Harnessing the FcRn Receptor: A Comprehensive Guide to Albumin Half-Life Extension for Therapeutic Proteins
This article provides a detailed technical overview of the neonatal Fc receptor (FcRn) for drug development professionals, covering its fundamental biology in IgG and albumin recycling, methodologies for engineering half-life...
Harnessing the FcRn Receptor: A Comprehensive Guide to Albumin Half-Life Extension for Therapeutic Proteins
Abstract
This article provides a detailed technical overview of the neonatal Fc receptor (FcRn) for drug development professionals, covering its fundamental biology in IgG and albumin recycling, methodologies for engineering half-life extended therapeutics, common optimization challenges, and comparative analysis of emerging technologies. It synthesizes current research and practical applications to guide the design of next-generation biologics with enhanced pharmacokinetics.
The FcRn Recycling Pathway: Foundational Biology for Half-Life Extension
Within the broader context of extending the therapeutic half-life of albumin and albumin-fusion biologics, a comprehensive understanding of the Neonatal Fc Receptor (FcRn) is fundamental. FcRn is the central regulator of albumin and IgG homeostasis, and its targeted manipulation is a cornerstone of modern half-life extension strategies.
Structure and Binding Mechanism
FcRn is a heterodimeric receptor composed of a transmembrane MHC class I-like α-chain and a soluble β2-microglobulin (β2m) light chain. The binding site for both IgG and albumin is formed at the interface of these two subunits. Crucially, binding is strictly pH-dependent, occurring with high affinity in the acidic environment (pH ~6.0) of endosomes, but not at neutral or physiological pH (pH 7.4) of the blood.
Table 1: Key Quantitative Parameters of Human FcRn
| Parameter | Value / Description | Notes |
|---|---|---|
| Gene | FCGRT (α-chain) | Chromosome 19q13.33 |
| Protein Complex | FcRn (α-chain) + β2-microglobulin | Non-covalent association |
| Molecular Weight | ~50 kDa (α-chain) + ~12 kDa (β2m) | Total ~62 kDa |
| IgG Binding Affinity (pH 6.0) | K_D ~ 10 nM - 200 nM | Varies by IgG subclass/species |
| Albumin Binding Affinity (pH 6.0) | K_D ~ 300 nM - 1 µM (human) | Species-specific |
| Binding pH Optimum | ≤ 6.0 | Dissociates at pH ≥ 7.0 |
Expression and Cellular Localization
FcRn is widely expressed in various tissues, fulfilling both protective (neonatal transport) and homeostatic (salvage) functions. Its cellular localization is primarily endosomal.
Table 2: FcRn Expression Profile Across Key Tissues/Cells
| Tissue / Cell Type | Primary Role | Localization |
|---|---|---|
| Vascular Endothelium | IgG/Albumin salvage | Acidic endosomes, sorting vesicles |
| Hepatocytes | Albumin homeostasis | Early/sorting endosomes |
| Intestinal Epithelium | Neonatal uptake; adult luminal sampling | Apical endocytic compartments |
| Proximal Tubule (Kidney) | Salvage from glomerular filtrate | Apical endosomes |
| Myeloid Cells (Macrophages) | Antigen presentation via IgG immune complexes | Phagosomes, endosomes |
| Placental Syncytiotrophoblasts | Maternal-to-fetal IgG transfer | Endosomes, vesicular networks |
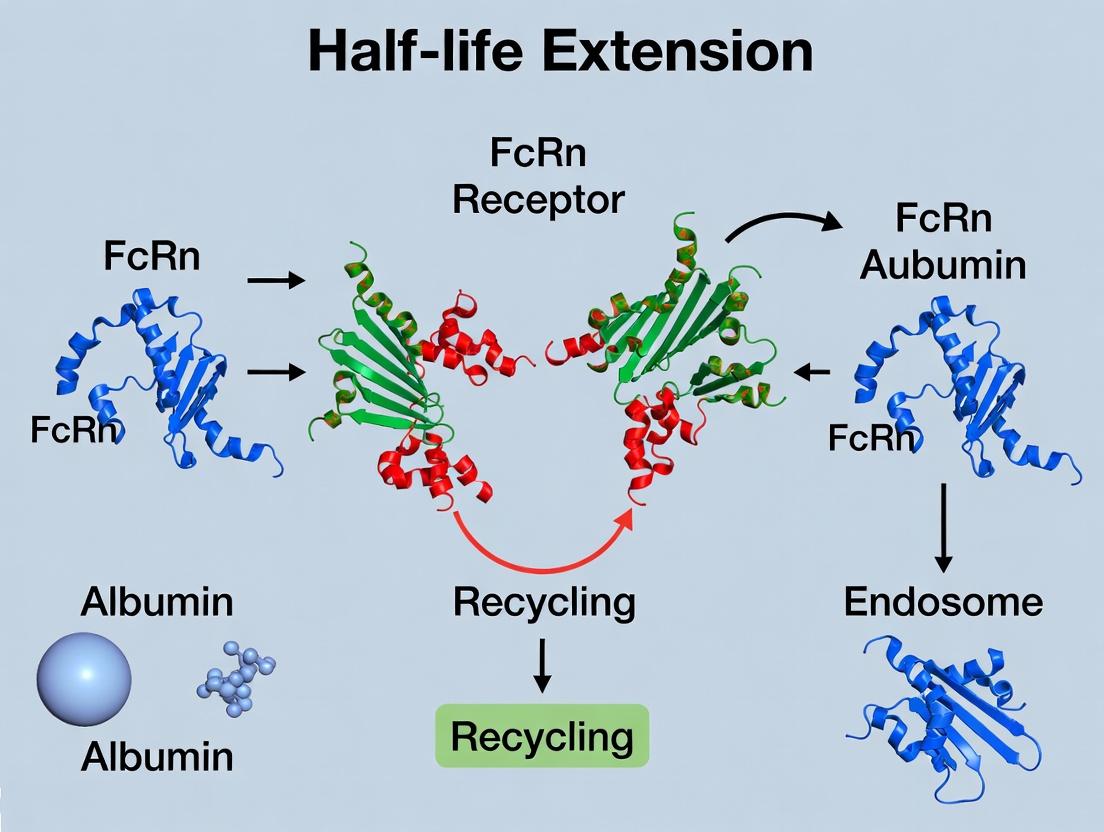
The Salvage Pathway: Visualizing FcRn Function
The core function of FcRn in extending half-life is visualized in the cellular salvage pathway.
Diagram Title: FcRn-Mediated Cellular Salvage of IgG and Albumin
Key Experimental Protocols in FcRn Research
Protocol 1: Surface Plasmon Resonance (SPR) for pH-Dependent Binding Kinetics
- Objective: Determine the binding affinity (KD) and kinetics (kon, k_off) of FcRn-IgG/FcRn-albumin interactions at different pH values.
- Methodology:
- Immobilize recombinant human FcRn onto a CMS sensor chip via amine coupling.
- Use HBS-EP (10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% P20) as running buffer. For acidic binding, use pH 5.5-6.0 buffer; for dissociation, switch to pH 7.4 buffer.
- Inject a concentration series of IgG or albumin analyte over the FcRn surface at pH 6.0 (association phase, 120-180 sec).
