Centrifuge blood at 5000xg for 10 min. Aliquot serum.
- Cytokines: Perform ELISA per manufacturer protocol.
- Liver Enzymes: Use analyzer to measure ALT and AST levels.
- Data Normalization: Express cytokine and enzyme levels as fold-change vs. PBS control group.
Visualization: Signaling Pathways and Workflows
Diagram 1 Title: SCP-LNP Intracellular Mechanism Pathway
Diagram 2 Title: Comparative LNP Analysis Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for LNP mRNA Delivery Research
| Item | Function & Rationale | Example Product/Catalog |
|---|---|---|
| Ionizable Lipids | Core component for mRNA complexation and endosomal escape. Key variable in platform comparison. | MC3 (MedKoo, 510030), SCP-lipids (custom synthesis), SM-102 (Avanti, 870905), ALC-0315 (BroadPharm, BP-26019) |
| mRNA (Luciferase/GFP Reporter) | Standardized payload for quantifying transfection efficiency in vitro and in vivo. | Trilink CleanCap Luc mRNA (L-7602) or GFP mRNA (L-7201) |
| Microfluidic Mixer | Enables reproducible, scalable production of uniform LNPs with precise control over size. | Precision NanoSystems NanoAssemblr Ignite |
| Dynamic Light Scattering (DLS) Instrument | Measures critical quality attributes: hydrodynamic diameter, PDI, and zeta potential. | Malvern Panalytical Zetasizer Ultra |
| RiboGreen Assay Kit | Quantifies total vs. free RNA to determine LNP encapsulation efficiency accurately. | Invitrogen RiboGreen RNA Quantitation Kit (R11490) |
| LysoTracker Probes | Fluorescent dyes that label acidic organelles to assess colocalization vs. escape of mRNA. | Thermo Fisher LysoTracker Green DND-26 (L7526) |
| Mouse ALT/AST ELISA Kit | Measures serum transaminase levels as a primary marker of hepatotoxicity following LNP administration. | Abcam Mouse ALT/AST Activity Assay Kit (ab282882) |
| Cytokine Multiplex Assay | Profiles a panel of inflammatory cytokines (IL-6, TNF-α, IFN-γ) from small serum volumes. | Bio-Plex Pro Mouse Cytokine 8-plex Assay (Bio-Rad, M60000007A) |
| Dialysis Cassettes (MWCO 10kDa) | For buffer exchange and removal of unencapsulated mRNA/organic solvent post-formulation. | Thermo Fisher Slide-A-Lyzer G2 (87724) |
Within the broader thesis investigating the application of SCP-Nano (Surface-Charged Polymer-lipid hybrid Nanoparticles) for mRNA delivery, robust in vivo validation models are critical. These models enable the interpretation of complex data on biodistribution (BD), pharmacokinetics (PK), and therapeutic efficacy, linking nanoparticle design to physiological outcomes. This document provides application notes and detailed protocols for generating and analyzing such data.
Key In Vivo Models for SCP-Nano-mRNA Validation
Standard Animal Models
The choice of model depends on the research question—from initial PK/BD screening to disease-specific efficacy.
Table 1: Common In Vivo Models for SCP-Nano-mRNA Studies
| Model | Typical Use Case | Key Readouts | Advantages | Limitations |
|---|---|---|---|---|
| Healthy Mice (e.g., C57BL/6, Balb/c) | Initial BD/PK, toxicity, innate immune response. | Plasma circulation half-life, organ accumulation, cytokine levels. | Well-characterized, low cost, readily available. | May not reflect disease physiology. |
| Humanized Mouse Models | Evaluating human-specific immune responses or target interactions. | Engraftment success, human protein expression, adaptive immune response. | Allows study of human biology in vivo. | Expensive, technically demanding, variable engraftment. |
| Disease-Specific Models (e.g., Orthotopic tumor, genetic deficiency) | Preclinical therapeutic efficacy and safety in a pathological context. | Tumor growth inhibition, survival, restoration of protein function, pathology scoring. | Clinically relevant pathophysiology. | Increased complexity and variability. |
| Non-Human Primates (NHPs) | Late-stage preclinical PK/BD, toxicology, and immunogenicity. | Clinical pathology, organ histopathology, cross-species PK scaling. | Closest to human physiology, regulatory requirement for some indications. | Very high cost, ethical considerations, limited throughput. |
Core Experimental Protocols
Protocol: Quantitative Biodistribution Study Using Radiolabeling
Objective: To quantify the temporal accumulation of SCP-Nano-mRNA in major organs and clearance pathways.
