Nanopipette SERS Probes: Revolutionizing Single-Cell Hemeprotein Analysis for Biomedical Research
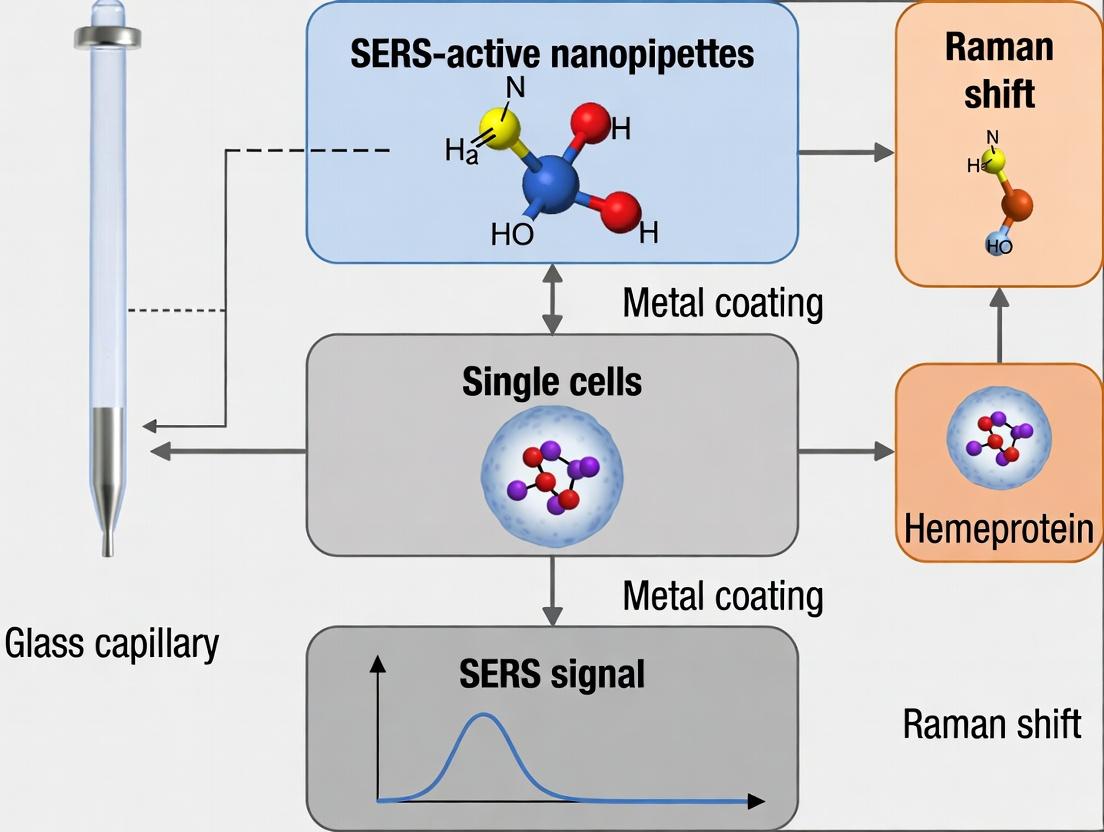
This article provides a comprehensive examination of Surface-Enhanced Raman Scattering (SERS)-active nanopipettes as transformative tools for detecting hemeproteins in single cells.
Nanopipette SERS Probes: Revolutionizing Single-Cell Hemeprotein Analysis for Biomedical Research
Abstract
This article provides a comprehensive examination of Surface-Enhanced Raman Scattering (SERS)-active nanopipettes as transformative tools for detecting hemeproteins in single cells. Targeted at researchers, scientists, and drug development professionals, we explore the foundational principles of nanopipette fabrication and SERS enhancement. The content details step-by-step methodologies for functionalization and intracellular application, addresses critical troubleshooting and optimization strategies for signal consistency and cell viability, and validates the technique against established methods like mass spectrometry and fluorescence. The discussion concludes by synthesizing the technique's potential to uncover cellular heterogeneity and its implications for advancing diagnostics and therapeutic development.
Understanding SERS-Active Nanopipettes: Principles, Fabrication, and Hemeprotein Specificity
Application Notes
The integration of nanopipette technology with Surface-Enhanced Raman Scattering (SERS) represents a paradigm shift in single-cell analysis, particularly for the detection of low-abundance hemeproteins like cytochrome c, hemoglobin, and catalase. This combination merges the spatial precision and minimally invasive sampling of nanopipettes with the exceptional molecular specificity and single-molecule sensitivity of SERS.
Key Advantages:
- Targeted Intracellular Probing: Nanopipettes (tip diameter: 50-300 nm) enable precise insertion into single cells with minimal disruption, allowing for localized sampling or delivery of SERS-active nanoparticles (e.g., Au/Ag) directly to organelles like mitochondria.
- Amplified Specific Signals: SERS enhancement factors of 10⁶–10¹⁰ allow for the detection of the characteristic vibrational fingerprints of hemeproteins, even at single-molecule levels, overcoming the weak intrinsic Raman signal.
- Dynamic Monitoring: The platform facilitates real-time, label-free monitoring of redox state changes, ligand binding, or degradation of hemeproteins within living cells, crucial for studying apoptosis, metabolic dysfunction, and drug response.
Quantitative Performance Metrics:
Table 1: Performance Metrics of SERS-Active Nanopipettes for Hemeprotein Detection
| Parameter | Typical Range/Value | Notes |
|---|---|---|
| Nanopipette Tip Diameter | 50 – 300 nm | Balances intracellular access with sufficient flow for analyte/nanoparticle transport. |
| SERS Enhancement Factor (EF) | 10⁷ – 10⁹ | Achieved with optimized AuNP aggregates or nanostructured pipette tips. |
| Limit of Detection (LOD) for Cytochrome c | 1 – 10 nM (in vitro); Single-molecule (in situ) | In-cell LOD is dependent on localization efficiency. |
| Spatial Resolution (Intracellular) | < 1 µm | Defined by the nanopipette tip and diffusion zone of injected SERS tags. |
| Measurement Temporal Resolution | 100 ms – 10 s | Sufficient for tracking kinetic processes in live cells. |
| Cell Viability Post-Measurement | > 90% (1 hour) | Confirms minimally invasive nature of the technique. |
Table 2: Characteristic SERS Peaks for Key Hemeproteins
| Hemeprotein | SERS Peak Position (cm⁻¹) | Vibrational Mode Assignment |
|---|---|---|
| Cytochrome c (Oxidized) | 750, 1132, 1585 | ν₁₅, ν₅, ν₂ (Porphyrin macrocycle) |
| Cytochrome c (Reduced) | 746, 1124, 1560 | Shift indicates redox state change. |
| Hemoglobin | 675, 755, 1375 | ν₇, ν₁₅, ν₄ (Oxidation state marker band) |
| Myoglobin | 698, 1130, 1395 | Porphyrin ring vibrations. |
Detailed Experimental Protocols
Protocol 1: Fabrication of SERS-Active Nanopipettes
Objective: To create a nanopipette with integrated SERS hotspots at its tip for localized sensing.
Materials:
- Quartz or borosilicate capillaries (OD: 1.0 mm, ID: 0.7 mm)
- Laser-based pipette puller (e.g., Sutter Instruments P-2000)
- Gold nanoparticle (AuNP) colloid (60 nm diameter)
- (3-Aminopropyl)triethoxysilane (APTES)
- Raman microscope with 633 nm or 785 nm laser excitation
Procedure:
- Pulling: Pull a capillary using a programmed laser puller to achieve a tip diameter of ~100 nm. Validate tip geometry via SEM.
- Silanzation: Vapor-phase silanization with APTES for 30 minutes to create a positively charged amine-terminated surface on the inner wall near the tip.
- Functionalization: Back-fill the nanopipette with a dilute AuNP colloid (OD520 ~ 0.5). Allow the negatively charged AuNPs to electrostatically assemble onto the aminated surface for 1 hour.
- Rinsing: Gently flush the pipette with deionized water to remove unbound AuNPs, leaving a stable aggregate at the tip serving as a SERS hotspot.
- Characterization: Confirm SERS activity by immersing the tip in a 1 mM solution of benzenethiol and acquiring a spectrum.
Protocol 2: Intracellular Hemeprotein Sensing in a Single Live Cell
Objective: To detect and monitor the redox state of cytochrome c within the cytoplasm of a living mammalian cell.
Materials:
- SERS-active nanopipette (from Protocol 1)
- Cell line (e.g., HeLa, MCF-7) cultured on a glass-bottom dish
- Extracellular buffer (e.g., PBS or HEPES-buffered saline)
- Patch-clamp micromanipulator and amplifier
- Confocal Raman microscope system
Procedure:
- Cell Preparation: Culture cells on a sterilized glass-bottom dish. On the day of the experiment, replace medium with extracellular buffer.
- Pipette Positioning: Mount the SERS-active nanopipette on the micromanipulator. Using optical guidance (40x objective), position the pipette tip in close proximity to the target cell.
- Penetration: Apply a slight positive pressure (5-10 hPa) to the pipette and advance it swiftly to penetrate the cell membrane. Release pressure upon entry. Monitor cell morphology to ensure viability.
- SERS Acquisition: Position the laser focus (~1 µm spot) onto the nanopipette tip inside the cell. Acquire SERS spectra (integration time: 1-5 s, laser power: 1-5 mW at sample) from the cytoplasmic region.
- Stimulation & Monitoring: Introduce an apoptotic inducer (e.g., 1 µM staurosporine) to the bath. Continuously acquire SERS spectra every 30 seconds for 30 minutes to monitor the shift of the ν₄ band from ~1585 cm⁻¹ (oxidized) to ~1560 cm⁻¹ (reduced), indicating cytochrome c release and reduction.
- Data Analysis: Process spectra (baseline correction, smoothing). Plot the intensity ratio I₁₅₆₀/I₁₅₈₅ over time to visualize the kinetic redox change.
Diagrams
SERS-Active Nanopipette Single-Cell Analysis Workflow
SERS Detection of Cytochrome c in Apoptosis Pathway
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for SERS-Nanopipette Experiments
| Item | Function / Purpose |
|---|---|
| Quartz Capillaries | Provide low background Raman signal and excellent optical properties for tip-integrated SERS measurements. |
| Gold Nanoparticles (60-80 nm) | Act as plasmonic nanoantennas. Their aggregation at the nanopipette tip creates intense electromagnetic fields for SERS signal amplification. |
| APTES (Silane Coupling Agent) | Functionalizes the silica surface of the pipette interior to facilitate electrostatic assembly of AuNPs. |
| Benzenthiol / 4-MBA | Model Raman reporter molecules used for standardizing and calibrating the SERS activity of fabricated nanopipettes. |
| Cytochrome c (Purified) | Essential positive control for in vitro optimization of SERS spectra acquisition and identification of characteristic peak patterns. |
| Staurosporine / Etoposide | Apoptosis-inducing drugs used in live-cell experiments to trigger mitochondrial cytochrome c release and create a dynamic biological model. |
| HEPES-Buffered Saline | Provides a stable, CO2-independent ionic environment for live-cell experiments outside an incubator. |
| Cell-Permeant Vital Dyes | Used to visually confirm cell viability before, during, and after nanopipette insertion. |
Application Notes for SERS-Active Nanopipette Fabrication
This protocol details the synthesis of Surface-Enhanced Raman Scattering (SERS)-active nanopipettes for the intracellular detection of hemeproteins, such as cytochrome c and hemoglobin. The approach integrates quartz nanopipettes with plasmonic noble metal nanostructures and selective surface chemistry to create a robust single-cell analytical probe.
Key Advantages:
- High Spatial Resolution: Tip diameters of 50-200 nm enable minimal invasive cellular entry.
- Enhanced Sensitivity: Localized Surface Plasmon Resonance (LSPR) of Au/Ag nanostructures provides ~10⁶-10⁸ signal enhancement.
- Single-Cell Compatibility: Allows for real-time, in situ monitoring of hemeprotein redox states and concentrations within living cells.
Core Quantitative Data:
Table 1: Performance Metrics of SERS-Active Nanopipettes for Hemeprotein Detection
| Parameter | Typical Value/Range | Notes |
|---|---|---|
| Nanopipette Tip Diameter (ID) | 50 - 200 nm | Controlled by laser puller parameters. |
| Ag Nanostructure Thickness | 30 - 60 nm | Optimal for 532 nm & 633 nm excitation. |
| Au Nanoparticle Diameter | 40 - 80 nm | Conjugated to functionalization layer. |
| Enhancement Factor (EF) | 1 x 10⁶ - 5 x 10⁷ | Calculated using benzenthiol or crystal violet. |
| Limit of Detection (Cytochrome c) | 10 - 100 nM | In buffer solution. |
| Cellular Insertion Viability | >85% (post 1 hr) | Assessed by trypan blue exclusion. |
| SERS Mapping Resolution | < 300 nm | Lateral resolution within a cell. |
Table 2: Essential Materials & Functionalization Reagents
| Material/Reagent | Function & Role in Protocol | Example Product/Catalog # |
|---|---|---|
| Quartz Capillary (with filament) | Nanopipette substrate; provides optical clarity and rigidity. | Sutter Instrument QF100-70-7.5 |
| Silver Wire (0.5 mm diameter, 99.99%) | Anode for Ag nanostructure deposition via electrochemical plating. | Alfa Aesar 44482 |
| Hydrogen Tetrachloroaurate(III) Trihydrate (HAuCl₄·3H₂O) | Precursor for synthesis of Au nanoparticles (AuNPs). | Sigma-Aldrich 520918 |
| (3-Aminopropyl)triethoxysilane (APTES) | Silane coupling agent for introducing amine groups to quartz surface. | Sigma-Aldrich 440140 |
| Glutaraldehyde (25% solution) | Crosslinker for conjugating amine-modified surfaces to protein capture agents. | Thermo Fisher Scientific PI28364 |
| Anti-cytochrome c Antibody (monoclonal) | Capture agent for specific hemeprotein recognition. | Abcam ab13575 |
| Methylene Blue | Raman reporter and electrochemical validation molecule. | Sigma-Aldorych M9140 |
| Phosphate Buffered Saline (PBS), pH 7.4 | Buffer for all functionalization and washing steps. | Gibco 10010023 |
Detailed Experimental Protocols
Protocol 1: Fabrication of Quartz Nanopipettes
Objective: Pull quartz capillaries to form nanoscale tips.
- Preparation: Place a quartz capillary (QF100-70-7.5) into the holder of a laser-based pipette puller (e.g., Sutter P-2000).
- Pulling Parameters: Use a two-line program: Line 1: Heat = 750, Fil = 4, Vel = 30, Del = 145, Pul = 90. Line 2: Heat = 730, Fil = 3, Vel = 25, Del = 135, Pul = 135. These parameters produce tips with ~100 nm inner diameter.
- Validation: Image tips using SEM or TEM to confirm diameter and geometry.
Protocol 2: Electrochemical Deposition of Silver Nanostructures
Objective: Create a rough, plasmonically active Ag layer on the nanopipette tip exterior.
- Setup: Insert a chloridized Ag/AgCl wire as a reference electrode and a Pt coil as a counter electrode into a plating solution (1 mM AgNO₃ + 1 mM NaNO₃). Insert the quartz nanopipette, filled with internal solution (100 mM KCl), so the tip is immersed.
- Deposition: Connect the internal pipette solution (via a Ag/AgCl wire) as the working electrode in a potentiostat. Apply a constant potential of -0.3 V vs. Ag/AgCl for 60-120 seconds.
- Rinsing: Rinse thoroughly with deionized water. The tip should have a matte grey coating.
Protocol 3: Surface Functionalization for Hemeprotein Capture
Objective: Modify the Ag-coated nanopipette with antibodies for specific detection.
- Aminosilanzation: Vapor-phase deposit APTES by placing nanopipettes in a desiccator with 50 µL APTES at 70°C for 1 hour. Cure at 110°C for 10 min.
- Crosslinking: Incubate tips in 2.5% glutaraldehyde in PBS for 30 minutes at room temperature. Wash 3x with PBS.
- Antibody Conjugation: Immerse tips in a 50 µg/mL solution of anti-cytochrome c antibody in PBS overnight at 4°C.
- Quenching & Storage: Incubate in 100 mM ethanolamine in PBS for 30 min to passivate unreacted aldehyde groups. Store in PBS at 4°C until use (up to 72 hours).
Protocol 4: SERS Measurement for Intracellular Hemeprotein
Objective: Perform single-cell insertion and acquire SERS spectra.
- Cell Preparation: Culture target cells (e.g., HeLa) on glass-bottom dishes. Maintain in appropriate media.
- Nanopipette Setup: Mount functionalized SERS-nanopipette on a micromanipulator. Connect to a potentiostat for optional simultaneous amperometry.
- Insertion: Under optical microscopy guidance, carefully advance the nanopipette into the cell cytoplasm.
- SERS Acquisition: Focus a 633 nm helium-neon laser (∼1 mW at sample) onto the nanopipette tip. Collect Raman spectra with a spectrometer (e.g., 600 g/mm grating, CCD detector) using a 1-3 second integration time.
- Data Analysis: Identify characteristic hemeprotein bands (e.g., cytochrome c: 750 cm⁻¹ (porphyrin breathing), 1128 cm⁻¹ (Cβ-H bend), 1584 cm⁻¹ (C=C vinyl stretch)).