- Switch flow to pH 7.4 buffer to monitor dissociation (120-300 sec).
- Regenerate the surface with a short pulse of glycine pH 2.0.
- Analyze data using a 1:1 Langmuir binding model to calculate kinetics.
Protocol 2: Cellular Transcytosis/Recycling Assay using Polarized Cells
- Objective: Quantify FcRn-mediated transport of IgG or albumin across a polarized epithelial monolayer (e.g., MDCK-II or human endothelial cells stably expressing FcRn).
- Methodology:
- Culture cells on Transwell filters until tight junctions form (monitor TEER).
- Radiolabel (I-125) or fluorescently label (e.g., Alexa Fluor) IgG/albumin.
- Add ligand to the apical chamber in pre-warmed, serum-free medium at pH 6.0.
- Incubate at 37°C. Sample from the basolateral chamber (for transcytosis) or re-collect from the apical chamber (for recycling) over a time course (e.g., 30, 60, 120 min).
- Quantify transported/recycled ligand using a gamma counter or fluorometer.
- Include controls: excess unlabeled ligand (competition), incubations at 4°C, or use of FcRn-knockdown cells.
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Research Reagents for FcRn Studies
| Reagent / Material | Function / Application | Example/Notes |
|---|---|---|
| Recombinant Human FcRn (α-chain + β2m) | In vitro binding studies (SPR, ELISA), crystallization. | Produced in HEK293 or CHO cells for proper folding/glycosylation. |
| pH-Sensitive Binding Buffers | Mimic endosomal (pH 5.5-6.2) and physiological (pH 7.4) conditions. | MES (pH 5.5-6.5) and HEPES (pH 7.0-7.5) buffers are standard. |
| FcRn-Specific Monoclonal Antibodies | Blocking function, detecting receptor in WB/IHC/Flow. | e.g., clone 1H5 (blocking), ADM31 (non-blocking, detecting). |
| FcRn-Knockout Mouse Model | In vivo validation of FcRn-dependent pharmacokinetics. | Fcgrt tm1Dcr strain; shows rapid clearance of IgG/albumin. |
| Polarized Epithelial Cell Lines | Study of transcytosis and recycling pathways. | MDCK-II, Caco-2, or engineered human endothelial cell lines. |
| Tracer IgG/Albumin | Labeled ligands for tracking cellular processing. | Human IgG1, HSA; labels: I-125, Alexa Fluor 488/647, HiLyte Fluor. |
| Protease Inhibitor Cocktail (Acidic) | Preserve ligands/FcRn during endosomal fractionation studies. | Essential for assays involving lysosomal inhibition (e.g., leupeptin). |
The precise manipulation of the FcRn-albumin interaction, informed by its structural details and cellular trafficking logic, remains a primary avenue for achieving superior pharmacokinetic profiles of next-generation biologic therapeutics.
This whitepaper provides an in-depth technical guide to the neonatal Fc receptor (FcRn) mediated salvage pathway, a critical physiological mechanism that extends the half-lives of immunoglobulin G (IgG) and albumin. Within the context of a broader thesis on FcRn receptor-based half-life extension research, understanding this endogenous recycling process is foundational for developing novel therapeutics with optimized pharmacokinetic profiles. FcRn functions as a cellular protector, rescuing these two most abundant plasma proteins from inevitable lysosomal degradation, thereby maintaining homeostatic concentrations and modulating immune function.
The FcRn Receptor: Structure and Function
FcRn is a heterodimeric receptor composed of a non-covalently associated α-chain (homologous to major histocompatibility complex class I molecules) and a β2-microglobulin light chain. Its binding to ligands is strictly pH-dependent: high-affinity binding occurs at the acidic pH (~6.0-6.5) of endosomes, while release occurs at neutral pH (~7.4) of the bloodstream or interstitial fluid. This elegant pH switch is the cornerstone of the recycling mechanism.
Detailed Salvage Pathway for IgG and Albumin
The recycling pathway is a coordinated intracellular journey.
Step 1: Cellular Uptake. IgG and albumin are taken into vascular endothelial cells and other cell types (e.g., hematopoietic, epithelial) via fluid-phase pinocytosis or, in some cases, non-specific endocytosis.
Step 2: Endosomal Sorting. The pinocytotic vesicles mature into early endosomes, which acidify via vacuolar ATPase pumps. The acidic environment induces conformational changes in both FcRn and its ligands, promoting high-affinity binding.
Step 3: Diverting from Degradation. Cargo-bound FcRn is actively sorted away from the default lysosomal degradation pathway. This sorting involves specific interactions with the cellular trafficking machinery, directing the FcRn-ligand complex into recycling tubules and vesicles.
Step 4: Return to Circulation. These recycling vesicles transit to the cell periphery, where the neutral extracellular pH triggers the release of IgG and albumin back into the bloodstream or interstitial space. The unoccupied FcRn is then free to re-enter the cycle.
Step 5: Degradation of Unbound Protein. Any IgG or albumin that does not bind to FcRn in the acidic endosome proceeds to late endosomes and ultimately to lysosomes, where they are catabolized into amino acids.
The following diagram illustrates this core salvage pathway.
Diagram 1: The FcRn-Mediated Salvage Pathway
Quantitative Data: Impact on Half-Life
The FcRn-mediated rescue profoundly impacts the serum half-life of IgG and albumin compared to proteins without this mechanism. The table below summarizes key comparative data.
Table 1: Impact of FcRn on Protein Half-Lives
| Protein | Typical Half-Life (Human) | Half-Life in FcRn Knockout/Inhibition | Key Binding Site on Protein |
|---|---|---|---|
| IgG1 | ~21 days | ~2-3 days | CH2-CH3 domain (Fc region), His310, His435 |
| Albumin | ~19 days | ~1-2 days | Domain III, His166, His242, His288 |
| IgG (Fab fragment) | ~0.5 days | Unchanged | N/A (Lacks Fc region) |
| Fc-fusion protein | Varies (days-weeks) | Drastically reduced | Engineered Fc domain |
Key Experimental Protocols for FcRn Research
Understanding and manipulating the FcRn pathway requires robust experimental models. Below are detailed methodologies for key assays.
Protocol 1: In Vitro pH-Dependent Binding Kinetics (Surface Plasmon Resonance - SPR) Objective: Quantify the binding affinity (KD) of IgG/albumin variants to FcRn at acidic vs. neutral pH. Materials: Biacore or equivalent SPR instrument, recombinant human/mouse FcRn, CMS sensor chip, HBS-EP running buffer. Procedure:
- Immobilize anti-His antibody on a CMS chip via amine coupling.
- Capture His-tagged FcRn onto the sensor chip surface via the anti-His antibody.
- Inject serial dilutions of the IgG or albumin analyte in acidic buffer (pH 5.5-6.0) over the flow cells.
- Monitor the association phase for 180 seconds.
- Switch to neutral buffer (pH 7.4) to initiate dissociation, monitoring for 300-600 seconds.
- Regenerate the surface with a mild acidic buffer (pH 4.5).
- Repeat with neutral pH running buffer for the entire cycle as a control.
- Analyze data using a 1:1 Langmuir binding model to calculate association (ka), dissociation (kd) rates, and equilibrium KD.