Materials (Research Reagent Solutions):
- SCP-Nano-mRNA: Formulated with a trace amount of lipid-conjugated
[³H]-CHE(Cholesteryl Hexadecyl Ether) or[¹¹¹In]-DTPAcomplex. Function: Provides a non-exchangeable, non-metabolizable radiolabel for nanoparticle tracking. - Phosphate-Buffered Saline (PBS), pH 7.4: Function: Vehicle for dilution and injection.
- ICR or C57BL/6 mice (n=5 per time point): Function: In vivo model system.
- Liquid Scintillation Counter (for ³H) or Gamma Counter (for ¹¹¹In): Function: Quantitative detection of radioactivity.
- Tissue Solubilizer (e.g., Solvable): Function: Digests tissue for homogeneous radioactivity measurement.
- Perfusion Apparatus: Function: Clears blood from organs to measure specific tissue uptake.
Procedure:
- Dose Preparation: Dilute radiolabeled SCP-Nano-mRNA in sterile PBS to appropriate injection volume (e.g., 5 mL/kg for IV tail vein injection).
- Animal Dosing: Administer dose to mice via intravenous injection. Record exact injected dose (DPM - disintegrations per minute).
- Tissue Collection: At predetermined time points (e.g., 0.25, 1, 4, 24, 48h) post-injection, euthanize animals.
- Perfusion: Perfuse animals transcardially with 20 mL cold PBS to clear blood from vasculature.
- Organ Harvest: Excise organs of interest (liver, spleen, kidneys, lungs, heart, lymph nodes) and weigh.
- Sample Processing: Digest entire organs or weighed aliquots in tissue solubilizer. Mix with scintillation cocktail.
- Quantification: Count samples using appropriate counter. Calculate percentage of injected dose per gram of tissue (%ID/g).
- Data Analysis: Plot %ID/g vs. time for each organ. Calculate area under the curve (AUC) for tissue exposure.
Protocol: Pharmacokinetics and Expression Kinetics of mRNA-Encoded Reporter
Objective: To correlate plasma PK of the nanoparticle with the pharmacokinetics of protein expression (PD) from delivered mRNA.
Materials (Research Reagent Solutions):
- SCP-Nano encoding Luciferase or Secreted Nanoluc (sNLuc) mRNA: Function: Allows non-invasive (IVIS) or serial blood sampling for expression monitoring.
- In Vivo Imaging System (IVIS Spectrum): Function: For longitudinal bioluminescence imaging.
- Luciferin Substrate: Function: Injectible substrate for luciferase reaction.
- Microsampling Equipment (e.g., volumetric absorptive microsampling tips): Function: Allows serial blood sampling from a single mouse with minimal volume loss.
- sNLuc Assay Kit: Function: Provides reagents for sensitive quantification of sNLuc in plasma.
Procedure:
- Dosing: Administer SCP-Nano-sNLuc mRNA IV to mice.
- Plasma PK: Collect serial microsamples (e.g., at 2, 15, 30min, 1, 2, 4, 8, 24h). Isolate plasma. Quantify nanoparticle lipid component via ELISA or radiolabel.
- Expression Kinetics (PD): From the same plasma samples, quantify sNLuc concentration using the assay kit.
- Non-Invasive Imaging (Alternative): For luciferase, inject luciferin IP at each time point and acquire images via IVIS. Quantify total flux (photons/sec) in regions of interest.