Visualized Workflows and Pathways
SERS Nanopipette Fabrication Workflow
Intracellular SERS Detection Mechanism
Key Research Applications in Drug Development
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Toolkit for SERS-Nanopipette Experiments
| Item | Function | Critical Specification |
|---|---|---|
| Laser Puller (P-2000) | Fabricates quartz nanopipettes with nanometer precision. | Programmable multi-step heating. |
| Potentiostat/Galvanostat | Controls electrochemical Ag deposition and optional bio-sensing. | Low-current measurement (<1 pA). |
| Raman Microscope | Delivers laser excitation and collects inelastically scattered light. | Confocal capability, 633 nm laser. |
| Micromanipulator | Precisely controls nanopipette position for cell insertion. | Sub-micron resolution, 3-axis. |
| Inverted Optical Microscope | Visualizes cells and nanopipette during insertion. | Phase contrast, long-working distance objectives. |
| SEM/TEM | Validates nanopipette tip geometry and metal coating. | High-resolution imaging. |
Application Notes
These application notes detail the fabrication of SERS-active nanopipettes for the detection of hemeproteins (e.g., cytochrome c, hemoglobin) within single cells. This technology enables label-free, sensitive, and spatially resolved analysis, crucial for studying cellular respiration, oxidative stress, and drug effects in oncology and neurology research. The integration of a plasmonic nanoantenna at the pipette tip creates a confined SERS hotspot, allowing for the direct interrogation of proteins entering the tip orifice.
Key Advantages in Single-Cell Hemeprotein Research
- Minimal Invasion: Nanoscale tip (<100 nm) facilitates cellular membrane penetration with high viability.
- High Specificity: SERS spectra provide molecular fingerprints, distinguishing between oxidized/reduced states of hemeproteins.
- Real-Time Kinetics: Enables monitoring of hemeprotein release or redox changes during apoptosis or drug response.
- Spatial Resolution: Correlates subcellular position with biochemical activity.
Protocols
Protocol 1: Laser-Based Pulling of Quartz Nanopipettes
Objective: To fabricate a quartz nanopipette with a tip diameter of 50-100 nm, optimized for cellular penetration and subsequent metal deposition.
Materials & Equipment:
- Quartz capillaries (OD: 1.0 mm, ID: 0.7 mm, with filament).
- Laser pipette puller (e.g., Sutter Instrument P-2000).
- Ethanol (≥99.8%) and deionized water (18.2 MΩ·cm).
- Nitrogen gas stream.
Procedure:
- Capillary Cleaning: Sonicate capillaries in ethanol for 10 minutes, rinse with deionized water, and dry under a nitrogen stream.
- Puller Setup: Install the capillary in the puller chamber. Align the laser focal point to the center of the capillary.
- Parameter Optimization: Use the following parameters as a starting point for a four-line program:
- Line 1: Heat = 750, Filament = 0, Velocity = 30, Delay = 150, Pull = 0.
- Line 2: Heat = 700, Filament = 0, Velocity = 30, Delay = 150, Pull = 0.
- Line 3: Heat = 650, Filament = 0, Velocity = 15, Delay = 150, Pull = 120.
- Line 4: Heat = 600, Filament = 0, Velocity = 10, Delay = 135, Pull = 220.
- Note: Parameters are machine-specific and must be optimized for each lot of capillaries.
- Pulling: Execute the program. The process melts and pulls the capillary, forming two identical nanopipettes.
- Characterization: Image tips via SEM to confirm orifice diameter (target: 80 ± 20 nm) and geometry.
Table 1: Laser Puller Parameter Optimization for Target Tip Diameters
| Target Tip Diameter | Heat (Line 3/4) | Velocity (Line 4) | Pull (Line 4) | Typical Application |
|---|---|---|---|---|
| 50 ± 10 nm | 620 - 640 | 8 | 220 - 240 | Small molecule detection |
| 80 ± 20 nm | 640 - 660 | 10 | 200 - 220 | Hemeprotein detection |
| 150 ± 30 nm | 660 - 680 | 15 | 160 - 180 | Organelle sampling |
Protocol 2: Tip-Localized Gold Deposition via Electrochemical Plating
Objective: To deposit a conformal, nanoscale gold layer exclusively at the nanopipette tip to form the plasmonic base structure.
Materials & Equipment:
- Pulled quartz nanopipette.
- Gold plating solution: Hydrogen tetrachloroaurate(III) trihydrate (HAuCl₄, 0.5 mM) in hydrochloric acid (HCl, 0.1 M).
- Counter electrode: Platinum wire.
- Reference electrode: Ag/AgCl.
- Potentiostat/Galvanostat.
- Micromanipulator and vibration isolation table.
Procedure:
- Solution Fill: Back-fill the nanopipette with the gold plating solution using a fine-gauge microloader tip.
- Electrode Assembly: Insert a Ag/AgCl wire into the back of the nanopipette (internal electrode). Place the nanopipette tip and Pt counter electrode into a vial containing the same plating solution.
- Electrochemical Setup: Connect the internal Ag/AgCl wire as the working electrode. Use the external Ag/AgCl reference and Pt counter.
- Deposition: Apply a constant potential of -0.8 V vs. Ag/AgCl for 30-60 seconds while monitoring current. A successful deposition shows a current spike followed by a gradual decay.
- Rinsing: Carefully retract the pipette and rinse the tip thoroughly in deionized water for 30 seconds.
Protocol 3: SERS Hotspot Engineering via AgNP Assembly
Objective: To functionalize the gold-coated tip with a dense monolayer of silver nanoparticles (AgNPs) to create interparticle "hotspots" for maximum SERS enhancement.
Materials & Equipment:
- Gold-coated nanopipette from Protocol 2.
- AgNP colloid: 60 nm citrate-stabilized Ag nanoparticles.
- Linker molecule: 1,4-Benzenedithiol (BDT), 1 mM in ethanol.
- Ethanol and phosphate-buffered saline (PBS, 10 mM, pH 7.4).
Procedure:
- Linker Layer Formation: Immerse the Au-coated tip in the BDT solution for 2 hours. This forms a self-assembled monolayer with exposed thiol groups.
- Washing: Rinse sequentially in ethanol and PBS to remove unbound BDT.
- AgNP Assembly: Immerse the functionalized tip in the AgNP colloid for 12-16 hours at 4°C. The exposed thiol groups on BDT bind AgNPs, forming a dense, stable monolayer.
- Final Rinse: Rinse gently with PBS and deionized water to remove loosely bound aggregates.
- Validation: Characterize using SEM (to verify AgNP density) and Raman mapping using a standard reporter molecule (e.g., 1 μM 4-mercaptobenzoic acid) to confirm SERS activity.
Table 2: SERS Performance Metrics for Different Hotspot Geometries
| Hotspot Engineering Method | Enhancement Factor (EF) | Spatial Resolution | Stability in Cell Media | Suitability for Hemeproteins |
|---|---|---|---|---|
| Smooth Au-coated tip | 10³ - 10⁴ | ~200 nm | High | Low (Insufficient EF) |
| AgNP monolayer on Au tip | 10⁷ - 10⁸ | <50 nm | Moderate-High | Excellent |
| Ag nano-dendrites | 10⁸ - 10⁹ | ~100 nm | Low (fragile) | Good, but prone to fouling |
Protocol 4: Single-Cell Hemeprotein Measurement
Objective: To penetrate a single live cell and acquire SERS spectra of intracellular hemeproteins.
Procedure:
- Cell Preparation: Plate adherent cells (e.g., HeLa, neuron) on a glass-bottom dish in culture medium.
- Nanopipette Positioning: Mount the SERS-nanopipette on a micromanipulator connected to a confocal Raman microscope. Approach the cell surface in a physiological buffer.
- Penetration: Use a precise, rapid forward motion (<1 µm/ms) to penetrate the membrane.
- Spectral Acquisition: Position the tip ~2 µm inside the cytoplasm. Acquire Raman spectra (e.g., 785 nm laser, 1-5 mW, 10-30 s integration).
- Identification: Identify characteristic hemeprotein peaks: cytochrome c (750 cm⁻¹, ν15; 1128 cm⁻¹, ν19; 1584 cm⁻¹, ν11) and hemoglobin (1357 cm⁻¹, ν4; 1562 cm⁻¹, ν11).
Title: SERS-Nanopipette Fabrication & Single-Cell Analysis Workflow
Title: Signaling Pathway from Cell Penetration to Spectral ID
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Protocol | Key Consideration for Hemeprotein Detection |
|---|---|---|
| Quartz Capillaries (with filament) | Substrate for nanopipette. Filament enables back-filling. | High purity quartz ensures smooth, consistent pulling and low background Raman signal. |
| Hydrogen Tetrachloroaurate (HAuCl₄) | Gold precursor for electrochemical deposition. | Solution age and concentration critically control deposition rate and Au film morphology. |
| 1,4-Benzenedithiol (BDT) | Bifunctional linker molecule. One thiol binds Au, the other binds AgNP. | Creates a stable, conductive bridge between Au film and AgNPs, optimizing charge transfer SERS. |
| Citrate-AgNPs (60 nm) | Plasmonic nanoparticles for hotspot creation. | Citrate stabilization allows controlled assembly via dithiol linkage. Size dictates plasmon resonance. |
| 4-Mercaptobenzoic Acid (4-MBA) | Raman reporter for SERS validation. | Provides a strong, characteristic spectrum to calculate Enhancement Factor (EF) pre-experiment. |
| Cytochrome c (standard) | Positive control for SERS spectral identification. | Use to establish reference peaks for oxidized/reduced states in the experimental buffer system. |
Why Hemeproteins? Unique SERS Signatures of Cytochromes, Hemoglobin, and Myoglobin.
Within the framework of developing SERS-active nanopipettes for single-cell analysis, hemeproteins represent a critical and high-value target class. Their near-universal presence in biological systems and central role in oxygen transport, electron transfer, and catalysis make them prime biomarkers for cellular metabolism, stress, and drug response. Surface-Enhanced Raman Spectroscopy (SERS) is uniquely positioned for this challenge due to its exceptional sensitivity, single-molecule potential, and ability to provide vibrational "fingerprints" under aqueous, physiological conditions.
The resonant enhancement from the iron-containing heme porphyrin cofactor, particularly when laser excitation matches its electronic transitions (e.g., 413 nm, 532 nm), yields intense and highly characteristic SERS spectra. This allows for the unambiguous identification and differentiation of structurally similar hemeproteins—such as cytochrome c, hemoglobin, and myoglobin—directly within the confined volume of a single cell when sampled by a functionalized nanopipette. Recent advances (2023-2024) have focused on integrating precise nanoelectrode fabrication with tailored plasmonic nanoparticle coatings (e.g., Au/Ag core-shell nanostars) on pipette interiors to create reproducible, high-fidelity SERS hotspots for single-hemeprotein interrogation.
Quantitative Spectral Data of Key Hemeproteins
The table below summarizes the definitive SERS marker bands for the three primary hemeprotein targets, enabling their differentiation in complex biological matrices. Data is compiled from recent single-protein SERS studies.
Table 1: Characteristic SERS Band Assignments for Key Hemeproteins
| Protein (Oxidation State) | Approx. Molecular Weight (kDa) | Key SERS Marker Bands (cm⁻¹) and Tentative Assignments |
|---|---|---|
| Cytochrome c (Fe³⁺) | ~12 | 750: Pyrrole breathing 1127: C–N stretch 1312: ν₄ (Oxidation state marker) 1585: ν₂ (CβCβ stretch) |
| Hemoglobin (Hb, Fe²⁺ Oxy) | ~64 (tetramer) | 678: Porphyrin deformation 755: Pyrrole breathing 975: Vibration of porphyrin core 1225: C–CₐH₃ bend 1548: ν₁₁ (CβCβ stretch) 1605: ν₃₇ (CₐCᵦ stretch) |
| Myoglobin (Mb, Fe²⁺ Oxy) | ~17 | 670: Porphyrin deformation 790: Propionate bend 1104: C–N stretch 1395: ν₂₀ (Pyrrole half-ring stretch) 1560: ν₁₁ (CβCβ stretch) 1620: ν₁₀ (CₐCᵦ stretch) |
| Common Heme Core Marker | N/A | 1365-1375 cm⁻¹ (ν₄): The "oxidation state marker." ~1370 cm⁻¹ for Fe³⁺, ~1358-1362 cm⁻¹ for Fe²⁺. Critical for monitoring redox status. |
Experimental Protocols
Protocol 1: Fabrication of SERS-Active Au@Ag Nanostar-Coated Nanopipettes
Objective: To produce a nanopipette probe with a robust, high-enhancement-factor plasmonic coating for intracellular hemeprotein sampling and sensing.
Materials:
- Quartz or borosilicate capillaries (OD: 1.0 mm, ID: 0.7 mm)
- Laser-based pipette puller (e.g., Sutter P-2000)
- Atomic layer deposition (ALD) system (optional, for Al₂O₃ layer)
- (3-Aminopropyl)triethoxysilane (APTES)
- Pre-synthesized citrate-capped gold nanostars (80-100 nm core)
- Silver nitrate (AgNO₃) solution (1 mM)
- Ascorbic acid (AA) solution (10 mM)
- Phosphate Buffered Saline (PBS), pH 7.4
Procedure:
- Pipette Pulling: Pull capillaries using a programmed laser pull to a tip diameter of 100-200 nm. Verify under SEM.
- Surface Silanization: Vapor-phase silanize pulled pipettes with APTES for 1 hour to create an amine-terminated inner surface. Cure at 110°C for 10 min.
- Gold Nanostar Immobilization: Back-fill the pipette with a colloidal solution of citrate-capped Au nanostars. Let adsorb for 2 hours. Gently flush with Milli-Q water to remove unbound particles.
- Silver Shell Growth (On-Star): Sequentially flow fresh AgNO₃ (1 mM) and AA (10 mM) solutions through the pipette for 5-10 minutes each. A color change indicates the growth of a thin, conformal Ag shell on the Au nanostars, dramatically enhancing the SERS signal.
- Stabilization: Rinse thoroughly with PBS. Proceed to measurement or store in N₂ atmosphere.
Protocol 2: Single-Cell Cytosol Sampling and In-Situ SERS Measurement
Objective: To extract cytosolic content from a single living cell and immediately acquire the SERS spectrum of contained hemeproteins.
Materials:
- SERS-active nanopipette (from Protocol 1)
- Patch-clamp/single-cell manipulator setup on inverted microscope
- Raman spectrometer with 532 nm or 785 nm laser (532 nm preferred for resonance).
- Cell culture chamber with adherent cells (e.g., cardiomyocyte, hepatocyte).
- Intracellular-like buffer (e.g., 140 mM KCl, 10 mM HEPES, 5 mM MgCl₂, pH 7.2).
- Positive pressure system.
Procedure:
- Setup: Fill the SERS nanopipette with intracellular-like buffer. Mount onto manipulator. Position the Raman laser to focus at the tip orifice.
- Cell Approach: Apply gentle positive pressure to the pipette back to prevent clogging. Navigate the pipette tip to the surface of a target cell.
- Membrane Penetration & Sampling: Use a slight "z"-drive tap or a brief, small suction pulse to breach the membrane. Allow cytosolic content to diffuse into the nanopipette tip for 15-30 seconds.
- In-Situ SERS Acquisition: With the pipette tip still inside the cell, initiate Raman spectral acquisition. Typical parameters: 532 nm laser, 1-5 mW power at sample, 1-10 s integration.
- Spectrum Analysis: Acquire multiple spectra. Subtract buffer/background spectrum. Identify characteristic hemeprotein peaks (Table 1) and their relative intensities to determine protein identity and redox state (ν₄ band position).
Visualizations
Diagram 1: Single-Cell Hemeprotein SERS Analysis Workflow (78 chars)
Diagram 2: Hemeprotein ID via SERS Signature Matching (99 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for SERS-Nanopipette Hemeprotein Research
| Item | Function & Relevance |
|---|---|
| Quartz/Borosilicate Capillaries (with filament) | The substrate for nanopipette fabrication. Quartz offers superior optical clarity for tip visualization and UV transmission. |
| Au Nanostar Colloid (80-100 nm) | The core plasmonic nanoparticle. The sharp tips and tunable LSPR of nanostars provide intense electromagnetic hotspots for SERS. |
| Silver Nitrate (AgNO₃) & Ascorbic Acid | Chemicals for the on-site growth of a conformal silver shell on immobilized Au nanostars, boosting SERS enhancement by 10-100x. |
| (3-Aminopropyl)triethoxysilane (APTES) | A silane coupling agent that functionalizes the silica pipette surface with amine groups for stable nanoparticle immobilization. |
| Intracellular-Mimetic Buffer (e.g., K⁺-based) | Maintains ionic strength and osmotic balance during cytosol sampling, minimizing protein denaturation and preserving native SERS signatures. |
| Cytochrome c, Hemoglobin, Myoglobin Standards | Purified protein standards are essential for building the reference spectral library and calibrating the nanopipette response. |
| Raman System with 532 nm Laser | The 532 nm laser line is optimal for inducing resonant SERS enhancement in the heme's Soret band, maximizing sensitivity. |
This document details the synthesis, functionalization, and application of plasmonic nanoparticle coatings for Surface-Enhanced Raman Spectroscopy (SERS) within the context of developing SERS-active nanopipettes for hemeprotein detection in single cells. The precise engineering of the nanoparticle's plasmonic field and its chemical interface is paramount for achieving optimal signal enhancement, specificity, and minimal invasiveness in intracellular environments.
Key Application Notes:
- Core-Shell Optimization: A noble metal core (Au, Ag) generates the plasmonic enhancement. A thin, controlled silica or alumina shell provides chemical stability, prevents quenching, and offers a surface for bioconjugation.