Protocol 2: Cellular Recycling Assay Using Human Endothelial Cells (e.g., HMEC-1) Objective: Directly measure the FcRn-mediated rescue of test IgG from degradation. Materials: HMEC-1 cells, serum-free medium, acidic wash buffer (pH 5.0), neutral wash buffer (pH 7.4), fluorescently-labeled or radioiodinated ([125]I) test IgG, cycloheximide (protein synthesis inhibitor). Procedure:
- Culture HMEC-1 cells to confluence in 24-well plates.
- Pre-block cells with excess unlabeled human IgG to saturate non-specific uptake.
- Incubate cells with a known concentration of labeled test IgG in serum-free, pH 7.4 medium at 37°C for 2h (pulse phase).
- Wash cells extensively with acidic buffer (pH 5.0) to remove surface-bound IgG.
- Add fresh pre-warmed medium containing cycloheximide and an excess of unlabeled IgG (to prevent re-capture of released labeled IgG).
- Incubate at 37°C for varying times (0-24h) to allow for recycling (chase phase).
- At each time point, collect the supernatant and lyse the cells.
- Measure radioactivity or fluorescence in the supernatant (recycled/catabolized) and cell lysate (retained) fractions using a gamma counter or plate reader.
- Calculate the percentage of internalized IgG that is released intact over time. Compare FcRn-binding variants to controls.
Protocol 3: In Vivo Pharmacokinetic Study in Humanized FcRn Transgenic or FcRn KO Mice Objective: Determine the serum half-life of an IgG/albumin variant in a whole-animal model with a humanized FcRn system or lacking FcRn. Materials: Human FcRn transgenic mice (e.g., B6.mFcRn-/-.hFcRn Tg32), wild-type and FcRn KO controls, test protein (IV formulation), EDTA-coated microtainers for blood collection. Procedure:
- Administer a single intravenous bolus of the test protein (e.g., 5 mg/kg) to groups of mice (n=5-8).
- Collect serial blood samples (e.g., 10 µL from tail vein) at predetermined time points (e.g., 5 min, 2h, 8h, 1, 2, 4, 7, 10, 14, 21 days post-dose).
- Process plasma by centrifugation.
- Quantify test protein concentration in each sample using a specific ELISA (e.g., anti-human Fc or anti-albumin capture).
- Perform non-compartmental pharmacokinetic analysis (using software like Phoenix WinNonlin) on mean concentration-time data to determine key parameters: Terminal half-life (t1/2), clearance (CL), and area under the curve (AUC).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for FcRn/Albin Half-Life Research
| Reagent/Category | Example Product/Source | Function in Research |
|---|---|---|
| Recombinant FcRn Protein | Sino Biological, R&D Systems | In vitro binding studies (SPR, ELISA), crystallization, screening assays. |
| pH-Sensitive Binding ELISA Kits | Theraclone Sciences, in-house formats | High-throughput screening of IgG/albumin variants for pH-dependent FcRn binding. |
| FcRn-Expressing Cell Lines | HMEC-1 (endothelial), MDCK-hFcRn | Cellular recycling and transcytosis assays; model systems for in vitro PK. |
| Humanized FcRn Mouse Models | Tg32, Tg276 (The Jackson Laboratory) | In vivo pharmacokinetic and efficacy studies in a physiologically relevant model. |
| FcRn Inhibitors/Blockers | Monoclonal anti-FcRn (e.g., Rozanolixizumab), Peptide 'Abdeg' | Tools to acutely inhibit FcRn in vivo, validating mechanism or modeling KO phenotype. |
| Site-Directed Mutagenesis Kits | Agilent QuikChange, NEB Q5 | Engineering point mutations in Fc/albumin to modulate FcRn binding affinity. |
| Long-Acting (Half-Life Extended) Reference mAbs | Trastuzumab, Palivizumab (wild-type) vs. engineered variants (e.g., YTE) | Critical benchmarks for comparing the performance of novel half-life extension technologies. |
Application in Drug Development: Half-Life Extension Technologies
The mechanistic understanding of FcRn recycling has been directly translated into biotherapeutic engineering. The primary strategies are:
- Fc Engineering: Introducing point mutations (e.g., M252Y/S254T/T256E - "YTE"; M428L/N434S - "LS") into the Fc region of monoclonal antibodies to increase binding affinity to FcRn at acidic pH, while maintaining proper pH-sensitive release.
- Albumin Fusion: Genetically fusing the therapeutic protein (e.g., cytokines, enzymes) to human albumin, thereby hijacking its natural FcRn-mediated recycling pathway.
- Albumin Binding Domains: Engineering small proteins, peptides, or antibody fragments (e.g., AlbuTag) with high affinity for albumin, creating a non-covalent complex that follows the albumin recycling route.
The competitive landscape of these approaches can be visualized through their developmental logic.
Diagram 2: Drug Development Strategies via FcRn
The FcRn salvage pathway is a master regulator of IgG and albumin homeostasis. Its detailed mechanistic understanding, from cellular trafficking to pH-sensitive kinetics, provides a powerful blueprint for modern drug design. Research in this field, as framed by this thesis, continues to evolve with more precise structural insights, novel cellular models, and sophisticated engineered variants. The experimental protocols and tools outlined here form the cornerstone for advancing this knowledge, driving the development of next-generation biologics with optimized half-lives, reduced dosing frequency, and improved patient outcomes.
Within the broader thesis of leveraging the neonatal Fc receptor (FcRn) for albumin half-life extension research, this whitepaper provides an in-depth analysis of the core mechanism: pH-dependent binding. FcRn's unique ability to bind IgG and albumin at acidic pH (endosome) and release at neutral pH (bloodstream) is fundamental to its role in endosomal sorting, recycling, and rescue from lysosomal degradation. This process is the cornerstone for developing long-lasting biotherapeutics. This guide details the structural biology, kinetics, experimental methodologies, and research tools essential for investigators in this field.
The neonatal Fc receptor is a major histocompatibility complex (MHC) class I-related heterodimer, composed of a heavy chain (α-chain) and beta-2-microglobulin (β2m). Its primary physiological function is to regulate the serum half-lives of IgG and albumin, two of the most abundant plasma proteins. This is achieved via a sophisticated intracellular salvage pathway. Disrupting FcRn-IgG interaction can treat autoimmune diseases, while engineering therapeutics to enhance FcRn affinity can dramatically extend their circulating half-life. This principle is central to albumin-fusion and Fc-engineered drug development.
Structural & Kinetic Basis of pH-Dependent Binding
Molecular Interactions at Acidic vs. Neutral pH
Binding is governed by protonation states of key histidine residues.
- For IgG: At pH ~6.0 (endosomal lumen), histidines (e.g., H310, H435 on IgG Fc) become protonated, enabling ionic bonds with negatively charged glutamates/aspartates (e.g., E115, E116) on FcRn. At pH ~7.4 (bloodstream), these histidines deprotonate, disrupting ionic interactions and leading to dissociation.
- For Albumin: The mechanism is analogous but involves distinct residues. At acidic pH, histidine residues in albumin's domain III (H464, H510) protonate, enabling binding to a similar pocket on FcRn. Neutral pH triggers release.
Quantitative Binding Kinetics
The affinity constants underscore the stark pH dependency.