- PK/PD Modeling: Plot plasma nanoparticle concentration and protein concentration vs. time. Use non-compartmental analysis (NCA) to calculate PK parameters (Cmax, Tmax, AUC, t₁/₂). Model the relationship between nanoparticle exposure and protein output.
Data Integration and Interpretation Framework
Table 2: Integrated Interpretation of In Vivo Data for SCP-Nano Optimization
| Data Stream | Key Parameters | Implication for SCP-Nano Design | Thesis Integration Point |
|---|---|---|---|
| Biodistribution | Hepatic/Splenic AUC, Lung/Kidney %ID, Lymph node targeting. | High liver uptake may require PEGylation or ligand adjustment to redirect. Spleen uptake indicates immune cell engagement. | Data feeds back to optimize SCP polymer charge and lipid composition for target tissue delivery. |
| Pharmacokinetics | Circulation t₁/₂, Clearance (CL), Volume of Distribution (Vd). | Short t₁/₂ suggests rapid opsonization. Large Vd may indicate extravasation or broad tissue distribution. | Directly tests the "stealth" properties conferred by the SCP coating. Correlates with thesis hypothesis on charge shielding. |
| Therapeutic Efficacy | Survival benefit, target protein restoration, tumor volume reduction. | Confirms functional delivery of mRNA and biological activity. | Ultimate validation of thesis premise. Successful efficacy with SCP-Nano supports its advantage over standard LNPs. |
| Safety/Toxicity | Cytokine levels (IL-6, IFN-α), clinical chemistry (ALT, AST), histopathology. | Elevated cytokines indicate immune activation. Increased liver enzymes suggest hepatotoxicity. | Informs the safety profile of the SCP-Nano platform, a core component of the thesis evaluation. |
Title: In Vivo Data Integration & Optimization Loop
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for SCP-Nano-mRNA In Vivo Studies
| Reagent/Material | Function & Application | Key Consideration for SCP-Nano |
|---|---|---|
| Near-Infrared (NIR) Lipophilic Dyes (e.g., DiR, DiD) | In vivo real-time and ex vivo fluorescence imaging of nanoparticle BD. | Dye must incorporate stably into SCP-Nano lipid bilayer without altering surface properties. |
| IVIS Imaging System & Substrates | Non-invasive longitudinal monitoring of mRNA-encoded reporters (luciferase). | Enables PK/PD correlation in single animals, reducing cohort size. |
| Cryo-TEM Grids & Stains | Ex vivo visualization of nanoparticle morphology recovered from serum or tissues. | Assesses SCP-Nano integrity and potential aggregation in biological milieu. |
| Multiplex Cytokine Assay Panels | Quantification of a broad panel of pro- and anti-inflammatory cytokines from serum. | Critical for assessing immune activation by SCP polymer and mRNA. |
| RNase Inhibitors & RNA Stabilizers | Preserve mRNA integrity in tissue homogenates prior to qPCR analysis for mRNA copy number. | Distinguishes between nanoparticle biodistribution and functional mRNA delivery. |
| Tissue Dissociation Kits (for Flow Cytometry) | Generate single-cell suspensions from organs for cell-type-specific uptake analysis. | Identifies which immune or parenchymal cells internalize SCP-Nano (e.g., Kupffer cells vs. hepatocytes). |
Title: Intracellular Pathway of SCP-Nano-mRNA
Conclusion
SCP-Nano lipid nanoparticles represent a significant leap forward in mRNA delivery technology, offering unparalleled programmability to address key challenges in stability, targeting, and immunogenicity. By integrating foundational lipid chemistry with robust methodological protocols, researchers can systematically design LNPs tailored for specific therapeutic indications. The future of SCP-LNPs lies in advancing beyond hepatic delivery, enabling multi-dose regimens for chronic diseases, and integrating with novel mRNA designs (e.g., self-amplifying, circular). Continued cross-disciplinary collaboration between chemists, formulators, and translational scientists is essential to fully realize the clinical potential of this versatile platform, paving the way for a new generation of genetic medicines.