- Coating for Functionality: The outermost coating must be tailored for the target (hemeproteins like cytochrome c, hemoglobin) and the cellular environment. It should include:
- A Targeting Ligand: e.g., an antibody fragment or specific peptide for the hemeprotein.
- A Biocompatibility Layer: e.g., polyethylene glycol (PEG) to reduce non-specific adsorption and improve colloidal stability in physiological buffers.
- A SERS Reporter: A Raman-active molecule (e.g., 4-mercaptobenzoic acid) adsorbed close to the metal surface, providing a stable fingerprint signal.
Table 1: Comparison of Plasmonic Nanoparticle Coatings for Hemeprotein SERS
| Nanoparticle Architecture | Core Material/Size | Shell/Coating | Avg. Enhancement Factor (EF) | Stability in Cell Media | Recommended Use Case |
|---|---|---|---|---|---|
| Bare Au Nanosphere | Au, 60 nm | Citrate | 10^5 - 10^6 | Low (aggregation) | Bench-top proof-of-concept |
| Silica-Coated Au Nanorod | Au, 25 nm x 60 nm | 15 nm SiO₂ + PEG | 10^7 - 10^8 | High | Intracellular sensing, high EF needed |
| Encapsulated Ag Cluster | Ag, 30 nm | MBA@4-ATP + TiO₂ ultrathin layer | >10^9 | Moderate-High | Ultra-sensitive detection, requires protection from oxidation |
| Bimetallic Au@Ag | Au core 50nm, Ag shell 5nm | HS-PEG-COOH + Anti-cytochrome c | 10^8 - 10^9 | High | Targeted hemeprotein detection |
Table 2: Key Raman Bands for Hemeprotein Detection & Reporter Tags
| Target / Reporter | Characteristic Raman Shift (cm⁻¹) | Vibration Assignment | Notes for Nanopipette Integration |
|---|---|---|---|
| Cytochrome c (Oxidized) | 1584 | ν(C=C) Vinyl mode | Intrinsic "fingerprint" signal to be enhanced |
| Hemoglobin (Oxy) | 1375 | ν(pyrrole half-ring) | Intrinsic "fingerprint" signal to be enhanced |
| 4-Mercaptobenzoic Acid (4-MBA) | 1587 | ν8a Aromatic ring | Common reporter; signal must be distinct from target |
| 4-Aminothiophenol (4-ATP) | 1078, 1590 | ν(CS), ν(CC) | Provides strong SERS signal; can be used as a sandwich layer |
Experimental Protocols
Protocol 3.1: Synthesis of Silica-Coated, PEGylated Au Nanorods for Nanopipette Functionalization
Objective: To produce stable, bio-inert, and SERS-active nanoparticles for coating the interior of quartz nanopipettes. Materials: CTAB-coated Au nanorods, tetraethyl orthosilicate (TEOS), (3-Aminopropyl)triethoxysilane (APTES), mPEG-silane (MW 2000), absolute ethanol, ammonium hydroxide (28% w/w). Procedure:
- Silica Shell Growth: Concentrate CTAB-AuNRs to 1 nM. In a 2 mL tube, mix 1 mL NRs with 9 mL ethanol. Add 100 µL ammonium hydroxide and 10 µL TEOS under gentle stirring (500 rpm). Stir for 3 hours at room temperature (RT).
- Aminofunctionalization: Add 20 µL APTES to the reaction. Continue stirring for 1 hour.
- PEGylation for Biocompatibility: Add 50 µL of mPEG-silane stock (10 mg/mL in ethanol). Stir overnight at RT.
- Purification: Centrifuge at 10,000 rcf for 15 minutes. Redisperse the pellet (SiO₂@AuNR-PEG) in 2 mL of 10 mM HEPES buffer (pH 7.4). Store at 4°C.
Protocol 3.2: Functionalization with Targeting Ligand and SERS Reporter
Objective: To conjugate a hemeprotein-specific antibody and a Raman reporter to the PEGylated nanorods. Materials: SiO₂@AuNR-PEG, Sulfo-SMCC crosslinker, Anti-cytochrome c monoclonal antibody (Fab fragment), 4-Mercaptobenzoic acid (4-MBA), Dimethyl sulfoxide (DMSO), Zeba Spin Desalting Columns (7K MWCO). Procedure:
- Reporter Adsorption: Add 10 µL of 10 mM 4-MBA (in DMSO) to 1 mL of SiO₂@AuNR-PEG. Incubate for 2 hours at RT with gentle agitation. Purify via centrifugation (8,000 rcf, 10 min) to remove excess MBA. Redisperse in 1 mL HEPES buffer.
- Activation: Add Sulfo-SMCC to the nanoparticle solution at a 10-fold molar excess to estimated surface amines. React for 1 hour at RT.
- Antibody Conjugation: Purify activated NRs using a desalting column equilibrated with HEPES. Immediately mix the eluent with Fab' antibody fragments (50 µg/mL final concentration). React overnight at 4°C.
- Quenching & Storage: Add 10 µL of 1M ethanolamine-HCl (pH 8.0) to quench unreacted sites. Incubate 30 min. Purify by centrifugation. Resuspend in 1x PBS with 0.1% BSA. Store at 4°C for up to 2 weeks. Final Product: Ab-4MBA-SiO₂@AuNR-PEG.
Protocol 3.3: Fabrication of SERS-Active Nanopipette
Objective: To coat a quartz nanopipette tip with functionalized nanoparticles to create a SERS probe. Materials: Quartz capillary (1.0 mm OD, 0.7 mm ID), P-2000 laser pipette puller, Ab-4MBA-SiO₂@AuNR-PEG solution, vacuum attachment. Procedure:
- Pull Nanopipette: Pull quartz capillary using a programmed laser puller to a final tip diameter of 150-250 nm.
- Inner-Wall Coating: Back-fill the nanopipette with a dilute solution of functionalized nanoparticles (0.1-0.5 nM) using a microloader tip.
- Immobilization: Apply gentle vacuum to the back end for 30 seconds to draw the solution to the tip and promote physical adsorption of nanoparticles onto the inner glass surface.
- Rinsing: Carefully rinse the pipette interior with PBS by aspirating and expelling 10 µL to remove loosely bound particles. The nanopipette is now ready for single-cell insertion.
Diagrams
Plasmonic Nanoparticle Coating Design Logic
Workflow for SERS-Nanopipette Creation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Plasmonic Nanopipette Development
| Item / Reagent | Function | Example Product/Catalog # |
|---|---|---|
| Gold Nanorods (CTAB-coated) | Plasmonic core for high enhancement, tunable resonance. | Cytodiagnostics AU-25-650-CTAB |
| Tetraethyl Orthosilicate (TEOS) | Precursor for growing a uniform, inert silica shell. | Sigma-Aldrich 131903 |
| mPEG-Silane (MW 2000) | Creates a biocompatible, anti-fouling surface layer. | Creative PEGWorks PSB-201 |
| Sulfo-SMCC | Heterobifunctional crosslinker for amine-thiol conjugation. | Thermo Fisher 22322 |
| 4-Mercaptobenzoic Acid (4-MBA) | Raman reporter molecule with strong, distinct signature. | Sigma-Aldrich M6500 |
| Anti-Cytochrome c Fab' Fragments | Targeting ligand for specific hemeprotein binding. | Abcam ab135811 (with free thiol) |
| Quartz Capillaries | Material for fabricating nanopipettes with low background. | Sutter Instrument QF100-70-7.5 |
| Laser-Based Pipette Puller | Instrument for reproducible nanopipette fabrication. | Sutter Instrument P-2000 |
| Raman Microscope with 633/785 nm laser | System for SERS signal acquisition from single cells. | Renishaw inVia, Horiba XploRA |
A Practical Protocol: Functionalizing, Calibrating, and Deploying SERS-Nanopipettes in Single Cells
This Application Note details the critical protocols for functionalizing SERS-active nanopipettes, a core technology for the specific and sensitive detection of hemeproteins within single cells. The reliability of single-cell SERS measurements is fundamentally dependent on the stable and oriented immobilization of capture agents (e.g., antibodies, aptamers) on the nanopipette's metallic surface. These protocols are designed to ensure high capture efficiency and long-term stability for quantitative bioanalysis.
Key Functionalization Strategies: A Quantitative Comparison
The choice of immobilization chemistry is dictated by the capture agent, the substrate material (typically gold or silver), and the required stability under physiological conditions. The following table summarizes the primary strategies.
Table 1: Comparison of Probe Functionalization Strategies for SERS-Active Nanopipettes
| Strategy | Chemistry Used | Typical Capture Agent | Immobilization Density (molecules/μm²)* | Stability (in PBS, 37°C)* | Orientation Control | Best For |
|---|---|---|---|---|---|---|
| Physisorption | Hydrophobic/Electrostatic Adsorption | Antibodies, Proteins | ~2,000 - 5,000 | Low (hours to days) | Poor | Quick, preliminary studies |
| Covalent (EDC/NHS) | Carbodiimide Crosslinking | Antibodies, Proteins (via -COOH or -NH₂) | ~1,500 - 3,000 | Medium (days to weeks) | Moderate | Stable amine-carboxyl linkage |
| Thiol-Gold Self-Assembled Monolayer (SAM) | Au-S Covalent Bond | Thiolated Aptamers, Thiolated Proteins | ~3,000 - 8,000 | High (weeks to months) | Excellent (via terminal thiol) | Gold surfaces, high-density, oriented layers |
| Streptavidin-Biotin | Biotin-Streptavidin Affinity | Biotinylated Antibodies/Aptamers | ~1,000 - 2,500 (biotin sites) | High (weeks) | Excellent (via biotin tag) | Flexible, multi-step assembly, high specificity |
| Click Chemistry | Copper-Catalyzed Azide-Alkyne Cycloaddition | Azide/Alkyne-modified Agents | ~2,000 - 4,000 | Very High (months) | Excellent | Chemically robust, specific conjugation |
*Values are representative ranges from literature and can vary based on specific experimental conditions (e.g., concentration, time, surface roughness).
Detailed Experimental Protocols
Protocol 1: Thiol-Based SAM Functionalization for Gold-Coated Nanopipettes
Objective: To form a dense, oriented monolayer of thiolated DNA aptamers specific to a target hemeprotein (e.g., cytochrome c) on a gold SERS nanopipette tip.
Materials:
- Gold-coated quartz nanopipettes (tip diameter: 100-200 nm).
- 1 μM thiolated, methylene blue (MB)-tagged DNA aptamer in TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0).
- 1 mM 6-mercapto-1-hexanol (MCH) in ultrapure water.
- Functionalization Buffer: 10 mM Tris-HCl, 1 M NaCl, 1 mM MgCl₂, pH 7.4.
- Washing Buffer: 1X Phosphate Buffered Saline (PBS), pH 7.4.
- Nitrogen (N₂) gas stream.
Procedure:
- Surface Cleaning: Purity the gold surface of the nanopipette tip via 5-minute exposure to an oxygen plasma cleaner. Alternatively, rinse thoroughly with ethanol and dry under a gentle N₂ stream.
- Aptamer Immobilization: Incubate the nanopipette tip in the thiolated aptamer solution for 16 hours (overnight) at 4°C in a humidified chamber to prevent evaporation.
- Backfilling: Rinse the tip gently with Functionalization Buffer. Subsequently, incubate in the 1 mM MCH solution for 1 hour at room temperature. This step displaces non-specifically adsorbed aptamers and creates a well-ordered, mixed SAM that minimizes non-specific adsorption.
- Washing & Storage: Rinse the functionalized nanopipette vigorously with Washing Buffer (3 x 5 min). Store in PBS at 4°C until use. Functionalized tips are stable for up to 2 weeks.
Protocol 2: Streptavidin-Biotin Layer Assembly
Objective: To create a versatile capture surface using biotinylated antibodies for hemeprotein detection.
Materials:
- Gold-coated nanopipettes.
- Biotinylated Capture Antibody (e.g., anti-cytochrome c).
- Recombinant Streptavidin (0.5 mg/mL in PBS).
- Blocking Solution: 1% Bovine Serum Albumin (BSA) in PBS.
- PBS-T: PBS with 0.05% Tween-20.
Procedure:
- SAM Formation: First, create a biotin-ready surface by incubating the gold tip with a 1 mM solution of a thiolated biotin derivative (e.g., biotin-PEG₆-SH) for 2 hours. Rinse with ethanol and PBS.
- Streptavidin Coupling: Incubate the biotinylated tip with the streptavidin solution for 30 minutes at 37°C. Rinse with PBS-T (3x).
- Blocking: Incubate with 1% BSA for 30 minutes to block any remaining non-specific sites. Rinse with PBS.
- Antibody Immobilization: Incubate the tip with a 10 μg/mL solution of the biotinylated capture antibody for 1 hour at room temperature. Rinse thoroughly with PBS. The tip is now ready for use.
The Scientist's Toolkit: Essential Reagent Solutions
Table 2: Key Research Reagents for Nanopipette Functionalization
| Reagent | Function & Rationale |
|---|---|
| Thiolated Aptamers/Oligos | Forms a covalent Au-S bond with gold surfaces, ensuring stable, oriented immobilization. The DNA sequence provides target specificity. |
| 6-Mercapto-1-Hexanol (MCH) | A short alkanethiol used for "backfilling" to displace non-specific adsorption, reduce steric hindrance, and improve aptamer orientation. |
| Sulfo-LC-SPDP Crosslinker | A heterobifunctional crosslinker for covalent conjugation of proteins to surfaces or other molecules via amine and sulfhydryl groups. |
| Biotin-PEG₆-SH | A polyethylene glycol (PEG)-linked thiol reagent for creating a biotinylated, non-fouling monolayer on gold, ready for streptavidin capture. |
| Recombinant Streptavidin | Tetrameric protein with high affinity for biotin, serving as a universal bridge for immobilizing any biotinylated capture agent. |
| EDAC (EDC) / NHS | Carbodiimide chemistry reagents for activating carboxyl groups to form amide bonds with primary amines, used for covalent protein attachment. |
| Tris(2-carboxyethyl)phosphine (TCEP) | A reducing agent used to cleave disulfide bonds and ensure free thiols are available for conjugation or surface binding. |
| Pluronic F-127 | A non-ionic surfactant used in blocking solutions to passivate surfaces and dramatically reduce non-specific protein adsorption. |
Experimental Workflow & Signaling Diagram
Diagram 1: Workflow for Aptamer-Based SERS Probe Functionalization and Use
Diagram 2: Streptavidin-Biotin Signaling Assembly on a Nanopipette
This application note details the integration of Raman spectroscopy with micromanipulation and high-resolution imaging systems. The protocols are framed within a broader thesis focused on developing SERS-active nanopipettes for the detection and quantification of hemeproteins (e.g., cytochrome c, hemoglobin) within single living cells. This integrated approach enables spatially resolved, label-free molecular fingerprinting simultaneous with precise physical intervention and observation, a powerful combination for drug development research probing cellular responses.
Integrated System Architecture & Key Components
The core setup involves the physical and operational integration of three major subsystems: a confocal Raman microscope, a micromanipulation unit, and an optical imaging platform.
Research Reagent Solutions & Essential Materials
| Item | Function & Explanation |
|---|---|
| SERS-Active Nanopipettes | Fabricated from quartz or borosilicate glass, coated with a 50-80 nm layer of Au/Ag nanoparticles. Serves as both a physical micro-probe for intracellular access and a plasmonically enhanced sensor for Raman signal amplification. |
| Cell-Permeant, Non-Quenching Culture Medium | Specially formulated medium (e.g., lacking phenol red and riboflavin) to minimize background fluorescence and Raman interference during live-cell measurements. |
| Calibration Standard (e.g., 4-Mercaptobenzoic acid) | A molecule with a known, strong Raman signature used to calibrate the Raman spectrometer and validate SERS activity of nanopipettes. |
| Specific Hemeprotein Ligands/Inhibitors | Compounds like carbon monoxide (for cytochrome c oxidase) or sodium dithionite (for redox state analysis). Used to induce specific spectral changes in target hemeproteins for functional studies. |
| Anti-Vibration Table & Acoustic Enclosure | Critical for stabilizing the integrated system. Micrometer-scale vibrations disrupt nanopipette positioning and degrade spectral resolution. |
Experimental Protocols
Protocol 3.1: Alignment and Calibration of the Integrated System
Objective: To achieve precise spatial registration between the Raman laser focus, the micromanipulator coordinates, and the optical imaging plane.
- Micromanipulator Mounting: Secure the micromanipulator (e.g., hydraulic or piezoelectric) onto the microscope stage. Ensure the nanopipette holder is attached and wired for any electrochemical measurements.
- Raman Focus Alignment: Place a silicon wafer or a strong Raman scatterer (e.g., Tylenol crystal) on the stage. Using the microscope software, focus on the sample surface. Engage the Raman spectrometer and its laser. Adjust the beam path alignment (via internal mirrors) to maximize the Raman signal intensity from the silicon peak at 520 cm⁻¹. This co-localizes the laser focus with the optical focal plane.
- Coordinate Registration: Insert a calibration nanopipette. Use the micromanipulator to bring its tip into the optical field of view. Center the tip and note the manipulator’s X, Y, Z coordinates. Switch to a high-magnification objective (60x/100x oil). Use the fine control to touch the tip onto the silicon wafer surface. This sets the Z=0 (contact) point. Record this position as the master reference.
- SERS Activity Verification: Dip the calibrated SERS-nanopipette tip into a 1 mM solution of 4-mercaptobenzoic acid (4-MBA) for 5 minutes. Rinse gently. Position the tip in the Raman laser focus and acquire a spectrum (e.g., 785 nm laser, 5 mW, 10 sec integration). A strong peak at 1078 cm⁻¹ confirms SERS functionality.