Table 1: Representative FcRn Binding Affinities (K_D)
| Ligand | pH | K_D (nM) | Method | Reference (Example) |
|---|---|---|---|---|
| Human IgG1 (Wild-type) | 6.0 | 50 - 2000 | Surface Plasmon Resonance (SPR) | (Chaudhury et al., 2003) |
| Human IgG1 (Wild-type) | 7.4 | >10,000 (weak/no bind) | SPR | (Chaudhury et al., 2003) |
| Engineered IgG (YTE mutant) | 6.0 | ~10-50 | SPR | (Robbie et al., 2013) |
| Human Serum Albumin (HSA) | 5.5-6.0 | 100 - 500 | Biolayer Interferometry (BLI) | (Andersen et al., 2012) |
| HSA | 7.4 | >>1000 | BLI | (Andersen et al., 2012) |
The Endosomal Sorting & Recycling Pathway: A Visual Guide
Title: FcRn-Mediated Endosomal Recycling Pathway
Key Experimental Protocols
Measuring pH-Dependent Binding Kinetics by Surface Plasmon Resonance (SPR)
Objective: Determine the association (kon) and dissociation (koff) rates and equilibrium dissociation constant (K_D) of FcRn-ligand interaction at different pH values. Detailed Protocol:
- Immobilization: FcRn (purified extracellular domain) is immobilized on a CMS sensor chip using standard amine coupling to achieve ~1000-2000 Response Units (RU).
- Running Buffer Preparation: Prepare HBS-EP buffers adjusted to pH 5.5, 6.0, 6.5, and 7.4. Filter and degas.
- Ligand Injection: Serial dilutions of IgG or albumin analyte are prepared in the corresponding pH buffer. Inject over the FcRn and reference flow cells for 180s (association phase) at a flow rate of 30 µL/min.
- Dissociation Phase: Switch flow to pH 7.4 running buffer for 300-600s to monitor dissociation. Note: For true physiological mimicry, a two-step dissociation can be used: first in the analyte pH buffer, then switch to pH 7.4 buffer.
- Regeneration: The chip surface is regenerated with a short pulse (30s) of pH 7.4 buffer or mild basic solution (e.g., 10mM NaOH).
- Data Analysis: Double-reference subtraction (reference cell & buffer blank). Kinetic parameters are calculated by globally fitting the sensograms to a 1:1 Langmuir binding model using the instrument's software (e.g., Biacore Evaluation Software).
Cellular Recycling Assay (In Vitro)
Objective: Quantify the FcRn-mediated recycling efficiency of a ligand in a cellular system (e.g., human endothelial or epithelial cells). Detailed Protocol:
- Cell Culture: Seed FcRn-expressing cells (e.g., human microvascular endothelial cells (HMEC-1) stably transfected with hFcRn) into 24-well plates.
- Loading/Pulse: Wash cells and incubate with a known concentration of test IgG/albumin (radiolabeled or fluorescently tagged) in acidic, serum-free medium (pH 6.0, 37°C) for 1-2 hours to allow binding and uptake.
- Acid Wash: Remove unbound ligand by washing cells with cold acidic buffer (pH 3.0-4.0) to strip surface-bound (but not internalized) ligand.
- Chase/Recycling: Add pre-warmed, ligand-free neutral medium (pH 7.4, 37°C) to initiate recycling. Collect the supernatant at defined time points (e.g., 15, 30, 60, 120 min).
- Lysate Collection: At the final time point, lyse cells to quantify ligand remaining intracellularly.
- Quantification: Measure ligand concentration in supernatants and lysates via gamma counting (for I-125 label) or fluorescence. Calculate the percentage of internalized ligand released into the neutral medium over time.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for FcRn Binding & Recycling Studies
| Item | Function / Application | Example Product / Note |
|---|---|---|
| Recombinant Human FcRn (α-chain + β2m) | The core reagent for in vitro binding studies (SPR, BLI). Must be heterodimeric and properly folded. | Sino Biological, AcroBiosystems, homemade from HEK293 cells. |
| Control IgGs (Wild-type & Mutants) | Positive (high pH-dependent bind, e.g., IgG1) and negative (low bind, e.g., IgG4, Fc mutants) controls for assays. | Commercial purified IgG, or expressed from vectors. |
| pH-Varied Buffer Systems | Critical for mimicking endosomal and plasma conditions. Must have matched ionic strength. | MES (pH 5.5-6.5), HEPES/Phosphate (pH 7.0-7.4). |
| FcRn-Expressing Cell Lines | For cellular recycling and transcytosis assays. | HMEC-1/hFcRn, MDCK-II/hFcRn, HEK293/hFcRn. |
| Anti-FcRn Antibodies (Blocking) | To confirm FcRn-specificity in cellular assays by inhibiting function. | Clone 1G3, 4C9, or ADM31. |
| Biosensor Chips & Systems | For label-free kinetic analysis. | Cytiva Biacore CMS chips, FortéBio Octet Streptavidin (SA) or Anti-Human Fc (AHQ) biosensors. |
| Labeled Ligands | For tracking in cellular assays. | Iodine-125 (I-125) for radioassays, Alexa Fluor 488/647 for fluorescence-based assays. |
| Protease Inhibitors (e.g., Leupeptin) | Used in cellular assays to differentiate recycling from degradation (inhibits lysosomal proteases). | Add to chase medium to block degradation, increasing recycled fraction. |
Advanced Research & Therapeutic Engineering
Understanding precise pH-switch mechanisms enables rational drug design. Strategies include:
- Fc Engineering: Introducing mutations (e.g., M252Y/S254T/T256E "YTE", M428L/N434S "LS") that increase affinity at pH 6.0 without compromising the off-rate at pH 7.4.
- Albumin Engineering & Fusion: Mutating albumin (e.g., K573P) or creating genetic fusions to hijack the FcRn recycling pathway.
- FcRn Inhibitors: Developing high-affinity antibodies (e.g., efgartigimod) or peptides that block FcRn-IgG interaction at all pH values, promoting IgG degradation for autoimmune disease treatment.
Title: Therapeutic Strategies Targeting the FcRn pH-Switch
Within the therapeutic landscape dominated by monoclonal antibodies, Fc-fusion proteins, and emerging albumin-conjugated biologics, precise modulation of IgG/albumin homeostasis is paramount. A central thesis in half-life extension research posits that selectively engaging the neonatal Fc receptor (FcRn) for recycling, while avoiding engagement of classical Fc gamma receptors (FcγRs) responsible for effector functions, is key to optimizing pharmacokinetics (PK) and pharmacodynamics (PD). This whitepaper delineates the structural, functional, and mechanistic distinctions between these receptor families, providing a technical guide for their targeted manipulation in drug development.