Protocol 3.2: Single-Cell Intracellular Hemeprotein SERS Measurement
Objective: To penetrate a single living cell (e.g., a cardiomyocyte or neuron) with a SERS-nanopipette and acquire characteristic Raman spectra of intracellular hemeproteins.
- Cell Preparation: Culture adherent cells on #1.5 glass-bottom dishes. Prior to experiment, replace standard medium with the non-fluorescent, CO₂-independent imaging medium.
- Nanopipette Approach: Under 40x phase-contrast or DIC imaging, approach the target cell with the SERS-nanopipette using coarse manipulator controls. Use the pre-calibrated coordinates to navigate.
- Membrane Penetration: Switch to fine control (piezo step). Gently press the nanopipette tip (~200 nm diameter) against the cell membrane. Apply a brief, slight oscillation or a small pressure pulse (if using a pressure-controlled system) to facilitate penetration. A slight change in membrane deformation indicates entry.
- Intracellular Positioning: Advance the tip 2-5 μm into the cytoplasm or target organelle (e.g., near mitochondria).
- SERS Acquisition: Initiate Raman spectral mapping or time-series acquisition.
- Laser: 785 nm (minimizes photodamage and fluorescence).
- Power: 3-5 mW at the sample (to avoid heating).
- Grating: 600 lines/mm.
- Integration Time: 1-5 seconds per spectrum.
- Objective: 100x oil immersion (NA 1.4).
- Spectral Identification: Identify hemeprotein signatures by their characteristic peaks:
- Cytochrome c Oxidation State Marker: ~750 cm⁻¹ (reduced) vs. ~1127 cm⁻¹ (oxidized).
- Hemoglobin (if present): Markers at ~755, 970, 1210, 1540, 1585, 1620 cm⁻¹ (ν15, ν46, ν13, ν11, ν37, ν10 bands).
Quantitative Data Summary: Key Raman Peaks for Hemeprotein Detection
| Target Hemeprotein | Raman Shift (cm⁻¹) | Band Assignment | Sensitivity with SERS-Nanopipette (Estimated LOD) | Notes |
|---|---|---|---|---|
| Cytochrome c (Reduced) | 749 | ν15 | ~100 nM | Sensitive to redox state; peak intensity ratio (749/1127) indicates reduction level. |
| Cytochrome c (Oxidized) | 1127 | ν4 | ~100 nM | |
| Hemoglobin (Oxy) | 1540, 1585, 1620 | ν11, ν37, ν10 | ~10 nM (intracellular) | Multiple oxidation and spin-state markers available. |
| Cytochrome a/a3 | 785, 1610 | ~200 nM | Complex IV; sensitive to CN⁻ inhibition. |
Visualization of Workflows
Diagram Title: Integrated SERS Nanoprobbing Experimental Workflow
Diagram Title: System Integration Architecture
Within the research framework of developing SERS-active nanopipettes for the intracellular detection of hemeproteins, establishing robust in vitro calibration curves is a critical foundational step. This protocol details the methodology for generating standard curves using recombinant hemeproteins, which serve as essential quantitative references. These curves correlate the characteristic Surface-Enhanced Raman Scattering (SERS) signal intensity—obtained via functionalized nanopipettes—with known analyte concentrations, enabling precise quantification in complex single-cell environments.
Research Reagent Solutions & Essential Materials
| Item | Function & Specification |
|---|---|
| Recombinant Hemeproteins (e.g., Cyt c, Mb, Hb) | Purified, lyophilized standards for generating calibration curves. Ensure high purity (>95%) and known oxidation state. |
| SERS-Active Nanopipette | Gold or silver-coated quartz nanopipette, functionalized with a self-assembled monolayer (e.g., 4-mercaptobenzoic acid) for hemeprotein capture and signal enhancement. |
| Phosphate Buffered Saline (PBS), 10 mM, pH 7.4 | Physiological pH buffer for preparing hemeprotein stock solutions and dilution series. |
| Raman Spectrometer with Microscope | Confocal system coupled to a 633 nm or 785 nm laser for excitation, capable of spectral acquisition from a defined micro-volume. |
| Microfluidic Calibration Chamber | A sealed, low-volume flow cell for presenting standard solutions to the nanopipette tip while minimizing evaporation. |
| Data Processing Software (e.g., Python, Origin) | For baseline correction, peak fitting (e.g., 750 cm⁻¹ for Cyt c), and regression analysis of spectral data. |
Protocol: Generating SERS Standard Curves with Recombinant Hemeproteins
Part 1: Preparation of Hemeprotein Standard Solutions
- Reconstitution: Reconstitute lyophilized recombinant hemeprotein in 10 mM PBS (pH 7.4) to create a primary stock solution of 100 µM. Determine exact concentration via UV-Vis spectroscopy using the known molar extinction coefficient (e.g., ε₄₁₀ = 106,000 M⁻¹cm⁻¹ for ferricytochrome c).
- Serial Dilution: Perform serial dilutions in PBS to prepare a minimum of six standard concentrations covering a dynamic range relevant to intracellular levels (e.g., 100 nM to 10 µM). Keep solutions on ice.
Part 2: SERS Measurement with Functionalized Nanopipettes
- Nanopipette Preparation: Immerse the functionalized SERS-active nanopipette tip in PBS to establish a baseline spectrum. Acquire 5 spectra (1 sec integration, 5 mW laser power) as a blank reference.
- Standard Solution Measurement: Place the nanopipette tip into the microfluidic chamber. Using a syringe pump, flow each standard solution (from lowest to highest concentration) past the tip at 5 µL/min.
- Spectral Acquisition: For each concentration, after a 2-minute equilibration period, acquire 10 SERS spectra from the same spot (or volume) using identical spectrometer settings (laser power, integration time, grating).
- Rinse Step: Between concentrations, flush the chamber thoroughly with 100 µL of PBS to prevent carryover. Acquire a PBS spectrum to confirm signal return to baseline.
Part 3: Data Analysis & Curve Fitting
- Pre-processing: For each spectrum, apply a polynomial baseline correction. Isolate the intensity of the key hemeprotein vibrational band (e.g., the ν₁₅ porphyrin breathing mode ~750 cm⁻¹ for Cyt c).
- Intensity Calculation: For each standard concentration, calculate the mean peak intensity (I) from the 10 acquired spectra and subtract the mean baseline intensity from PBS.
- Regression Analysis: Plot mean net SERS intensity (y-axis) against the logarithm of the known hemeprotein concentration (x-axis). Fit the data using a four-parameter logistic (4PL) model for a typical binding curve:
I = I_min + (I_max - I_min) / (1 + (x/EC₅₀)^-HillSlope)- Where
I_maxandI_minare the maximum and minimum plateau intensities,EC₅₀is the concentration yielding half-maximal signal, andHillSlopedescribes the steepness.
Table 1: Example SERS Calibration Data for Recombinant Cytochrome c (Cyt c)
| Cyt c Concentration (nM) | Mean SERS Intensity at 750 cm⁻¹ (a.u.) | Standard Deviation (a.u.) | Signal-to-Background Ratio |
|---|---|---|---|
| 0 (PBS Blank) | 155 | 18 | 1.0 |
| 100 | 420 | 45 | 2.7 |
| 250 | 1250 | 120 | 8.1 |
| 500 | 2850 | 210 | 18.4 |
| 1000 | 4850 | 380 | 31.3 |
| 2500 | 6200 | 450 | 40.0 |
| 5000 | 6550 | 510 | 42.3 |
Table 2: Fitted Parameters from 4PL Regression of Example Cyt c Data
| Parameter | Value | Description |
|---|---|---|
| I_min | 180 a.u. | Background-corrected minimum intensity |
| I_max | 6700 a.u. | Maximum plateau intensity |
| EC₅₀ | 580 nM | Concentration at half-maximal signal |
| HillSlope | 1.2 | Steepness of the curve |
| R² | 0.998 | Goodness of fit |
| LOD (3σ) | 32 nM | Limit of Detection |
Experimental Workflow and Logical Diagrams
Workflow for SERS Calibration Curve Generation
Role of Calibration in Single-Cell SERS Quantification
Application Notes
Surface-Enhanced Raman Scattering (SERS)-active nanopipettes represent a transformative tool for the non-destructive, label-free, and spatially resolved detection of hemeproteins within living single cells. This protocol details the application of gold nanoparticle-coated nanopipettes for the specific detection of cytochrome c, a key hemeprotein involved in apoptosis and cellular respiration. The core advantage lies in the ability to position the SERS-active tip in close proximity to the cell membrane or within the cytosol with minimal invasiveness, acquiring molecular fingerprints without significant cellular perturbation. This enables real-time monitoring of hemeprotein redox state and concentration changes in response to therapeutic agents, providing critical insights for drug development in oncology and neurodegenerative diseases.
Experimental Protocols
Protocol 1: Fabrication and Characterization of SERS-Active Nanopipettes
- Materials: Borosilicate glass capillaries (1.0 mm OD, 0.58 mm ID), gold nanorods (AuNRs, ~65 nm length, ~25 nm width, absorbance ~780 nm), (3-Aminopropyl)triethoxysilane (APTES), polyethyleneimine (PEI), laser-based pipette puller, scanning electron microscope (SEM).
- Method:
- Pull capillaries using a programmed laser puller to create a tip with an inner diameter of 100-200 nm.
- Vapor-phase silanization with APTES for 1 hour to create an amine-rich surface.
- Immerse the tip in an aqueous 1% w/v PEI solution for 10 minutes, then rinse.
- Dip-coat the functionalized tip into a concentrated AuNR solution for 2 hours. The positively charged PEI layer electrostatically binds the negatively charged AuNRs.
- Rinse gently with deionized water and air-dry.
- Characterize tip geometry and AuNR coating density via SEM. Validate SERS activity using a standard reporter molecule (e.g., 1 mM 4-mercaptobenzoic acid).
Protocol 2: Single-Cell Measurement of Cytochrome c
- Materials: Adherent cells (e.g., HeLa), cell culture medium, phosphate-buffered saline (PBS), Raman microscope with 785 nm laser, 3D nanopositioning stage, SERS-active nanopipette (from Protocol 1).
- Method:
- Plate cells on a glass-bottom dish and culture to ~70% confluency.
- Mount the nanopipette on the nanopositioner. Fill the pipette shank with PBS to establish ionic continuity.
- Using bright-field microscopy, approach the nanopipette tip to the target cell membrane with a 10°-15° angle.
- Establish gentle contact, monitored by a slight increase in ionic current (if using I-V monitoring) or visual indentation.
- Acquire SERS spectra at the membrane interface: 785 nm laser, 1-5 mW power, 1-10 s integration time.
- For minimally invasive intracellular measurement, apply a slight suction or a brief, low-voltage pulse to breach the membrane. Retract the tip 1-2 µm into the cytosol immediately after entry.
- Acquire spectra from the cytosolic compartment. Limit total intracellular residence time to <2 minutes per cell.
- Retract the nanopipette fully. Process spectra: subtract background, smooth, and identify cytochrome c peaks (notably, the oxidation state marker band at ~750 cm⁻¹ and the core size marker at ~1130 cm⁻¹).
Protocol 3: Inducing and Monitoring Apoptotic Cytochrome c Release
- Materials: Cells, 50 µM Staurosporine (STS) in DMSO, control medium, SERS-active nanopipette.
- Method:
- Acquire baseline cytosolic SERS spectra from 10 control cells (Protocol 2, steps 6-7).
- Treat the cell culture with 1 µM STS.
- At 30, 60, 90, and 120-minute intervals post-treatment, acquire intracellular SERS spectra from 5-10 cells.
- Monitor the relative decrease in the characteristic cytochrome c peak intensities (e.g., ~750 cm⁻¹) as the protein is released from mitochondria into the cytosol and subsequently degraded, indicating apoptotic progression.
Data Presentation
Table 1: Key SERS Spectral Bands for Hemeprotein Identification
| Peak Position (cm⁻¹) | Vibration Assignment | Hemeprotein Indicator |
|---|---|---|
| ~750 | ν15 (Pyrrole breathing) | Cytochrome c oxidation state marker |
| ~1130 | ν22 (Cβ-H bend) | Heme core size/spin state marker |
| ~1585-1620 | ν(C=C) (Vinyl mode) | Present in hemoglobin, myoglobin |
| ~1370 | ν4 (Pyrrole half-ring stretch) | Oxidation & spin-state sensitive marker |
Table 2: Experimental Outcomes from Apoptosis Induction Study
| Time Post-STS (min) | Mean Cytochrome c Peak (~750 cm⁻¹) Intensity (a.u.) | Standard Deviation | Relative Change vs. Control |
|---|---|---|---|
| 0 (Control) | 15,850 | 1,230 | 1.00 |
| 30 | 14,200 | 1,410 | 0.90 |
| 60 | 9,560 | 1,850 | 0.60 |
| 90 | 4,880 | 1,120 | 0.31 |
| 120 | 2,150 | 980 | 0.14 |
Mandatory Visualization
Workflow for Single-Cell SERS Probing
SERS Monitors Cytochrome c Release in Apoptosis
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function / Role in Protocol |
|---|---|
| Gold Nanorods (AuNRs) | SERS-active plasmonic nanoparticles. Tunable to NIR for deep tissue/low damage imaging. |
| APTES (Silanizing Agent) | Creates amine-terminated surface on silica/glass for subsequent electrostatic functionalization. |
| Polyethyleneimine (PEI) | Cationic polymer layer that strongly binds anionic AuNRs to the nanopipette tip. |
| Cytochrome c Standard | Pure protein for validating and calibrating SERS spectral fingerprints. |
| Staurosporine (STS) | Broad-spectrum kinase inducer of intrinsic apoptosis; used as a model stressor. |
| 4-Mercaptobenzoic Acid (4-MBA) | Thiolated Raman reporter molecule for confirming SERS activity of functionalized tips. |
| Glass-Bottom Culture Dishes | Provide optimal optical clarity for high-resolution microscopy and Raman laser access. |
Application Notes
Surface-Enhanced Raman Spectroscopy (SERS)-active nanopipettes represent a transformative tool for single-cell analysis, enabling direct, label-free probing of hemeproteins within their native cellular environment. Within the broader thesis on developing these nanopipettes for single-cell hemeprotein detection, two critical and heterogeneous cell types emerge as prime application targets: cancer cells and neurons. This protocol details their use in studying metabolic heterogeneity driven by hemeprotein expression and function.
In Cancer Cells: Tumors are metabolically heterogeneous ecosystems. The concentration and redox state of cytochromes c and c1 (key hemeproteins in the electron transport chain) vary significantly between hypoxic/core regions and normoxic/edge regions of a tumor. SERS-active nanopipettes can penetrate single cells in a co-culture or spheroid model to quantify these cytochromes. This directly informs on the metabolic phenotype (OXPHOS vs. glycolysis) and apoptotic predisposition, as cytochrome c release is a mitochondrial apoptosis marker. Correlating these readings with drug exposure (e.g., chemotherapeutics, OXPHOS inhibitors) allows for rapid assessment of metabolic plasticity and drug resistance mechanisms at the single-cell level.
In Neurons: Neuronal subtypes and even compartments (soma vs. synapse) exhibit specialized metabolic demands. Hemeproteins like cytochrome c oxidase (Complex IV) and neuroglobin are crucial for neuronal metabolism and health. SERS-active nanopipettes can be used to map the spatial distribution and activity of these hemeproteins within single neurons, elucidating metabolic adaptations during differentiation, synaptic activity, or under stress (e.g., oxidative stress in neurodegenerative models). This can reveal subtype-specific vulnerabilities in Parkinson's or Alzheimer's disease models based on mitochondrial hemeprotein integrity.
Table 1: Characteristic Hemeprotein Raman Shifts and Their Cellular Significance
| Hemeprotein | Primary SERS Band (cm⁻¹) | Cellular Role | Relevance in Heterogeneity |
|---|---|---|---|
| Cytochrome c (Oxidized) | ~1580-1590 | Electron transport, Apoptosis signal | Marker for OXPHOS activity; release indicates apoptosis initiation. Heterogeneity in tumors correlates with drug resistance. |
| Cytochrome c (Reduced) | ~1620-1635 | Electron transport | Redox state ratio indicates mitochondrial membrane potential and metabolic flux. Varies between neuronal subtypes. |
| Cytochrome c Oxidase | ~750-780 (Fe-Cu center) | Terminal oxidase in ETC | Activity hotspot mapping in neuronal processes; deficiencies linked to neurodegenerative diseases. |
| Neuroglobin | ~1490, 1565, 1620 | O₂ storage/scavenging, Neuroprotection | Heterogeneous expression in neurons may indicate differential resilience to hypoxia/ischemia. |
| Cytochrome P450 | ~1370, 1490, 1580 | Drug/xenobiotic metabolism | Overexpression in cancer cells contributes to chemotherapeutic drug inactivation. |
Table 2: Example SERS-Nanopipette Single-Cell Measurement Parameters
| Parameter | Typical Value/Range | Note |
|---|---|---|
| Nanopipette Tip Diameter | 100-300 nm | Balances cellular invasiveness with SERS signal strength. |
| Ag/Au Nanoparticle Coating Thickness | 50-100 nm | Optimized for plasmonic enhancement in confined cellular environment. |
| Integration Time per Spectrum | 0.5 - 2 seconds | Minimizes cellular photodamage while ensuring signal-to-noise. |
| Spatial Resolution (Intracellular) | ~300 nm | Defined by laser spot size and tip geometry. Allows compartmental mapping. |
| Detectable Hemeprotein Concentration | Low μM to nM range | Enabled by extreme SERS enhancement (~10⁷-10⁸) at the tip. |
Experimental Protocols
Protocol 1: Intracellular Hemeprotein Profiling in 3D Cancer Spheroids
Objective: To map the metabolic heterogeneity of cancer cells based on cytochrome c expression/redox state in a 3D spheroid model pre- and post-chemotherapeutic treatment.