Structural and Functional Dichotomy
The fundamental divergence lies in their cellular distribution, binding sites on IgG, pH dependency, and ultimate cellular fate.
| Feature | FcRn | FcγRs (I, II, III subfamilies) |
|---|---|---|
| Primary Role | Homeostasis & Transcytosis | Immune Effector Functions |
| Binding Site on IgG | CH2-CH3 domain interface | Lower hinge/CH2 region (overlapping but distinct) |
| Critical Binding pH | Acidic (≤6.5) for high affinity; Neutral (7.4) for release | Neutral/Alkaline (7.0-7.4) for high affinity |
| Cellular Expression | Vascular endothelium, monocytes, hepatocytes, epithelia | Myeloid cells (macrophages, DCs, neutrophils), NK cells, B cells |
| Key Signaling Motifs | None (no intracellular signaling domains) | ITAM (FcγRI, IIA, IIIA) or ITIM (FcγRIIB) |
| Ligand Engagement Fate | Recycling/Transcytosis (protective) | Phagocytosis, ADCC, ADCP, Degradation, Cell Activation |
Quantitative Binding and Functional Data
Recent studies using Surface Plasmon Resonance (SPR) and cellular assays quantify these distinctions.
Table 1: Comparative Binding Affinities (SPR Data)
| Receptor | Affinity for Human IgG1 (KD) | pH of Measurement | Notes |
|---|---|---|---|
| hFcRn | ~50-400 nM | pH 6.0 | High affinity; negligible binding at pH 7.4. |
| hFcγRI (CD64) | ~10^-10 M | pH 7.4 | High affinity, monomeric binding. |
| hFcγRIIa (H131) | ~3 x 10^-7 M | pH 7.4 | Low affinity, aggregate-induced binding. |
| hFcγRIIIa (V158) | ~5 x 10^-7 M | pH 7.4 | Low affinity, critical for ADCC. |
Table 2: Cellular Outcome Metrics
| Assay Type | FcRn Engagement Outcome | FcγR (ITAM) Engagement Outcome |
|---|---|---|
| IgG Half-life (in vivo) | Extended from days to ~21 days (human) | Reduced via clearance of immune complexes. |
| Antigen Presentation | Enhanced cross-presentation (pH-dependent). | Robust presentation via phagocytic/endocytic pathway. |
| Cytokine Release | Minimal to none. | Significant (e.g., TNF-α, IL-6, IFN-γ). |
| ADCC (in vitro % lysis) | 0-5% (background). | 20-60% (with engineered high-affinity variants). |
Detailed Experimental Protocols
Protocol: pH-Dependent FcRn Binding ELISA
Objective: Quantify IgG/Fc-fusion protein binding to FcRn at acidic vs. neutral pH.
- Coating: Immobilize recombinant hFcRn (5 µg/mL in PBS) on a high-binding ELISA plate overnight at 4°C.
- Blocking: Block with PBS containing 3% BSA and 0.05% Tween-20 for 2 hours.
- IgG Binding (Acidic): Add IgG serial dilutions in MES buffer (pH 6.0). Incubate 2 hours. Include wild-type IgG and Fc-mutated variants (e.g., M428L/N434S).
- Wash: Wash plates with pH 6.0 buffer.
- Detection (Neutral Release): Add HRP-conjugated anti-human Fc antibody in PBS (pH 7.4). Incubate 1 hour. The pH shift minimizes detection of non-specifically retained IgG.
- Development & Readout: Develop with TMB substrate, stop with H₂SO₄, read absorbance at 450nm.
Protocol: FcγRIIIa (CD16a) Reporter Bioassay for ADCC Potential
Objective: Measure cell-based signaling activation via FcγRIIIa.
- Cell Preparation: Use engineered Jurkat T-cells stably expressing human FcγRIIIa (V158 variant) and an NFAT-response element driving luciferase.
- Target Cell Seeding: Seed target cells expressing the antigen of interest in a white-walled 96-well plate.
- Antibody Addition: Add serial dilutions of the test therapeutic antibody.
- Effector Cell Addition: Add FcγRIIIa reporter cells at an effector:target ratio of 10:1.
- Incubation: Incubate for 6 hours at 37°C, 5% CO₂.
- Signal Detection: Add Bio-Glo Luciferase Assay Reagent, measure luminescence. Data is reported as Relative Light Units (RLU) vs. antibody concentration.
Pathway and Workflow Visualizations
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Supplier Examples | Function in FcRn/FcγR Research |
|---|---|---|
| Recombinant hFcRn (α-chain + β2m) | Sino Biological, AcroBiosystems | Essential for in vitro binding studies (SPR, ELISA) to measure pH-dependent affinity. |
| FcγR Tetramers (FcγRI, IIa/b, IIIa) | Thermo Fisher, BioLegend | Detect and quantify IgG binding to specific FcγRs via flow cytometry, minimizing avidity issues of monomeric binding. |
| ADCC Reporter Bioassay Kits (FcγRIIIa) | Promega | Standardized, cell-based system to measure antibody-induced FcγR signaling and effector function potential. |
| hFcRn Transgenic Mouse Models | The Jackson Laboratory, genOway | In vivo model for human IgG PK studies, predicting human half-life extension of engineered variants. |
| SPR/Biacore Sensor Chips (CM5, SA) | Cytiva | Immobilize FcRn or FcγRs for real-time, label-free kinetic analysis of antibody binding interactions. |
| Human IgG Fc Variant Libraries | Custom synthesis (e.g., Genscript) | Collections of Fc mutants (e.g., Xtend, YTE, LS) for high-throughput screening of binding properties. |
| pH-Sensitive Fluorophore Dyes (e.g., pHrodo) | Thermo Fisher | Label antigens or immune complexes to track phagocytosis (ADCP) and lysosomal acidification in real time. |
The neonatal Fc receptor (FcRn) is a critical regulator of immunoglobulin G (IgG) and albumin homeostasis, governing their extended serum half-lives via a pH-dependent recycling mechanism. Research into leveraging FcRn for half-life extension of biologics and albumin-fusion therapeutics forms the cornerstone of a broader thesis on next-generation protein engineering. A central, often underappreciated, challenge in this field is the profound species differences in FcRn biology, which can confound the interpretation of preclinical pharmacokinetic (PK) studies and derail translation to human clinical outcomes.
Comparative Biology of FcRn Across Species
FcRn function is conserved, but its specific expression patterns, binding kinetics, and cellular handling vary significantly. These differences necessitate careful species selection for in vitro and in vivo studies.
Table 1: Key Species-Specific Characteristics of FcRn Biology
| Characteristic | Human | Cynomolgus Monkey | Mouse | Rat | Transgenic Mouse (e.g., hFcRn) |
|---|---|---|---|---|---|
| Gene Symbol | FCGRT | FCGRT | Fcgrt | Fcgrt | Fcgrt (KO); human FCGRT (TG) |
| β2m Dependence | Yes | Yes | Yes | Yes | Yes (with human β2m) |
| Tissue Expression | Broad: endothelium, epithelium, myeloid cells | Similar to human | High in endothelium, liver, placenta | High in yolk sac, kidney, intestine | Pattern depends on promoter (often ubiquitous) |
| pH-Dependent Binding | Strong binding at pH ≤6.5; rapid release at pH 7.4 | Similar to human | Strong binding at pH ≤6.0; release at pH 7.0-7.4 | Similar to mouse | Mirrors human pH profile |
| Affinity for hIgG (KD at pH 6.0) | ~300-800 nM | ~200-600 nM | Very weak (>10 µM) | Very weak | ~300-800 nM (for hIgG) |
| Primary PK Study Utility | Target (clinical) | Gold-standard non-human primate model | Limited for human IgG PK; good for murine IgG | Limited for human IgG PK | Preferred rodent model for human IgG PK |
Experimental Protocols for Characterizing FcRn Interactions
Protocol 1: Surface Plasmon Resonance (SPR) for Binding Kinetics Objective: Determine species-specific affinity (KD) and kinetics (ka, kd) of IgG/albumin variants for recombinant FcRn at endosomal pH. Methodology:
- Immobilization: Capture or directly immobilize recombinant FcRn (e.g., human, mouse, cyno) onto a CMS sensor chip via amine coupling.