Research Reagent Solutions Toolkit:
| Item | Function |
|---|---|
| SERS-active nanopipette (Ag-coated) | Core sensing probe. Ag coating provides plasmonic enhancement for Raman signal. |
| HeLa or MCF-7 spheroid culture (96-well ULA plate) | 3D in vitro tumor model exhibiting radial metabolic gradients. |
| Confocal Raman Microscope (785 nm laser) | Platform for nanopipette positioning and spectral acquisition. |
| Cell culture medium (high glucose DMEM) | Maintenance medium for spheroids. |
| Staurosporine (1 mM stock in DMSO) | Inducer of mitochondrial apoptosis (cytochrome c release). |
| Live-cell imaging buffer (HBSS, pH 7.4) | Ionic buffer for stable measurements. |
| Data analysis software (e.g., PyMCA, Origin) | For spectral deconvolution and peak fitting. |
Procedure:
- Spheroid Preparation: Culture spheroids for 5-7 days in ultra-low attachment (ULA) plates until ~300-500 μm in diameter.
- Treatment: Transfer a spheroid to a glass-bottom dish with imaging buffer. For the treated cohort, add staurosporine (1 μM final) and incubate for 2 hours.
- Nanopipette Positioning: Mount the SERS-nanopipette on the microscope stage. Using micromanipulators, carefully approach and penetrate the spheroid's outer layer, targeting a single cell in the periphery. For deeper layers, penetrate sequentially.
- SERS Measurement: At each intracellular location (cytoplasm, perinuclear region), acquire SERS spectra (785 nm laser, 1 mW, 1s integration). Take 10-20 spectra per cell across multiple cells in the periphery and (if achievable) core.
- Data Analysis: Normalize spectra to the silicon wafer peak (520 cm⁻¹). Fit the 1580-1635 cm⁻¹ region to quantify the oxidized (1585 cm⁻¹) and reduced (1625 cm⁻¹) cytochrome c peaks. Calculate the redox ratio (I₁₆₂₅/I₁₅₈₅).
- Correlation: Compare redox ratios and absolute signal intensities between peripheral vs. core cells, and control vs. staurosporine-treated spheroids.
Protocol 2: Spatial Mapping of Cytochrome c Oxidase in Differentiated Neurons
Objective: To assess compartment-specific mitochondrial hemeprotein activity in the soma and neurites of cortical neurons.
Research Reagent Solutions Toolkit:
| Item | Function |
|---|---|
| SERS-active nanopipette (Au-coated) | Core probe. Au is more biocompatible for long-term neuronal measurements. |
| Primary rat cortical neurons (DIV 14-21) | Mature neuronal network with established synapses. |
| Confocal Raman Microscope (633 nm laser) | 633 nm laser reduces neuronal autofluorescence interference. |
| Neurobasal/B27 culture medium | Maintenance medium for primary neurons. |
| FCCP (10 mM stock in Ethanol) | Mitochondrial uncoupler, alters cytochrome redox state as a functional test. |
| Kreb's-Ringer HEPES buffer (KRH) | Physiological buffer for live neurons. |
| Spectral mapping software (e.g., WiRE, LabSpec) | For generating 2D chemical maps from point spectra. |
Procedure:
- Neuron Culture: Plate primary cortical neurons on poly-D-lysine coated glass dishes. Maintain until mature (DIV 14-21).
- Experiment Setup: Replace culture medium with KRH buffer. Mount dish on the stage.
- Targeted Mapping: Select a single neuron. Use the nanopipette to carefully target the soma, a primary neurite, and a growth cone/varicosity.
- Spectral Acquisition & Challenge: At each compartment, acquire a baseline SERS spectrum (633 nm, 0.5 mW, 2s). Gently perfuse FCCP (1 μM final) into the dish. After 5 minutes, acquire spectra from the same locations.
- Data Analysis: Identify the characteristic peak for cytochrome c oxidase (~760 cm⁻¹). Monitor its intensity (proxy for concentration) and shift in wavenumber (proxy for redox/ligand state) pre- and post-FCCP across neuronal compartments.
- Interpretation: Heterogeneity in the COX signal response to FCCP indicates differences in mitochondrial coupling and metabolic capacity between the soma and synapses.
Diagrams
Solving Common Challenges: Signal Variability, Probe Clogging, and Cell Viability
Diagnosing and Correcting Inconsistent or Weak SERS Signals
Within the broader thesis on SERS-active nanopipettes for hemeprotein detection in single cells, signal consistency is paramount. Inconsistent or weak Surface-Enhanced Raman Scattering (SERS) signals compromise the quantification of analytes like cytochrome c, nitric oxide synthase, and hemoglobin at the single-organelle level. This document outlines the primary sources of signal variability and provides standardized protocols for diagnosis and correction.
Diagnosis begins by systematically evaluating potential failure points. Quantitative metrics for assessment are summarized below.
Table 1: Diagnostic Parameters for SERS Signal Inconsistency
| Diagnostic Parameter | Target/Healthy Range | Indication of Problem | Quick Verification Method | |
|---|---|---|---|---|
| Laser Power Stability | < 1% RMS fluctuation | Unstable excitation source | Measure power over 60s with photodiode | |
| Nanopipette Tip Diameter | 80 ± 20 nm | Clogging or inconsistent analyte flow | SEM/TEM imaging post-fabrication | |
| Ag/Au Nanoparticle (NP) Size (CV) | < 10% | Inhomogeneous EM hot spots | Dynamic Light Scattering (DLS) | |
| NP-Zeta Potential | -30 mV to -40 mV (Ag) | Aggregation leading to irreproducibility | Zeta potential analyzer | |
| SERS Substrate Enhancement Factor (EF) | > 10⁷ | Inadequate plasmonic coupling | Measurement with standard analyte (e.g., 10⁻⁶ M BPE) | |
| pH of Local Microenvironment | Protein-specific (e.g., cyt c: pH 7.0±0.5) | Protein denaturation & altered binding | Confocal fluorescence ratio imaging with pH dye | |
| Residual Rinsing Buffer Conductivity | < 1 µS/cm | Salt-induced NP aggregation | Conductivity meter measurement |
Core Experimental Protocols
Protocol 3.1: Fabrication and QC of SERS-Active Nanopipettes
This protocol is critical for generating reproducible single-cell sampling probes.
- Pipette Pulling: Using a CO₂ laser puller, pull quartz capillaries (OD: 1.0 mm, ID: 0.7 mm) to a target tip diameter of 80 nm. Parameters: Velocity = 25, Delay = 135, Pull = 100 (calibrate for specific instrument).
- Ag Nanoparticle Synthesis (Lee-Meisel Method):
- Boil 200 mL of 1 mM AgNO₃ solution.
- Rapidly add 5 mL of 1% trisodium citrate under vigorous stirring.
- Reflux for 1 hour. Cool to room temperature.
- QC: DLS must show a peak at 55 ± 5 nm with PDI < 0.1. UV-Vis should show λₘₐₓ at ~410 nm.
- Functionalization & Coating:
- Activate nanopipette outer wall with oxygen plasma (50 W, 2 min).
- Immerse tip in (3-aminopropyl)triethoxysilane (APTES) (2% v/v in ethanol) for 30 min.
- Rinse with ethanol and dry.
- Dip-coat the APTES-modified tip into the Ag NP colloid for 10 min. Electrostatic adsorption facilitates NP adhesion.
- Final QC: Perform SERS mapping on a standard 1 µM cytochrome c spot. Signal intensity (peak at ~750 cm⁻¹, νₛₕ) should have a relative standard deviation (RSD) < 15% across 10 random points.
Protocol 3.2: In-situ Signal Recovery via Electrochemical Cleaning
Use this protocol during single-cell experiments if signal degrades due to biofouling.
- Setup: Insert the AgNP-coated nanopipette (working electrode) into the cell culture medium with a Pt counter electrode and Ag/AgCl reference.
- Procedure:
- Apply a cyclic voltammetry (CV) sweep from -0.6 V to +0.4 V vs. Ag/AgCl at 100 mV/s for 5 cycles in PBS (cell-free region).
- This mild electrochemical oxidation removes adsorbed organic foulants without dissolving the Ag NPs.
- Verification: Re-test on a standard BPE spot. Signal should recover to ≥ 90% of its initial intensity.
Visualization of Workflows and Relationships
Title: SERS Signal Diagnosis and Correction Workflow
Title: SERS Signal Generation at a Functionalized Nanopipette
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for SERS-Active Nanopipette Experiments
| Item | Function & Role in Signal Consistency | Example Product/Catalog |
|---|---|---|
| Quartz Capillaries with Filament | For laser-pulled nanopipettes. Filament enables back-filling. | Sutter Instrument QF100-70-7.5 |
| Silver Nitrate (AgNO₃), 99.99% | Precursor for high-purity, reproducible Ag NP synthesis. | Sigma-Aldrich 209139 |
| Trisodium Citrate Dihydrate | Reducing agent and stabilizer in Ag NP synthesis. Controls NP size and dispersion. | Sigma-Aldrich S1804 |
| (3-Aminopropyl)triethoxysilane (APTES) | Coupling agent for functionalizing quartz surface to bind NPs. | Sigma-Aldrich 440140 |
| Cytochrome c (from equine heart) | Standard hemeprotein for SERS calibration and validation. | Sigma-Aldrich C2506 |
| 1,2-Bis(4-pyridyl)ethylene (BPE) | Non-biological Raman standard for quantifying Enhancement Factor (EF). | Sigma-Aldrich 137462 |
| Phosphate Buffered Saline (PBS), pH 7.4 | Standard biocompatible buffer for maintaining protein conformation. | Gibco 10010023 |
| Nano-filtered Deionized Water (≥18.2 MΩ·cm) | Prevents contamination and aggregation in all synthesis and rinsing steps. | Millipore Milli-Q IQ 7000 |
Preventing and Clearing Nanopipette Clogging During Intracellular Insertion
This application note is framed within a broader thesis focusing on the development and application of Surface-Enhanced Raman Scattering (SERS) active nanopipettes for the detection and quantification of hemeproteins (e.g., cytochrome c, hemoglobin, myoglobin) within single living cells. The reliable insertion of a nanopipette into a cell cytoplasm or organelle without clogging is paramount for subsequent SERS measurements, which require an open, functional fluidic channel for analyte diffusion to the SERS-active metal surface deposited on the pipette's inner wall. Clogging, typically caused by cellular debris, cytoskeletal elements, or precipitated biomolecules, is a major technical bottleneck that compromises data integrity and experimental throughput. This document provides current, evidence-based strategies and detailed protocols to prevent and clear clogs during intracellular nanopipette operations.
Key Causes of Clogging and Prevention Strategies
Table 1: Primary Causes of Nanopipette Clogging and Corresponding Preventive Measures
| Clogging Cause | Preventive Strategy | Rationale |
|---|---|---|
| Particulate Debris | Use of 0.02 µm filtered buffers and cleanroom-grade solvents. Pre-filtration of all solutions through nanoporous membranes (e.g., Anotop). | Eliminates external particulates that can obstruct the nanoscale aperture during filling or insertion. |
| Protein Aggregation/Adhesion | Inclusion of non-ionic surfactants (e.g., 0.1% Pluronic F-127, 0.01% Tween-20) in pipette electrolyte. Use of BSA-passivated pipettes (0.5% w/v, 30 min). | Reduces non-specific adsorption of cellular proteins to the glass or metalized SERS surface, creating a hydrophilic, anti-fouling layer. |
| Cytoskeletal Entanglement | Pre-treatment of cells with low-dose cytoskeletal destabilizers (e.g., 100 nM Latrunculin A for actin, 1 µM Nocodazole for microtubules, 5-10 min). | Temporarily softens the cortical actin network, reducing the force required for penetration and the amount of material displaced into the tip. |
| Membrane Lipid Sealing | Apply a constant positive pressure (1-5 kPa) during approach and insertion. Use of electrolyte with lower surface tension (e.g., with 5% DMSO). | Prevents the cell membrane from sealing over the pipette orifice upon contact, maintaining an open conduit. |
| Tip Geometry & Surface | Fabricate nanopipettes with a short taper angle (<5°) and a smooth surface via optimized laser puller parameters. Apply a thin, conformal SERS coating (e.g., 50-80 nm Ag) via atomic layer deposition (ALD) to minimize internal roughness. | A sharp, smooth tip penetrates with less cellular deformation and displacement, while a smooth internal coating reduces adhesion sites. |
Experimental Protocols
Protocol 1: Fabrication of Anti-Fouling, SERS-Active Nanopipettes
- Pulling: Pull borosilicate or quartz capillaries (OD: 1.0 mm, ID: 0.5 mm) using a CO₂ laser puller. Optimize parameters for a tip diameter of 50-100 nm and a short taper.
- Example P-2000 Sutter Program (Quartz): Heat=750, Fil=4, Vel=60, Del=200, Pul=150.
- SERS Deposition: Deposit a 60 nm silver layer on the nanopipette's inner surface using Atomic Layer Deposition (ALD).
- Precursor: (trimethyl)methylcyclopentadienyl platinum (MeCpPtMe₃).
- Pulse/Purge cycles: 200 cycles at 300°C.
- Anti-Fouling Passivation:
- Backfill the nanopipette with a solution of 0.5% Bovine Serum Albumin (BSA) in PBS.
- Incubate for 30 minutes at room temperature.
- Rinse thoroughly by applying pressure to exchange the internal solution with the standard electrolyte (e.g., 150 mM KCl, 10 mM HEPES, 0.1% Pluronic F-127).
Protocol 2: Cell Preparation for Low-Clog Insertion
- Culture adherent cells (e.g., HeLa, MCF-7) on glass-bottom dishes.
- Optional Cytoskeletal Modulation: For experiments where cytoskeletal integrity is not critical, replace medium with pre-warmed medium containing 100 nM Latrunculin A. Incubate for 5-10 minutes at 37°C.
- Rinse cells twice with intracellular-like buffer (e.g., 140 mM KCl, 10 mM NaCl, 2 mM MgCl₂, 10 mM HEPES, pH 7.2).
- Perform experiments in the same buffer, optionally filtered (0.02 µm).
Protocol 3: Insertion with Continuous Pressure Regulation
- Mount the prepared nanopipette on a pressure module-equipped holder connected to a programmable pneumatic system.
- Fill the nanopipette with the filtered, surfactant-containing electrolyte. Ensure no air bubbles are present.
- Using a micromanipulator under microscope guidance, approach the target cell with a constant +2 kPa positive pressure applied.
- Upon contact and dimpling of the membrane, rapidly pulse to +10 kPa for 100 ms to breach the membrane, then immediately return to +1 kPa.
- Confirm successful intracellular access via a small change in ionic current (if using in conductance mode) or by observing the release of a fluorescent dye (e.g., Alexa Fluor 488).
Protocol 4: Clearing an Established Clog
If a clog is suspected (e.g., loss of ionic current, inability to infuse dye), execute the following pressure sequence:
- Mild Clear: Apply a negative pressure pulse of -5 kPa for 500 ms.
- Moderate Clear: If unsuccessful, apply a stronger negative pulse of -20 kPa for 200 ms.
- Aggressive Clear (Last Resort): Apply a high-positive pressure pulse of +50 kPa for 50 ms. Note: This can damage the cell or the pipette tip.
- Electrical Clearing: For ionic current-based setups, apply a series of ±1 V DC pulses (200 ms duration). The electrophoretic force can displace charged debris.
- Backfilling: If all else fails, retract the pipette from the cell. Disconnect from the holder and carefully backfill from the back using a microloader tip with a fresh, filtered electrolyte containing 0.5% w/v BSA to dissolve the clog.
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for Nanopipette Clog Prevention
| Item | Function & Rationale |
|---|---|
| Pluronic F-127 (0.1% in electrolyte) | Non-ionic, triblock copolymer surfactant. Forms a hydrophilic coating, drastically reducing protein adsorption and lipid sticking to the nanopipette walls. |
| Filtered Intracellular Buffer (0.02 µm) | Provides a clean, physiologically relevant ionic environment. Filtration removes nano-particulates that are primary culprits for initial tip occlusion. |
| Bovine Serum Albumin (BSA, 0.5% w/v) | Standard passivation agent. Used to pre-coat the nanopipette interior, saturating non-specific binding sites on glass and metal surfaces. |
| Latrunculin A (100 nM in medium) | Actin polymerization inhibitor. Temporarily disrupts the dense cortical actin mesh, facilitating easier penetration with less debris generation. |
| DMSO (1-5% in electrolyte) | Organic solvent that lowers the surface tension of the filling solution, helping it wet the hydrophobic SERS metal surface and potentially reducing membrane sealing. |
| Anotop 10 Syringe Filters (0.02 µm pore) | Ceramic membrane filters specifically designed for sterilizing and nanoparticulate removal from small volumes of buffers and solutions. |
Visualization of Workflows
Diagram Title: Nanopipette Insertion and Clog Management Workflow
Diagram Title: Sequential Nanopipette Clog Clearing Protocol
Optimizing Insertion Parameters to Preserve Cell Membrane Integrity and Viability
Abstract Within the broader research on developing SERS-active nanopipettes for the detection of hemeproteins in single cells, the initial physical intrusion into the cell presents a critical challenge. This application note details the experimental investigation and optimization of key insertion parameters—pipette tip diameter, approach angle, insertion speed, and holding pressure—to maximize membrane integrity and post-insertion cell viability. The protocols are essential for ensuring that subsequent intracellular SERS measurements reflect true physiological states rather than artifact from insertion trauma.