- Running Buffer: Use a buffer simulating endosomal conditions (e.g., 100 mM sodium phosphate, 150 mM NaCl, pH 5.5-6.0).
- Analyte Injection: Inject a concentration series (e.g., 0-1000 nM) of the IgG or albumin variant over the FcRn surface at a flow rate of 30 µL/min.
- Dissociation & Regeneration: Allow dissociation in running buffer, then regenerate the surface with a pulse of HBS-EP+ buffer at pH 7.4.
- Data Analysis: Fit the resulting sensograms to a 1:1 Langmuir binding model using BIAevaluation or similar software to calculate ka, kd, and KD.
Protocol 2: In Vivo PK Study in Humanized FcRn Transgenic Mouse Model Objective: Evaluate the serum half-life of a human IgG or albumin-fusion therapeutic candidate. Methodology:
- Animal Model: Use homozygous B6.Cg-Fcgrt*tm1Dcr Tg(FCGRT)32Dcr/DcrJ mice (or equivalent), which lack mouse FcRn and express human FcRn.
- Dosing: Administer a single intravenous bolus of the test article (e.g., 5 mg/kg) via the tail vein (n=5-6 per group).
- Serial Bleeding: Collect blood samples (≤10% total volume) at predefined time points (e.g., 0.083, 1, 6, 24, 72, 168, 240, 336 hours post-dose).
- Sample Analysis: Process serum and quantify test article concentration using a validated ligand-binding assay (e.g., ELISA).
- PK Analysis: Fit serum concentration-time data using non-compartmental analysis (NCA) to estimate key parameters: terminal half-life (t1/2), clearance (CL), and volume of distribution (Vd).
Visualizations
Title: FcRn-Mediated Recycling Pathway for IgG and Albumin
Title: Translational PK Workflow Accounting for Species Differences
The Scientist's Toolkit: Key Research Reagents & Materials
Table 2: Essential Reagents for FcRn-Focused Preclinical Research
| Reagent/Material | Function & Application | Key Considerations |
|---|---|---|
| Recombinant FcRn Proteins (Human, Cyno, Mouse) | In vitro binding assays (SPR, ELISA). Critical for screening variant libraries. | Must be complexed with β2-microglobulin. Verify correct pH-dependent binding profile. |
| hFcRn Transgenic Mice (e.g., B6.Cg-Fcgrttm1Dcr Tg32Dcr) | Primary in vivo model for predicting human IgG/albumin PK. | Confirm genotype. Backcrossing to desired background strain may be required. |
| Cynomolgus Monkeys (Macaca fascicularis) | Non-human primate PK and toxicology studies. Most predictive non-human model. | High cost and ethical considerations. Use of purpose-bred animals is mandatory. |
| pH-Sensitive Cell-Based Assays (e.g., using BeWo, HUVEC, or engineered cells) | Functional assessment of FcRn-mediated recycling and transcytosis. | Requires precise control of buffer pH during pulse and chase phases. |
| Anti-FcRn Blocking Antibodies (e.g., anti-hFcRn mAbs) | In vivo validation of FcRn-specific mechanism; tool to modulate PK. | Select species-specific antibodies. Can be used to saturate FcRn and accelerate clearance. |
| Biotinylated IgG/Albumin Ligands | Tracer molecules for cell surface binding assays or histochemistry. | Ensure biotinylation does not alter FcRn binding characteristics. |
Engineering Strategies for FcRn-Mediated Half-Life Extension: From Design to Delivery
The neonatal Fc receptor (FcRn) is central to the long half-life of IgG antibodies and serum albumin, protecting them from lysosomal degradation via a pH-dependent recycling mechanism. Within the broader thesis of FcRn-mediated half-life extension research, the engineering of Fc fusion proteins represents a pivotal strategy to improve the pharmacokinetics (PK) of therapeutic biologics. This whitepaper provides an in-depth technical guide to designing Fc domains with enhanced FcRn affinity at acidic pH (endosomal, ~pH 6.0) while maintaining weak binding at neutral pH (physiological, ~pH 7.4), a prerequisite for extended serum half-life.
Core Principles of Fc-FcRn Interaction
FcRn binds the Fc region of IgG at the CH2-CH3 domain interface. Key interaction residues on Fc include Ile253, His310, His433, Asn434, and Tyr436. Enhancing affinity primarily involves introducing mutations that stabilize the Fc-FcRn complex at pH 6.0 without increasing neutral pH binding, which would block recycling.
Table 1: Key Fc Residues for FcRn Engineering
| Fc Residue (Kabat) | Role in FcRn Binding | Common Mutations for Enhanced Affinity |
|---|---|---|
| Thr250 | Hydrogen bonding | Thr250Glu, Thr250Met |
| Met252 | Hydrophobic interaction | Met252Tyr, Met252Phe |
| Thr256 | Side chain packing | Thr256Asp, Thr256Glu |
| His310 | pH-sensitive interaction | His310Glu (stabilizes acidic interaction) |
| Asn434 | Critical salt bridge | Asn434His, Asn434Trp, Asn434Tyr |
| His433 | Adjacent to Asn434 | His433Lys, His433Phe |
| Ile253 | Hydrophobic contact | Ile253Ala, Ile253Ser |
| Tyr436 | Peripheral contact | Tyr436His |
Experimental Protocols for Evaluating FcRn Binding and Half-Life
Protocol 3.1: Surface Plasmon Resonance (SPR) for pH-Dependent Binding Kinetics
Objective: Measure the binding affinity (KD) of engineered Fc variants to human FcRn at pH 6.0 and pH 7.4. Methodology:
- Immobilization: Recombinant human FcRn is immobilized on a CMS sensor chip via amine coupling.
- Running Buffer: HBS-EP buffer adjusted to pH 6.0 or 7.4.
- Analyte: Purified Fc variant proteins at concentrations ranging from 0 nM to 1000 nM.
- Cycle: Inject analyte for 180s (association), then switch to running buffer for 600s (dissociation) at a flow rate of 30 µL/min.
- Regeneration: Inject pH 7.4 HBS-EP to fully dissociate complex.
- Analysis: Data is fit to a 1:1 Langmuir binding model. The ideal variant shows high-affinity binding at pH 6.0 (KD < 100 nM) and minimal binding at pH 7.4 (KD > 10,000 nM).
Protocol 3.2: In Vivo Pharmacokinetic Study in Humanized FcRn Transgenic Mice
Objective: Determine the serum half-life of lead Fc fusion variants. Methodology:
- Model: Use B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr/DcrJ mice (expressing human FcRn).
- Dosing: Administer a single 5 mg/kg intravenous bolus of each Fc fusion protein (n=5 per group).
- Sampling: Collect retro-orbital blood samples at 5 min, 30 min, 2h, 8h, 24h, 72h, 168h, 240h, and 336h post-dose.
- Analysis: Quantify serum concentrations via Fc-specific ELISA. Pharmacokinetic parameters (terminal half-life, AUC, clearance) are calculated using non-compartmental analysis (WinNonlin).
Table 2: Representative Data for Engineered Fc Variants
| Fc Variant | SPR KD at pH 6.0 (nM) | SPR KD at pH 7.4 (nM) | Murine Terminal t½ (h) | Fold Increase vs WT |
|---|---|---|---|---|
| WT Fc (Control) | 890 | >50,000 | 189 | 1.0 |
| M252Y/S254T/T256E (YTE) | 12 | >50,000 | 411 | 2.2 |
| M428L/N434S (LS) | 24 | >50,000 | 452 | 2.4 |
| N434H | 8 | 42,000 | 398 | 2.1 |
| N434W | 1.5 | 15,000 | 510 | 2.7 |
| N434Y | 4 | 29,000 | 467 | 2.5 |
| Triple Mutant (M252Y/N434Y/Y436H) | 0.9 | 8,000 | 625 | 3.3 |
Note: Data is illustrative, compiled from recent literature.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for FcRn-Fc Research
| Reagent / Material | Provider Examples | Function in Research |
|---|---|---|
| Recombinant Human FcRn (His-tagged) | Sino Biological, R&D Systems | The key ligand for in vitro binding assays (SPR, BLI). |
| Human IgG Fc (Wild-type) | GenScript, Thermo Fisher | Critical negative/positive control in binding and cellular assays. |
| Humanized FcRn Transgenic Mice | The Jackson Laboratory (Strain #014565) | The in vivo gold-standard model for predicting human PK of Fc variants. |
| Anti-Human Fc (HRP conjugate) | Sigma-Aldrich, Abcam | Detection antibody for quantifying Fc fusion concentrations in ELISA and serum samples. |
| ProteOn XPR36 or Biacore 8K SPR System | Bio-Rad, Cytiva | Instrumentation for high-throughput, precise kinetic analysis of Fc-FcRn interactions. |
| pH-Adjusted HBS-EP Buffer Packs | Cytiva, Teknova | Ensures consistent pH conditions (6.0 vs 7.4) for binding assays. |
| HEK293F or ExpiCHO-S Cells | Thermo Fisher | Preferred mammalian expression systems for producing high-quality Fc variant proteins. |
| Protein A/G Affinity Resin | Thermo Fisher, GenScript | Standard purification method for Fc-containing fusion proteins from culture supernatant. |
Advanced Design Strategies and Future Directions
Beyond point mutations, strategies include:
- Heterodimeric Fc Engineering: Creating "one-arm" high-affinity Fc designs to facilitate the recycling of bispecific molecules.
- Allosteric Modulators: Introducing mutations distal to the binding interface that subtly modulate Fc flexibility and pH sensitivity.
- Machine Learning-Guided Design: Utilizing structural and sequence data to predict novel affinity-enhancing combinations beyond known hotspots.
Validation requires an iterative cycle of in silico design → in vitro affinity screening → in vivo PK testing.
Designing optimal Fc domains for enhanced FcRn affinity is a mature but rapidly evolving discipline. The combination of established mutations (e.g., YTE, LS, N434W/Y) with high-throughput screening and in vivo models provides a robust framework for achieving significant half-life extension for Fc fusion therapeutics. Future work will focus on fine-tuning dissociation at neutral pH and applying these principles to novel biologic formats, further advancing the central thesis of FcRn-based pharmacokinetic optimization.
This technical guide explores the strategic exploitation of the neonatal Fc receptor (FcRn)-mediated recycling pathway to extend the plasma half-life of therapeutic proteins. By engineering fusion proteins or chemical conjugates that bind to endogenous albumin, or by creating recombinant albumin-fusion therapeutics, researchers can directly co-opt albumin's natural long half-life (~19 days in humans). This whitepaper details the molecular mechanisms, experimental methodologies, and current data underpinning this critical half-life extension technology, framed within ongoing FcRn-albumin research.
The extended half-life of serum albumin is governed by its pH-dependent binding to the FcRn. Following pinocytosis, albumin binds FcRn in the acidic endosome (pH ~6.0), is rescued from lysosomal degradation, and is recycled back to the cell surface where neutral pH (7.4) triggers its release. This review focuses on two primary strategies to harness this pathway: 1) Genetic fusion to albumin or albumin-binding domains, and 2) Covalent conjugation to albumin or albumin-binding molecules.
Table 1: Comparative Half-Life Extension via Albumin Interaction
| Therapeutic Format | Example Molecule | Species Tested | Approx. Plasma Half-Life (vs. Native) | Primary Mechanism |
|---|---|---|---|---|
| Native Albumin | HSA | Human | ~19 days | FcRn recycling |
| IgG1 | Therapeutic mAb | Human | ~21 days | FcRn recycling |
| GLP-1 Agonist | Liraglutide | Human | ~13 hrs (vs. min for GLP-1) | HSA non-covalent binding & FcRn |
| Albumin Fusion | Albiglutide (GLP-1-HSA) | Human | ~5 days | Direct FcRn engagement |
| Albumin-Binding Domain Fusion | ABD-fused bispecific | Mouse | ~2.5 days (vs. hrs for scFv) | Non-covalent HSA binding |
| PASylation | PAS-IL-1Ra | Rat | ~60 hrs (vs. 1-2 hrs) | Increased hydrodynamic radius |
| Fatty Acid Conjugation | Insulin detemir | Human | ~5-7 hrs (vs. 4-6 min) | Reversible HSA binding |
| Recombinant Fusion | rHSA-IFNα | Human | ~48 hrs (vs. 4-8 hrs for IFNα) | Direct FcRn engagement |
Table 2: Key FcRn Binding Affinity Data (Surface Plasmon Resonance)
| Ligand | pH | KD (µM) | ka (1/Ms) | kd (1/s) | Reference Context |
|---|---|---|---|---|---|
| Human Albumin (HSA) | 6.0 | 0.5 - 1.2 | ~2.0 x 10^5 | ~0.15 | Wild-type binding |
| HSA (H464Q mutant) | 6.0 | >50 | N/D | N/D | Disrupted FcRn binding |
| Mouse Albumin (MSA) | 6.0 | ~0.8 | ~1.8 x 10^5 | ~0.14 | Murine model studies |
| ABD (Albumin-Binding Domain) | 7.4 | ~0.01 | ~1 x 10^6 | ~0.01 | High-affinity binding at neutral pH |
| IgG1 (Fc) | 6.0 | ~0.5 - 2 | ~1.5 x 10^5 | ~0.1 | Comparison standard |
Core Experimental Protocols
Protocol: Evaluating pH-Dependent FcRn Binding via SPR
Objective: Quantify binding kinetics of albumin-fusion proteins to human FcRn at acidic vs. neutral pH.
Materials:
- Biacore T200 or equivalent SPR instrument.
- Series S CMS sensor chip.
- Recombinant human FcRn (extracellular domain).
- Test articles: HSA, HSA-fusion, albumin-binding conjugate.