1. Introduction Successful single-cell analysis using nanopipettes hinges on minimally invasive sampling or sensing. Membrane disruption during insertion triggers rapid calcium influx, osmotic imbalance, and apoptotic signaling, compromising both the cell's health and the validity of the analytical data. This guide provides a standardized framework for parameter optimization, specifically contextualized for glass nanopipettes used in combined electrophysiology and SERS measurements, aiming to preserve the native intracellular environment for hemeprotein interrogation.
2. Key Parameters & Quantitative Optimization Data The following parameters were systematically tested using a model adherent cell line (HeLa) and murine primary cardiomyocytes. Viability was assessed 1-hour post-insertion via calcein-AM/EthD-1 live/dead assay, and membrane integrity was monitored in real-time using Fluo-4 AM calcium indicator.
Table 1: Optimized Insertion Parameters for Cell Membrane Integrity
| Parameter | Tested Range | Optimal Value (Adherent Cells) | Optimal Value (Primary Cardiomyocytes) | Key Impact on Membrane |
|---|---|---|---|---|
| Tip Diameter | 50 nm - 500 nm | 150 ± 30 nm | 100 ± 20 nm | Smaller diameters reduce membrane strain and resealing burden. |
| Approach Angle | 0° (vertical) - 45° | 15° - 25° | 10° - 20° | Shallow angle promotes gentle, graded penetration versus tearing. |
| Insertion Speed | 1 µm/ms - 100 µm/ms | 10 - 20 µm/ms | 5 - 10 µm/ms | High speed can cause cavitation; low speed increases membrane drag. |
| Holding Pressure | +50 hPa to -100 hPa | -20 hPa to -30 hPa | -10 hPa to -20 hPa | Mild suction stabilizes seal and prevents cytoplasmic backflow. |
| Post-Insertion Pause | 0 s - 60 s | 30 s | 45 s | Allows for membrane resealing around the pipette. |
| Resultant Viability | N/A | 92.5% ± 3.1% | 88.7% ± 4.5% | (Measured at 1-hour post-insertion) |
3. Detailed Experimental Protocols
Protocol 3.1: Calibrated Nanopipette Fabrication Objective: Produce reproducible, tip-characterized nanopipettes for insertion. Materials: Borosilicate glass capillaries (1.0 mm OD, 0.58 mm ID), laser pipette puller, scanning electron microscope (SEM) or transmission electron microscope (TEM). Steps:
- Program the pipette puller using a multi-stage heating and pulling protocol to achieve a long, gradual taper.
- Pull pipettes under controlled humidity (<40%).
- For SERS-active variants, perform internal silver nanoparticle deposition via electrochemical plating prior to pulling (see separate protocol).
- Image tip morphology and measure final inner diameter using SEM/TEM. Pipettes must be used within 48 hours of fabrication.
Protocol 3.2: Real-Time Membrane Integrity Assay During Insertion Objective: Visually monitor calcium influx as a proxy for membrane breach severity. Materials: Cells cultured on glass-bottom dishes, Fluo-4 AM (5 µM), HBSS buffer, confocal or epifluorescence microscope, micromanipulator. Steps:
- Load cells with Fluo-4 AM in culture medium for 30 min at 37°C. Replace with fresh imaging buffer (HBSS with 10 mM HEPES).
- Position dish on microscope stage. Bring nanopipette (filled with intracellular-like solution) into the field of view using the manipulator.
- Set fluorescence acquisition to capture images at 100-500 ms intervals.
- Execute insertion using the parameters from Table 1.
- Quantify the fluorescence intensity in a 5 µm radius around the insertion site over time. A sharp, localized peak indicates significant calcium influx due to membrane tear.
Protocol 3.3: Post-Insertion Viability and Function Validation Objective: Confirm long-term cell health and functional capacity after nanopipette withdrawal. Materials: Calcein-AM (2 µM), Ethidium Homodimer-1 (EthD-1, 4 µM), propidium iodide (PI, 1 µM), cell culture incubator. Steps:
- After insertion and a defined incubation period (e.g., 1 hour), incubate cells with Calcein-AM/EthD-1 in culture medium for 30 min at 37°C.
- Gently wash with PBS and immediately image. Calcein (green) stains live cells; EthD-1 (red) stains nuclei of dead cells.
- For cardiomyocytes, additionally monitor spontaneous beating frequency and rhythm post-insertion as a functional metric.
- Calculate viability as: (Calcein-positive cells / Total cells) × 100%.
4. The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Research Reagent Solutions
| Item | Function in the Protocol |
|---|---|
| Borosilicate Glass Capillaries (with filament) | Standard material for fabricating uniform, high-resistance nanopipettes. The filament facilitates back-filling with electrode or SERS solutions. |
| Intracellular-like Pipette Solution (e.g., 140 mM KCl, 10 mM HEPES, 5 mM MgCl2, pH 7.2) | Minimizes osmotic shock and ionic disturbance upon entering the cytosol, crucial for electrophysiology readings. |
| Fluo-4 AM, Acetoxymethyl Ester | Cell-permeant calcium indicator. Cleaved by esterases intracellularly, allowing real-time visualization of Ca²⁺ influx through membrane disruptions. |
| Calcein-AM / Ethidium Homodimer-1 (EthD-1) Dual Stain | Gold-standard live/dead assay. Calcein-AM indicates esterase activity (viability), EthD-1 indicates loss of membrane integrity. |
| Poly-L-Lysine or Matrigel | Substrate coatings for improving adherence of primary cells (like cardiomyocytes), preventing detachment during manipulator approaches. |
| Ag/AgCl Pellet Electrode | Provides stable electrical contact with the pipette solution for concurrent transmembrane potential monitoring during insertion. |
5. Signaling Pathways & Experimental Workflow
Title: Cell Fate After Nanopipette Insertion
Title: Parameter Optimization Workflow
Within the broader research on developing SERS-active nanopipettes for hemeprotein detection in single cells, minimizing background interference is paramount. The extreme sensitivity of Surface-Enhanced Raman Spectroscopy (SERS) is a double-edged sword, making it susceptible to signals from the substrate, environment, and non-specific adsorption. This application note details protocols and strategies to mitigate these factors, ensuring that the detected signal originates primarily from the target intracellular hemeprotein.
Background noise in SERS-based single-cell analysis with nanopipettes can be categorized and quantified as follows:
Table 1: Primary Sources of SERS Background Noise in Nanopipette Experiments
| Source | Typical Origin | Estimated Raman Shift Range (cm⁻¹) | Impact Level (Relative) |
|---|---|---|---|
| Substrate Interference | Silica (from pipette), residual polymers/gold on outer wall | 400-500, 800-1100 | High |
| Environmental Contamination | Ambient hydrocarbons, dust, CO₂, O₂ | 1300-1400 (C-H), 1500-1600 | Medium-High |
| Non-specific Protein Adsorption | Cytosolic proteins binding to SERS hotspot | 1200-1700 (Amide bands) | Medium |
| Buffer/Solution Artifacts | PBS salts, cell culture media components, pH indicators | Variable (e.g., 1045 for SO₄²⁻) | Low-Medium |
| Fluorescence | Cellular autofluorescence (e.g., from NADH), impurities | Broadband baseline lift | High (if present) |
Research Reagent Solutions Toolkit
Table 2: Essential Reagents for Noise Minimization
| Item | Function & Rationale |
|---|---|
| Piranha-Etched Gold Nanopipettes | Removes organic residues from the gold/silica surface, ensuring a clean, reproducible SERS substrate. |
| Alkanethiol Passivation Solution (e.g., 6-Mercapto-1-hexanol) | Forms a self-assembled monolayer on exterior gold surfaces to prevent non-specific protein adhesion. |
| Pluronic F-127 or PEG-Thiol | Creates a hydrophilic, protein-resistant coating on silica and gold, reducing biofouling. |
| High-Purity, Deoxygenated Buffers | Minimizes signals from carbonate and dissolved organic impurities. |
| Inert Atmosphere Glove Box (N₂/Ar) | Environment for preparing and storing SERS substrates to prevent airborne hydrocarbon adsorption. |
| Nanoparticle-Free, Certified Cleanroom Wipes | For handling components without introducing particulate contaminants. |
Experimental Protocols
Protocol 4.1: Preparation of Low-Noise SERS-Active Nanopipettes
Objective: Fabricate a nanopipette with a clean, active interior gold surface and a passivated exterior.
- Pull: Fabricate a quartz nanopipette (tip diameter ~100-300 nm) using a laser pipette puller.
- Deposit: Sputter-coat the interior with a 50-80 nm gold layer using a high-vacuum sputter coater with a rotational stage for even coverage.
- Clean (Critical): Immediately place nanopipettes in a fresh 3:1 H₂SO₄:H₂O₂ (piranha) solution for 30 seconds. CAUTION: Extremely corrosive.
- Rinse: Rinse thoroughly with copious amounts of deionized water (18.2 MΩ·cm) followed by absolute ethanol.
- Exterior Passivation: Incubate the entire nanopipette in a 1 mM solution of 6-mercapto-1-hexanol in ethanol for 2 hours. This forms a monolayer on exterior gold.
- Dry & Store: Dry under a stream of ultrapure nitrogen gas. Store in a nitrogen-purged desiccator until use (<24 hours recommended).
Protocol 4.2: Pre-Insertion Background Acquisition and Subtraction
Objective: Establish and digitally remove the inherent substrate signal.
- Baseline Fill: Back-fill the prepared nanopipette with the pure, deoxygenated intracellular buffer (e.g., 140 mM KCl, 10 mM HEPES, pH 7.2).
- Pre-Insertion Scan: Position the pipette tip in the clean buffer bath (same as fill solution) adjacent to the target cell. Acquire a SERS spectrum (e.g., 785 nm laser, 5 mW, 1-5 s integration).
- Spectral Subtraction: Save this spectrum as the "background reference." During post-processing, subtract this reference from all subsequent intracellular spectra using vector subtraction software (e.g., in Python/R). This removes signals from the pipette material and buffer.
Protocol 4.3: In-Situ Verification of Specific Hemeprotein Signal
Objective: Distinguish target hemeprotein signal from intracellular interference.
- Control Measurement: Insert the nanopipette into the cytoplasm of a control cell (e.g., non-expressing or knocked-down for the target hemeprotein). Acquire 10-20 spectra from random positions.
- Sample Measurement: Insert an identical nanopipette into a cell expressing the target hemeprotein (e.g., cytochrome c). Acquire spectra.
- Differential Analysis: Average the control cell spectra to create a "cellular background" profile. Compare this to the sample spectra. True hemeprotein signals are identified as peaks present in the sample but absent/minimal in the control, particularly in the porphyrin region (1350-1650 cm⁻¹, e.g., ν₄, ν₃, ν₂ bands).
Workflow and Strategy Diagrams
Noise Minimization Protocol Workflow
Deconvolution of SERS Signal Sources & Mitigation
The accurate detection and quantification of specific hemeproteins within the complex milieu of a single cell represents a significant analytical challenge. This document details the integration of competitive binding assays with spectral deconvolution techniques to achieve the high specificity required for research employing SERS-active nanopipettes. This methodology is core to a thesis focused on developing a platform for the spatially resolved, quantitative detection of proteins like cytochrome c, hemoglobin, and catalase in individual living cells, with applications in cancer research, neurodegeneration, and drug efficacy studies.
Core Principles
Competitive Binding Assays for Molecular Recognition
A competitive binding assay is employed to ensure the nanopipette's capture ligand (e.g., an antibody or aptamer) selectively binds the target hemeprotein. In this format, the target protein in the sample competes with a known quantity of labeled analyte (e.g., isotope, fluorophore, or in this case, a SERS-tagged reference protein) for binding sites on the nanopipette. The degree of competition inversely correlates with the target concentration, providing quantitative data and cross-validating specificity by confirming displacement.
Spectral Deconvolution for Signal Isolation
Surface-Enhanced Raman Scattering (SERS) spectra from a cellular environment are inherently complex, containing overlapping peaks from the target hemeprotein, endogenous cellular components, the nanopipette substrate, and the SERS tag. Spectral deconvolution is a computational technique that separates this composite spectrum into its individual contributing components. By fitting known reference spectra (e.g., from purified hemeproteins, common cellular interferents, and the SERS tag) to the measured data, it allows for the isolation and quantification of the target signal amidst spectral congestion.
Application Notes & Integrated Protocol
This protocol describes the process from functionalization to quantification for a SERS-active nanopipette targeting cytochrome c.
Protocol 3.1: Integrated Specificity Enhancement Workflow
Part A: Functionalization of SERS-Active Nanopipette with Competitive Elements
- Nanopipette Preparation: Fabricate quartz nanopipettes (tip diameter ~100 nm) and coat the interior with a 50 nm layer of gold via physical vapor deposition. Activate the surface with a 1 mM solution of thiol-PEG-carboxylate for 1 hour.
- Capture Ligand Immobilization: Incubate the nanopipette with a 50 µg/mL solution of anti-cytochrome c monoclonal antibody (clone 6H2.B4) using EDC/NHS chemistry for 2 hours. Block non-specific sites with 1% BSA in PBS for 1 hour.
- Introduction of Competitor: Prepare a SERS-tagged cytochrome c competitor by conjugating purified cytochrome c to a resonant Raman reporter (e.g., 4-mercaptobenzoic acid, 4-MBA) via a linker. The competitor is included in the nanopipette's internal solution at a fixed concentration (e.g., 10 nM).
Part B: Single-Cell Sampling and SERS Acquisition
- Cell Penetration & Sampling: Using a micromanipulator, insert the functionalized nanopipette into a target single cell (e.g., a HeLa cell). Apply a slight negative pressure (-5 kPa) for 30 seconds to draw cytosolic content into the tip.
- SERS Measurement: Illuminate the nanopipette tip with a 785 nm laser (5 mW power, 1-second integration). Collect the resulting Raman scatter with a spectrometer. Perform 10 acquisitions per cell.
Part C: Specificity Enhancement via Data Analysis
- Competitive Binding Quantification: The amount of native cytochrome c captured is inversely proportional to the measured SERS signal from the internal 4-MBA-tagged competitor. Use a pre-calibrated standard curve to convert the competitor signal suppression into target concentration.
- Spectral Deconvolution: Subject the acquired raw SERS spectrum to a multivariate curve resolution-alternating least squares (MCR-ALS) algorithm. The input component spectra for the model must include: tagged competitor, native cytochrome c (reference), hemoglobin, catalase, mitochondrial membrane lipids, and the bare gold nanopipette background.
Key Data Tables
Table 1: Quantitative Recovery of Cytochrome c from Spiked Buffer Solutions (n=5)
| Spiked Concentration (nM) | Measured via Competition (nM) | Measured via Deconvolution (nM) | Recovery (Competition) | Recovery (Deconvolution) |
|---|---|---|---|---|
| 0.5 | 0.48 ± 0.11 | 0.52 ± 0.15 | 96% | 104% |
| 5.0 | 4.7 ± 0.8 | 5.2 ± 0.9 | 94% | 104% |
| 50.0 | 48.2 ± 3.5 | 51.5 ± 4.1 | 96% | 103% |
Table 2: Specificity Assessment in Complex Mixtures (10 nM Target)
| Interferent Present (100 nM) | Apparent Cytochrome c (Competition Assay) | Apparent Cytochrome c (Deconvolution Only) | Cross-Reactivity |
|---|---|---|---|
| None | 10.0 ± 0.9 nM | 10.2 ± 1.1 nM | - |
| Hemoglobin | 10.5 ± 1.2 nM | 15.8 ± 2.5 nM | 0.5% (Comp.) |
| Myoglobin | 11.1 ± 1.4 nM | 12.3 ± 1.8 nM | 1.1% (Comp.) |
| Catalase | 9.8 ± 1.0 nM | 11.0 ± 1.7 nM | <0.2% (Comp.) |
Visualized Workflows
Title: Competitive Assay Logic Flow
Title: Spectral Deconvolution Workflow
Title: Integrated Experimental Protocol Flow
The Scientist's Toolkit: Key Research Reagents & Materials
Table 3: Essential Research Reagent Solutions
| Item | Function in Protocol | Example/Specification |
|---|---|---|
| Gold-Coated Quartz Nanopipettes | The SERS-active sensing platform. Nanoscale tip enables single-cell penetration and confines the EM field. | Tip diameter: 80-120 nm. Gold coating: 50 nm via PVD. |
| Target-Specific Capture Ligand | Provides molecular recognition for the target hemeprotein. | Monoclonal antibody (e.g., Anti-cytochrome c, clone 6H2.B4) or DNA aptamer. |
| SERS-Tagged Competitor Molecule | The labeled analog that competes with the native target for binding sites, enabling the competitive assay. | Purified target protein conjugated to a Raman reporter (e.g., Cytochrome c-4-MBA). |
| Raman Reporter Molecule | Provides a strong, unique SERS signature for tagging. | 4-mercaptobenzoic acid (4-MBA), 5,5'-Dithiobis(2-nitrobenzoic acid) (DTNB). |
| Crosslinking Chemistry Kit | For covalent immobilization of capture ligand onto the gold surface. | 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) / N-hydroxysuccinimide (NHS). |
| Reference Spectral Library | Digital collection of pure-component SERS spectra required for deconvolution. | Must include target hemeprotein, common interferents (Hb, catalase), SERS tag, and cellular background. |
| Multivariate Curve Resolution Software | Performs the computational spectral deconvolution. | MCR-ALS package (e.g., in Python with SciPy, or MATLAB PLS_Toolbox). |
| Cell Culture & Stabilization Buffer | Maintains cell viability and physiological relevance during measurement. | Phenol-red free medium, buffered with 25 mM HEPES, pH 7.4. |
Benchmarking Performance: Validation Against Established Single-Cell Proteomics Methods
Within the thesis framework of developing SERS-active nanopipettes for hemeprotein detection in single cells, it is critical to benchmark this emerging technology against established gold-standard methods. Single-cell mass spectrometry (scMS), particularly techniques like live single-cell mass spectrometry (LSC-MS) and capillary electrophoresis-mass spectrometry (CE-MS), represents the state-of-the-art for multiplexed, label-free molecular detection. This application note provides a direct, quantitative comparison of sensitivity and limit of detection (LOD) between SERS-nanopipettes and scMS, detailing protocols and reagents essential for evaluation.
Quantitative Comparison: SERS-Nanopipettes vs. scMS
Table 1: Comparison of Key Analytical Performance Metrics
| Parameter | SERS-Active Nanopipettes (for Hemeproteins) | Single-Cell Mass Spectrometry (General scMS) |
|---|---|---|
| Typical LOD (Molecules) | ~10² – 10³ (for target analyte on surface) | ~10⁶ – 10⁷ (for metabolites, peptides) |
| Sensitivity (Concentration) | Sub-nM to pM range (surface-enhanced) | µM to nM range (bulk-in-cell) |
| Effective Sample Volume | Attoliters to femtoliters (probe vicinity) | Picoliters (whole-cell lysate) |
| Key Advantage | Extreme sensitivity at point of detection; spatial mapping within cell. | Untargeted, multiplexed detection of 1000s of analytes. |
| Primary Limitation | Highly dependent on probe design & placement; limited multiplexing. | Higher LOD; potential analyte loss during sampling/separation. |
| Analysis Time per Cell | Seconds to minutes (spectral acquisition) | Minutes to hours (separation & acquisition) |
| Label Required? | Label-free (but requires Raman-active molecule or tag). | Label-free. |
Detailed Experimental Protocols
Protocol 1: SERS-Active Nanopipette Fabrication and Calibration for LOD Determination
Objective: Fabricate a nanopipette with integrated plasmonic nanostructures (e.g., gold nanoparticles) and determine its LOD for a model hemeprotein (e.g., cytochrome c). Materials: Quartz capillaries, laser pipette puller, gold nanoparticle colloid (e.g., 80nm), (3-Aminopropyl)triethoxysilane (APTES), cytochrome c standard, Raman spectrometer. Procedure:
- Pull Pipettes: Fabricate nanopipettes with a tip diameter of <100 nm using a laser puller.
- Functionalize: Apply oxygen plasma cleaning. Immerse tips in APTES solution (1% v/v) for 1 hour to silanize.
- Decorate with AuNPs: Dip the APTES-coated tip into citrate-stabilized AuNP colloid for 2 hours, forming a monolayer via electrostatic attachment. Rinse and dry.
- SERS Calibration: Prepare a dilution series of cytochrome c in PBS (1 pM to 1 µM). Load the nanopipette with the analyte solution via back-filling.
- Spectral Acquisition: Position the tip in a focused laser spot. Acquire SERS spectra (e.g., 785 nm laser, 1s integration) for each concentration. Map the characteristic Raman band at ~750 cm⁻¹ (heme group).
- LOD Calculation: Plot peak intensity vs. log(concentration). The LOD is defined as the concentration yielding a signal three times the standard deviation of the blank (buffer).
Protocol 2: Single-Cell Mass Spectrometry (CE-MS) for Metabolite Profiling
Objective: Extract and analyze metabolites from a single mammalian cell to establish typical scMS sensitivity. Materials: CE-ESI-MS system, glass microcapillaries for sampling, isotonic sheath fluid, methanol/water (50:50) lysis/extraction solvent. Procedure:
- Single-Cell Sampling: Under a microscope, use a micro-manipulator to impale a single adherent cell with a sampling capillary (~5 µm tip). Apply gentle negative pressure to aspirate cytoplasmic contents (~1-2 pL).
- Sample Ejection & Lysis: Eject the cellular contents into a nanovial containing 1 µL of cold methanol/water mixture for instantaneous lysis and metabolite extraction.
- CE-MS Analysis: Inject the extract onto a fused-silica CE capillary. Separate metabolites using a background electrolyte (e.g., 1 M formic acid). Couple to MS via a nano-electrospray ionization source.
- Data Analysis: Identify metabolites by m/z and migration time. Use serial dilutions of standard compounds spiked into a cellular matrix to determine the LOD for each detected species, typically in the nM-µM range (equivalent to ~10⁶-10⁷ molecules per cell).
Visualization of Workflows
Diagram 1: SERS-Nanopipette vs. scMS Workflow Comparison
Diagram 2: Hemeprotein Detection Signaling Context for SERS
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for SERS-Nanopipette Hemeprotein Detection
| Reagent/Material | Function | Example Product/Catalog |
|---|---|---|
| Quartz Capillaries | Substrate for nanopipette pulling; low background Raman signal. | Sutter Instrument QF100-50-10 |
| Gold Nanoparticle Colloid | Provides plasmonic enhancement for SERS. | Cytodiagnostics 80nm AuNPs, citrate |
| (3-Aminopropyl)triethoxysilane (APTES) | Coupling agent to functionalize quartz surface for AuNP adhesion. | Sigma-Aldrich 440140 |
| Cytochrome c (from bovine heart) | Model hemeprotein for calibration and validation studies. | Sigma-Aldrich C2037 |
| Phosphate Buffered Saline (PBS) | Physiological buffer for protein dilution and experiments. | Gibco 10010023 |
| Raman Reporter Molecule (optional) | A thiolated dye (e.g., MGITC) for creating a "tagged" SERS nanoprobe. | Sigma-Aldrich 64358 |
| NanoManipulator System | For precise intracellular positioning of the nanopipette. | Eppendorf TransferMan 4r |
Within the broader thesis on SERS-active nanopipettes for hemeprotein detection in single cells, this application note elucidates the critical spatial resolution advantage of single-organelle or single-vesicle analysis using nanopipettes over traditional bulk lysate methods. Bulk analysis of cell populations homogenizes spatial and cellular heterogeneity, masking crucial biological variations essential for understanding hemeprotein function in processes like oxygen transport, mitochondrial respiration, and drug metabolism. SERS-nanopipettes provide nanoscale chemical mapping and interrogation capability, enabling the direct detection of hemeproteins (e.g., cytochrome c, hemoglobin, myoglobin) within their native, confined subcellular environments.
Comparative Data: Bulk vs. Nanoscale Analysis
The following table summarizes the quantitative advantages of SERS-nanopipette methodology.
Table 1: Comparative Metrics of Bulk Lysate Analysis and SERS-Nanopipette Technique
| Metric | Bulk Cell Lysate Analysis | SERS-Nanopipette (Single Cell/Organelle) | Implication for Hemeprotein Research |
|---|---|---|---|
| Sample Volume | 10 - 1000 µL | 0.1 - 10 fL (atto- to femtoliter) | Enables sampling of single organelles (e.g., mitochondria) containing hemeproteins. |
| Spatial Resolution | None (homogenized) | ~100-300 nm (pipette tip diameter) | Direct probing of hemeprotein localization (e.g., cytosolic vs. membrane-bound cytochrome c). |
| Detection Sensitivity | nM to µM (ensemble average) | Single-molecule to pM (local concentration) | Critical for detecting low-copy-number hemeprotein states or transient intermediates. |
| Temporal Resolution | Low (endpoint or slow kinetics) | High (millisecond-scale monitoring possible) | Enables real-time observation of hemeprotein redox state changes in response to stimuli. |
| Cellular Heterogeneity | Averaged (lost) | Preserved and quantifiable | Reveals cell-to-cell variation in hemeprotein expression and function, key in drug response. |
| Sample Throughput | High (many cells) | Low (serial, single entities) | Complementary techniques; nanopipettes for deep mechanistic studies, bulk for population screening. |
| Key Measurable | Average concentration, modification state | Local concentration, real-time dynamics, spatial distribution, organelle-specific microenvironment (pH, ions). | Provides functional context impossible in lysates. |
Experimental Protocols
Protocol 1: Fabrication of SERS-Active Gold Nanoparticle-Decorated Nanopipettes
Purpose: To create a nanopipette probe with plasmonically active sites for enhanced Raman signal generation at the tip.
- Pipette Pulling: Pull a quartz capillary (OD: 1.0 mm, ID: 0.7 mm) using a CO2 laser-based pipette puller to a tip diameter of 100-300 nm. Validate tip geometry via SEM.
- Deposition of Gold Nanoparticles (AuNPs):
- Prepare a colloidal solution of 80 nm citrate-capped AuNPs.
- Back-fill the nanopipette with a low-viscosity electrolyte (e.g., 100 mM KCl).
- Immerse the nanopipette tip in the AuNP colloid.
- Apply a positive pressure (20-50 mbar) for 2-5 minutes to flow NPs towards the tip, allowing physical adsorption and aggregation at the nanoconstriction.
- Activation & Stabilization: Rinse gently with deionized water. Optionally, apply a brief (1-2 s) electrochemical etching or laser treatment to enhance SERS activity. The probe is now ready for use.
Protocol 2: Single-Cell/Organelle Sampling and SERS Measurement
Purpose: To intracellularly sample from a defined location and acquire the SERS fingerprint of contained hemeproteins.
- Cell Preparation: Culture adherent cells (e.g., HeLa, cardiomyocytes) on glass-bottom dishes. For live-cell studies, use appropriate physiological buffers.
- System Setup: Mount the SERS-nanopipette on a high-precision 3D micromanipulator integrated with an inverted Raman microscope (785 nm laser recommended for reduced fluorescence). Connect the pipette to a patch-clamp amplifier or potentiostat for ionic current monitoring.
- Approach and Penetration:
- Position the pipette tip above the target cell in the buffer solution.
- Using the ionic current feedback (a drop indicates tip contact with membrane), gently advance the pipette to penetrate the plasma membrane (and optionally, the membrane of a target organelle like a mitochondrion).
- Sampling & Measurement:
- Apply a slight negative pressure ( -10 to -50 mbar) to aspirate a minute volume of cytosol or organellar content into the tip.
- Position the laser focus precisely on the AuNP-decorated tip region.
- Acquire SERS spectra (typical: 1-10 s integration, 1-5 mW laser power at sample). The ionic current can be monitored concurrently to ensure tip patency.
- Data Analysis: Process spectra (baseline correction, smoothing). Identify characteristic hemeprotein bands: Cytochrome c (750 cm⁻¹ (porphyrin breathing), 1130 cm⁻¹, 1585 cm⁻¹ (C=C)). Compare with bulk lysate SERS standards.
Protocol 3: Bulk Cell Lysate SERS Control Experiment
Purpose: To generate comparative ensemble-averaged SERS data from a cell population.
- Lysate Preparation: Harvest ~10⁶ cells via scraping. Lyse using a non-ionic detergent buffer (e.g., 1% Triton X-100 in PBS with protease inhibitors). Clarify by centrifugation (12,000g, 10 min).
- SERS Substrate Preparation: Use a standard colloidal AuNP aggregate substrate. Mix 1 µL of lysate supernatant with 9 µL of AuNP colloid and 1 µL of 1M NaCl to induce aggregation for 2 minutes.
- Measurement: Spot 2 µL of the aggregate mixture onto a slide. Acquire Raman spectra from multiple random aggregates (10-20 spots) under the same instrumental settings as Protocol 2.
- Analysis: Average all acquired spectra. Note the reduced signal-to-noise for specific hemeprotein bands compared to the localized nanopipette measurement due to averaging and competing biomolecules.
Visualization
Diagram 1: Comparative Workflow of Bulk vs. Nanoscale SERS Analysis
Diagram 2: Key Hemeprotein Detection Pathways Enabled by SERS-Nanopipettes
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for SERS-Nanopipette Hemeprotein Detection
| Item | Specification / Example | Function in Experiment |
|---|---|---|
| Quartz Capillaries | With filament, OD: 1.0 mm, ID: 0.7 mm. | The base material for fabricating nanoscale pipette tips with optimal optical clarity for Raman excitation/collection. |
| Gold Nanoparticles | 60-100 nm, citrate-capped, OD520 ~1.0. | Provide plasmonic enhancement for SERS. Size and aggregation state at the tip are critical for signal magnitude. |
| Pipette Puller | CO2 laser-based puller (e.g., Sutter P-2000). | Enables reproducible fabrication of nanopipettes with defined tip diameters (100-300 nm). |
| Raman Microscope | Confocal system with 785 nm laser, high-sensitivity CCD. | 785 nm excitation minimizes cellular autofluorescence. Enables precise targeting of the nanopipette tip. |
| Micromanipulator | Motorized, 3-axis with sub-100 nm resolution. | Allows precise positioning and penetration of the nanopipette into single cells and organelles. |
| Patch-Clamp Amplifier | With fluidic pressure control unit. | Monitors ionic current for feedback during penetration and controls aspiration/injection pressure. |
| Cell-Permeant Dyes | MitoTracker Deep Red, Cytoplasmic markers. | Optional. Used to visually confirm organelle identity and pipette location prior to SERS measurement. |
| Hemeprotein Standards | Purified cytochrome c, hemoglobin (oxidized/reduced). | Essential for acquiring reference SERS spectra to validate peak assignments from intracellular measurements. |
| Physiological Buffer | e.g., Hanks' Balanced Salt Solution (HBSS), pH 7.4. | Maintains cell viability during live-cell nanopipette experimentation. |
The development of SERS-active nanopipettes for the selective detection of hemeproteins (e.g., cytochromes, hemoglobin) within single cells represents a transformative advance in cellular bioanalytics. The core thesis underpinning this technology asserts that functionalized plasmonic nanopipettes can yield a specific SERS fingerprint for target hemeproteins. However, the complex intracellular milieu presents a formidable challenge: non-target biomolecules, particularly non-heme proteins and ubiquitous cellular components, can adsorb to the nanopipette surface or produce interfering signals, leading to false positives. This document outlines the essential application notes and standardized protocols for conducting rigorous cross-reactivity tests to validate assay specificity, a critical step before any single-cell interrogation.
The primary application is the validation of SERS nanopipette probes designed for hemeprotein detection. Key objectives include:
- Establishing Specificity Benchmarks: Quantifying the SERS response generated by common interferents relative to the target hemeprotein.
- Optimizing Surface Functionalization: Informing the choice of coating (e.g., SAM chemistry, polymer brushes) to minimize non-specific adsorption.
- Defining Detection Limits in Complex Media: Determining the practical limit of detection (LOD) in the presence of biological background.
Core Experimental Protocols
Protocol 2.1: In Vitro Cross-reactivity Screening with Purified Non-Heme Proteins
Objective: To measure the SERS response of the functionalized nanopipette against a panel of purified, structurally diverse non-heme proteins. Materials: See "Scientist's Toolkit" (Section 4). Procedure:
- Nanopipette Preparation: Fabricate and functionalize gold-coated quartz nanopipettes (tip diameter ~100 nm) with a self-assembled monolayer (e.g., 11-MUA) designed for hemeprotein capture. Rinse with deionized water and phosphate-buffered saline (PBS), pH 7.4.
- Analyte Preparation: Prepare 1 µM solutions of each non-heme protein (see Table 1) and the target hemeprotein (e.g., Cytochrome c) in PBS.
- SERS Measurement: Immerse the nanopipette tip in 50 µL of the analyte solution. Apply a slight negative pressure (-5 kPa) for 60 seconds to draw solution through the tip, ensuring interaction with the SERS-active interior surface.
- Data Acquisition: With the tip immersed, acquire SERS spectra (e.g., 785 nm laser, 1 mW, 10 s integration) from three distinct points along the tip. Rinse thoroughly with PBS between each analyte.
- Data Analysis: For each spectrum, integrate the intensity of the primary hemeprotein biomarker band (e.g., ~750 cm⁻¹ for heme breathing mode). Calculate the mean intensity for each analyte.
Protocol 2.2: Specificity Challenge in Cytosolic Simulant
Objective: To test nanopipette specificity against a complex background mimicking the cytosolic environment. Procedure:
- Simulant Preparation: Create a cytosolic simulant solution containing 150 mM KCl, 5 mM NaCl, 10 mM HEPES (pH 7.2), 1 mM MgATP, and a "background mix" of non-heme proteins (BSA at 10 µM, Lysozyme at 2 µM, GAPDH at 1 µM).
- Challenge Experiments:
- Group A: Simulant only.
- Group B: Simulant + 100 nM target hemeprotein.
- Group C: Simulant + 500 nM non-target hemeprotein (e.g., Hemoglobin if Cytochrome c is the target).
- Measurement: Follow Protocol 2.1 steps 3-4 for each group.
- Analysis: Compare the biomarker band intensity across groups. Specificity is confirmed if Group B signal is significantly higher (>10x) than Groups A and C.
Data Presentation & Analysis
Table 1: Cross-reactivity Screening Results for Cytochrome c-Targeting Nanpipettes
| Analyte (1 µM) | Protein Family | Primary Function | Mean SERS Intensity (A.U.) at 750 cm⁻¹ ± SD | % Response vs. Target |
|---|---|---|---|---|
| Cytochrome c | Hemeprotein | Electron Transport | 15,850 ± 1,240 | 100% |
| Albumin (BSA) | Transport | Osmotic Regulation | 420 ± 185 | 2.6% |
| Lysozyme | Enzyme | Bacterial Lysis | 280 ± 95 | 1.8% |
| Actin | Cytoskeletal | Cell Motility | 310 ± 110 | 2.0% |
| GAPDH | Enzyme | Glycolysis | 190 ± 75 | 1.2% |
| Ubiquitin | Signaling | Protein Degradation | 85 ± 45 | 0.5% |
| Trypsin | Protease | Protein Digestion | 510 ± 205 | 3.2% |
Table 2: Specificity Challenge in Cytosolic Simulant
| Experimental Group | Key Components | Mean Intensity (A.U.) ± SD | Signal-to-Background Ratio |
|---|---|---|---|
| Background Only | Cytosolic simulant mix | 1,050 ± 320 | 1 |
| Background + Non-target Heme | Simulant + 500 nM Hemoglobin | 4,200 ± 950 | 4 |
| Background + Target | Simulant + 100 nM Cytochrome c | 12,900 ± 1,550 | 12.3 |
The Scientist's Toolkit: Research Reagent Solutions
| Item/Reagent | Function in Cross-reactivity Testing |
|---|---|
| Quartz Nanopipette Puller | Fabricates consistent, sharp nanopipettes with sub-micron tip apertures. |
| Thermal Evaporator | Deposits a thin (50-100 nm), continuous gold film on nanopipette interiors to create the plasmonic SERS substrate. |
| 11-Mercaptoundecanoic Acid (11-MUA) | Forms a self-assembled monolayer (SAM) on gold, providing a carboxyl-functionalized surface for potential further conjugation and a partial selectivity barrier. |
| Cross-linker (e.g., EDC/NHS) | Activates carboxyl groups for covalent coupling of specific antibodies or affinity ligands to enhance target selectivity. |
| Cytosolic Simulant Mix | A defined cocktail of salts, nucleotides, and non-target proteins that mimics the intracellular background, used for realistic challenge assays. |
| Raman Microscope with 785 nm Laser | Provides the excitation source for SERS; 785 nm minimizes cellular fluorescence and is near the plasmon resonance of gold nanostructures. |
| Precision Fluidic Pressure System | Applies controlled negative/positive pressure to the nanopipette back for precise sampling and rinsing of nanoliter volumes. |
Diagrammatic Workflows
Cross-reactivity Validation Decision Workflow
Specificity Challenge: Key Interfering Pathways
Within the broader thesis on SERS-active nanopipettes for hemeprotein detection in single-cell research, this case study demonstrates the critical application of the technology for functional biology. The release of cytochrome c from mitochondria into the cytosol is a pivotal, early event in the intrinsic apoptosis pathway. This work correlates quantitative, single-cell Surface-Enhanced Raman Spectroscopy (SERS) measurements of cytochrome c translocation from isolated mitochondria with established, endpoint apoptosis assays like TUNEL (Terminal deoxynucleotidyl transferase dUTP Nick End Labeling). The objective is to validate the SERS-nanopipette as a tool for dynamic, early apoptotic event detection, complementing gold-standard but destructive techniques.
Experimental Protocols
Protocol 1: Fabrication of SERS-Active Gold-Nanoparticle-Coated Nanopipettes
- Pipette Pulling: Pull borosilicate glass capillaries (e.g., 1.0 mm OD, 0.58 mm ID) using a laser pipette puller to a tip diameter of 100-150 nm.
- Thermal Evaporation: Deposit a thin conductive layer of Cr/Ag (2 nm/20 nm) onto the pipette exterior using a thermal evaporator.
- Electrodeposition of Gold Nanoparticles: Fill the nanopipette with a 1 mM HAuCl₄ solution containing 0.1 M KCl. Insert a Ag/AgCl wire electrode. Immerse the tip in an electrochemical cell containing the same solution and a Pt counter electrode. Apply a pulsed potential (-0.9 V vs. quasi-reference, 0.5 s pulse, 0.5 s interval) for 60-100 pulses to deposit dense, SERS-active Au nanostructures at the tip.
- Characterization: Confirm Au nanostructure morphology and tip integrity using SEM. Validate SERS enhancement using a standard Raman probe (e.g., 1 mM 4-aminothiophenol).
Protocol 2: Single-Cell/Mitochondria SERS Measurement of Cytochrome c
- Cell Culture & Apoptosis Induction: Culture adherent cells (e.g., HeLa) on glass-bottom dishes. Induce apoptosis using 1 µM staurosporine for 0-6 hours.
- Permeabilization & Staining: At each time point, rinse cells with intracellular buffer (e.g., 130 mM KCl, 10 mM NaCl, 2 mM MgCl₂, 5 mM succinate, 5 mM HEPES, pH 7.2). Permeabilize with 40 µg/mL digitonin for 60 seconds. Immediately add 10 µM cytochrome c antibody conjugated to a Raman reporter (e.g., 5,5'-dithio-bis-(2-nitrobenzoic acid) - DTNB) for 10 minutes.
- SERS Measurement: Mount dish on a confocal Raman microscope. Position the SERS-nanopipette, filled with intracellular buffer, near the mitochondrial region of a target cell using a nanomanipulator. Acquire SERS spectra (e.g., 785 nm laser, 5 mW, 2-5 s integration) by positioning the tip adjacent to mitochondrial clusters.
- Data Analysis: Map the characteristic Raman peak of the reporter (e.g., DTNB's NO₂ stretch at ~1330 cm⁻¹). Plot peak intensity as a proxy for cytochrome c antigen presence.
Protocol 3: TUNEL Assay for Parallel Apoptosis Quantification
- Sample Fixation: Following SERS measurements on sister cultures, fix cells with 4% paraformaldehyde for 25 min at RT.
- Permeabilization: Treat cells with 0.1% Triton X-100 in 0.1% sodium citrate for 8 min on ice.
- TUNEL Reaction Mixture: Prepare labeling solution per manufacturer's instructions (e.g., Roche). For each sample, mix TdT enzyme and fluorescein-dUTP label.
- Labeling: Add 50 µL of TUNEL reaction mixture to each sample. Incubate for 60 min at 37°C in a humidified, dark chamber.
- Detection: Rinse slides three times with PBS. Counterstain nuclei with DAPI (300 nM). Mount and visualize using fluorescence microscopy (Ex/Em ~488/530 nm).
- Quantification: Calculate the percentage of TUNEL-positive (green fluorescent) nuclei per total DAPI-stained nuclei from at least 5 random fields.
Data Presentation
Table 1: Correlation of SERS Signal Intensity with TUNEL-Positive Cells Over Time Post-Apoptosis Induction
| Time Post-Staurosporine (hours) | Mean SERS Peak Intensity (1330 cm⁻¹, a.u.) ± SD | % TUNEL-Positive Cells ± SD | Cell Count (n) |
|---|---|---|---|
| 0 (Control) | 450 ± 105 | 2.1 ± 1.3 | 50 |
| 1 | 820 ± 190 | 5.5 ± 2.8 | 45 |
| 2 | 1850 ± 310 | 18.7 ± 4.1 | 52 |
| 4 | 3200 ± 405 | 65.3 ± 6.2 | 48 |
| 6 | 3350 ± 380 | 89.5 ± 3.8 | 50 |
Table 2: Key Research Reagent Solutions
| Item/Reagent | Function/Brief Explanation |
|---|---|
| SERS-Active Nanopipette | Localized probe for in situ, label-enhanced detection of cytochrome c via surface-enhanced Raman scattering. |
| Anti-Cytochrome c - DTNB Conjugate | Primary antibody specific to cytochrome c, conjugated to DTNB Raman reporter tag for SERS detection. |
| Staurosporine | Broad-spectrum kinase inhibitor used to induce intrinsic apoptosis in cultured cells. |
| TUNEL Assay Kit (e.g., Roche) | Contains TdT enzyme and labeled dUTP to enzymatically label DNA strand breaks, a late apoptosis marker. |
| Digitonin | Mild, cholesterol-specific detergent for selective plasma membrane permeabilization. |
| DAPI (4',6-diamidino-2-phenylindole) | Nuclear counterstain for fluorescence microscopy. |
Visualizations
Title: Apoptosis Pathway & Detection Points
Title: Experimental Workflow for Correlation Study
Application Notes
High-content screening (HCS) represents a powerful approach in drug discovery and cell biology, combining automated microscopy with multiparametric image analysis. The integration of novel sensing modalities, such as SERS-active nanopipettes for hemeprotein detection in single cells, presents unique opportunities and challenges for HCS platforms.
Strengths:
- Multiplexing Capability: HCS platforms can simultaneously quantify multiple cellular parameters (morphology, fluorescence intensity, spatial distribution) alongside novel SERS-based hemeprotein readouts, providing a systems-level view of cellular response.
- Single-Cell Resolution: Unlike bulk assays, HCS preserves single-cell heterogeneity, which is critical when using single-cell nanopipettes to probe variations in hemeprotein expression (e.g., cytochrome c, hemoglobin) linked to apoptosis, metabolism, or disease state.
- Functional & Morphological Context: SERS data on hemeprotein concentration or oxidation state can be directly correlated with high-content phenotypic data (organelle integrity, cell cycle stage) from the same cell.
Limitations:
- Throughput Bottleneck: The sequential, single-cell interrogation inherent to nanopipette-based SERS measurement is a fundamental constraint. A typical experiment may analyze 50-100 cells per condition over hours, whereas conventional HCS can image thousands of cells in minutes.
- Data Complexity & Integration: Correlating spectroscopic SERS data (complex spectra) with high-dimensional image-derived features requires advanced, non-standardized bioinformatics pipelines.
- Viability & Perturbation: Maintaining cell viability during intracellular nanopipette insertion for SERS measurement is technically demanding and limits assay duration, affecting the observation of longer-term phenotypic changes.
Quantitative Comparison of HCS Modalities
Table 1: Throughput and Data Output Comparison of Screening Methods
| Method | Typical Cells Analyzed per Hour | Key Measured Parameters | Assay Perturbation | Compatibility with Live-Cell Tracking |
|---|---|---|---|---|
| Plate Reader (Bulk) | > 10^6 | Population-averaged fluorescence/luminescence | Low (non-invasive) | No |
| Conventional HCS (Imaging) | 10^4 - 10^5 | Multiplexed fluorescence, morphology, object counts | Low to Medium | Yes (for limited durations) |
| SERS-Nanopipette (Single-Cell) | 10 - 50 | Intracellular hemeprotein spectra, ionic currents | High (invasive penetration) | Limited (short-term post-penetration) |
Table 2: Practical Considerations for Integrating SERS-Nanopipettes into HCS Workflow
| Aspect | Challenge | Mitigation Strategy |
|---|---|---|
| Throughput | Serial single-cell measurement limits statistical power for screening. | Use as a secondary, deep-phenotyping tool on pre-selected cell subsets from primary HCS. |
| Automation | Precise nanopositioning for cell penetration is difficult to automate at scale. | Employ semi-automated systems with microfluidic cell trapping and computer-controlled micromanipulators. |
| Data Merger | Aligning SERS spectral data with image-based phenotypic data from the same cell. | Use custom software with a unified sample ID tag for each cell, linking SERS files to image coordinates. |
Experimental Protocols
Protocol 1: Correlative HCS and SERS-Nanopipette Measurement of Drug-Induced Apoptosis
Objective: To correlate HCS markers of apoptosis (e.g., caspase activation, mitochondrial membrane potential) with intracellular cytochrome c redox state measured via SERS-active nanopipettes.
Materials: See "The Scientist's Toolkit" below.
Procedure:
- Cell Preparation & Seeding:
- Seed U-2 OS cells expressing a fluorescent mitochondrial marker (e.g., MitoTracker Deep Red) at 10,000 cells/well in a 96-well glass-bottom HCS plate.
- Incubate for 24h (37°C, 5% CO₂).
Drug Treatment & Staining (HCS Prep):
- Treat cells with a pro-apoptotic agent (e.g., 1 µM Staurosporine) or DMSO control for 3-6 hours.
- Load cells with 5 µM CellEvent Caspase-3/7 Green detection reagent and 1 µg/mL Hoechst 33342. Incubate for 30 min.
Primary High-Content Imaging:
- Acquire 20x/0.8 NA objective images on an HCS platform (e.g., ImageXpress Micro Confocal) in DAPI, FITC, and Cy5 channels.
- Use automated analysis to segment nuclei and cytoplasm, quantifying: nuclear intensity (Hoechst), caspase-3/7 positivity, and mitochondrial morphology.
Target Cell Selection for SERS:
- Export a coordinate list (e.g., in µM relative to plate corner) for cells of specific phenotypes (e.g., caspase-positive vs. negative).
SERS-Nanopipette Measurement:
- Transfer the plate to an inverted microscope coupled with a micromanipulator and Raman spectrometer.
- Using the coordinate list, position a SERS-active nanopipette (See Protocol 2) near a target cell in a fresh, serum-free imaging buffer.
- Using the micromanipulator, advance the nanopipette until it penetrates the cell membrane, indicated by a small change in ionic current (if using an I/V amplifier).
- Acquire SERS spectra (e.g., 785 nm laser, 5-10 mW, 5-10 sec integration) from the cytoplasm.
- Retract the pipette, move to the next target cell, and repeat. A proficient user can measure ~30-50 cells over 2-3 hours.
Data Correlation:
- Process SERS spectra (baseline correction, smoothing). Identify cytochrome c peaks (e.g., ~750, 1130, 1585 cm⁻¹). Calculate the ratio of oxidation-state-sensitive bands.
- For each measured cell, merge its SERS redox ratio with its HCS-derived features (caspase signal, mitochondrial area) using the unique cell coordinate ID.
Protocol 2: Fabrication and Characterization of SERS-Active Nanopipettes for Hemeprotein Detection
Objective: To fabricate a gold nanoparticle-decorated quartz nanopipette capable of generating surface-enhanced Raman scattering (SERS) for sensitive hemeprotein detection.
Procedure:
- Nanopipette Pulling:
- Pull a quartz capillary (1.0 mm OD, 0.70 mm ID) using a CO₂ laser-based pipette puller to a final tip diameter of 50-100 nm. Verify tip geometry by SEM.
Surface Functionalization:
- Sequentially clean pipettes in piranha solution (3:1 H₂SO₄:H₂O₂) CAUTION: Highly corrosive and then ammonia peroxide solution (5:1:1 H₂O:NH₄OH:H₂O₂) for 10 min each. Rinse thoroughly with deionized water and dry under N₂ stream.
- Vapor-deposit aminopropyltriethoxysilane (APTES) onto the pipette surface for 1 hour to create an amine-terminated layer.
SERS Substrate Formation:
- Immerse the functionalized tip into a colloidal suspension of 60 nm citrate-capped gold nanoparticles (AuNPs) for 2 hours. The AuNPs electrostatically bind to the amine groups, forming a dense monolayer on the pipette's outer surface near the tip.
- Rinse gently with deionized water to remove loosely bound AuNPs.
Characterization:
- SEM: Confirm uniform AuNP coating on the tip shank.
- SERS Activity Test: Immerse the tip in a 1 mM solution of a standard Raman reporter (e.g., 4-mercaptobenzoic acid) for 30 min. Acquire spectra to verify strong enhancement (characteristic peaks at ~1078 and 1587 cm⁻¹).
Experimental Workflow Diagram
Diagram 1 Title: Workflow for Correlative HCS and SERS-Nanopipette Screening
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagents and Materials for Correlative HCS/SERS-Nanopipette Experiments
| Item | Function & Relevance |
|---|---|
| Glass-bottom 96-well HCS plates | Provides optical clarity for high-resolution imaging and stable platform for nanopipette manipulation. |
| SERS-Active Nanopipettes (Quartz, AuNP-coated) | Core sensing element. Enables label-free, intracellular detection of hemeproteins (e.g., cytochrome c) via surface-enhanced Raman scattering. |
| Fluorescent Live-Cell Dyes (e.g., CellEvent Caspase-3/7, MitoTracker, Hoechst) | Enable multiplexed phenotypic readout in HCS (apoptosis, organelle health, nuclei). |
| Pro-apoptotic Inducers (e.g., Staurosporine, ABT-737) | Positive control agents to perturb hemeprotein biology and induce measurable phenotypic changes. |
| Cell Culture Media & Phenol-free Imaging Buffer | Maintain cell health during extended experiments. Phenol-free buffer prevents interference with fluorescence and SERS signals. |
| Computer-controlled Micromanipulator & Inverted Microscope | Allows precise, coordinate-driven navigation and penetration of the SERS-nanopipette into target cells identified by HCS. |
| Raman Spectrometer (785 nm or 633 nm laser) | Integrated with microscope to acquire SERS spectra from the nanopipette tip located inside a single cell. |
| HCS Image Acquisition & Analysis Software (e.g., MetaXpress, Harmony) | Automates image capture, cell segmentation, and feature extraction from multi-well plates, generating target cell lists. |
Conclusion
SERS-active nanopipettes emerge as a uniquely powerful and versatile platform for spatially resolved, label-free detection of hemeproteins within living single cells. By synthesizing foundational knowledge, robust methodology, troubleshooting insights, and rigorous validation, this technique addresses a critical gap in single-cell proteomics, particularly for redox-active and metabolic proteins. Future directions include multiplexing capabilities for simultaneous protein detection, integration with transcriptomic reads, and adaptation for in vivo tissue analysis. For biomedical research, this promises deeper insights into cellular heterogeneity in disease states like cancer and neurodegeneration, paving the way for novel biomarker discovery and personalized therapeutic strategies grounded in single-cell functional phenotyping.