- Running Buffers: HBS-EP+ pH 6.0 (10 mM MES, 150 mM NaCl, 3 mM EDTA, 0.05% P20); HBS-EP+ pH 7.4.
- Amine coupling reagents: EDC, NHS, ethanolamine-HCl.
Method:
- FcRn Immobilization: Activate CMS chip surface with 1:1 EDC/NHS for 420s. Dilute hFcRn to 10 µg/mL in 10 mM sodium acetate pH 5.0. Inject for 300s to achieve ~5000 RU. Deactivate with ethanolamine.
- Kinetic Analysis: Dilute albumin-fusion analytes in respective pH buffers (6.0 and 7.4). Inject at 5 concentrations (e.g., 0.5, 1, 2, 4, 8 µM) at 30 µL/min for 180s association, followed by 600s dissociation in pH-matched buffer.
- Regeneration: After each cycle, regenerate surface with two 30s pulses of HBS-EP+ pH 7.4.
- Data Processing: Double-reference data. Fit to a 1:1 Langmuir binding model using Biacore Evaluation Software to derive ka, kd, and KD.
Protocol: In Vivo Pharmacokinetic Study in hFcRn Transgenic Mice
Objective: Assess half-life extension of an albumin-conjugated drug.
Materials:
- B6.mFcRn-/-.hFcRn Tg32 homozygous mice (express human FcRn).
- Test article: Alexa Fluor 680-labeled albumin-conjugate.
- Control: Alexa Fluor 750-labeled unconjugated protein.
- IVIS Spectrum or similar for fluorescence imaging; LC-MS/MS for quantitative analysis.
Method:
- Dosing: Administer a co-formulated mixture (2 mg/kg each conjugate and control) via tail vein injection (n=5/group).
- Serial Blood Collection: Collect ~20 µL blood via submandibular bleed at 2 min, 1, 4, 8, 24, 48, 72, 96, 120 hrs post-dose.
- Sample Analysis:
- Fluorescence: Plasma fluorescence measured (Ex/Em 680/720 and 750/780 nm). Calculate concentration from standard curve.
- LC-MS/MS: For precise quantification, digest plasma samples, isolate signature peptide, and analyze via MRM.
- PK Modeling: Use non-compartmental analysis (WinNonlin) to calculate AUC0-inf, clearance (CL), volume of distribution (Vd), and terminal half-life (t1/2).
Visualizations
Diagram 1: Therapeutic Co-option of the FcRn-Albumin Recycling Pathway
Diagram 2: R&D Workflow for Albumin-Based Half-Life Extension
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for FcRn-Albumin Research
| Reagent / Material | Supplier Examples | Function & Brief Explanation |
|---|---|---|
| Recombinant Human FcRn (extracellular domain) | Sino Biological, R&D Systems | Critical ligand for in vitro binding studies (SPR, ELISA) to validate pH-dependent interaction. |
| hFcRn Transgenic Mouse Model (B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr) | The Jackson Laboratory (Stock #014565) | In vivo model expressing human FcRn; essential for predictive PK studies of albumin-interacting therapeutics. |
| Surface Plasmon Resonance (SPR) Instrument | Cytiva (Biacore), Sartorius | Gold-standard for label-free, real-time kinetic analysis of protein-protein interactions (e.g., FcRn:Albumin-Fusion KD). |
| Albumin Depletion Kit (Human Serum) | Thermo Fisher, Sigma-Aldrich | Removes endogenous albumin from serum/plasma for cleaner analysis of conjugated drug fractions or for assay controls. |
| Site-Specific Conjugation Kits (e.g., Cys-34 Maleimide, Lysine) | Abzena, ProteoGenix | Enables reproducible chemical conjugation of payloads to specific residues on albumin or albumin-fusion proteins. |
| Human Hepatocyte Cell Line (HCCT-1 or HepG2) | ATCC | Expresses FcRn endogenously; used for cell-based recycling and transcytosis assays to model in vivo rescue. |
| Stable Cell Line for HSA-Fusion (CHO-DG44) | Thermo Fisher, Lonza | Preferred mammalian host for high-yield, consistent production of recombinant albumin-fusion proteins with proper folding. |
| Anti-HSA (Conformation-Specific) Antibodies | Antibodies-Online, Abcam | Detect native, folded HSA in complexes; crucial for ELISA, Western blot, and pharmacokinetic immunoassays. |
| pH-Sensitive Fluorophore (e.g., pHrodo) | Thermo Fisher | Labels endocytic vesicles; used in imaging assays to track internalization and intracellular trafficking of albumin-conjugates. |
Within the context of FcRn receptor-mediated albumin half-life extension research, engineering the neonatal Fc receptor (FcRn) binding interface of immunoglobulin G (IgG) has become a pivotal strategy for modulating antibody pharmacokinetics. Mutations such as M252Y/S254T/T256E (YTE) and M428L/N434S (LS) are designed to enhance pH-dependent binding, improving endosomal recycling and extending serum half-life. This whitepaper provides an in-depth technical analysis of how these mutations alter binding kinetics at the early endosomal pH of 6.0 versus the physiological pH of 7.4, detailing experimental methodologies, quantitative data, and practical research tools for characterization.
The extended half-life of IgG and albumin is governed by their interaction with FcRn, a process strictly dependent on pH. At the slightly acidic pH (~6.0) of the endosome, FcRn binds with high affinity to the Fc region of IgG, diverting it from the lysosomal degradation pathway and recycling it back to the cell surface. Upon exposure to the neutral pH (~7.4) of the blood, the complex rapidly dissociates, releasing the IgG back into circulation. Fc engineering aims to enhance this interaction by increasing affinity at pH 6.0 while maintaining minimal binding at pH 7.4, thereby optimizing the efficiency of the recycling process.
Core Diagram: FcRn-Mediated IgG Recycling and Mutagenesis Sites
Key Fc Engineering Mutations: Mechanism and Impact
The YTE and LS mutations are located at the Fc-FcRn interface, introducing residues that enhance electrostatic and hydrophobic interactions specifically under acidic conditions.
- YTE (M252Y/S254T/T256E): This triple mutation introduces a tyrosine and a negatively charged glutamate, strengthening hydrogen bonding and electrostatic interactions with FcRn residues (e.g., His161, Asp130) that are protonated at pH 6.0.
- LS (M428L/N434S): These substitutions enhance hydrophobic packing (M428L) and introduce a serine (N434S) that can form a hydrogen bond with FcRn Glu115, significantly increasing affinity at pH 6.0 with a more modest effect on pH 7.4 dissociation.
Quantitative Binding Kinetics Data
Binding kinetics are primarily measured using surface plasmon resonance (SPR) or biolayer interferometry (BLI). The key parameters are the dissociation constant (KD), association rate (ka), and dissociation rate (kd) at both pH conditions.
Table 1: Comparative Binding Kinetics of Fc Variants to Human FcRn
| Fc Variant | pH | KD (nM) | ka (1/Ms) | kd (1/s) | Fold Improvement in KD (vs WT at pH 6.0) | Reference Model |
|---|---|---|---|---|---|---|
| Wild-type (WT) | 6.0 | 300 - 600 | 1.0e5 - 2.0e5 | 6.0e-3 - 1.2e-2 | 1x | IgG1 baseline |
| Wild-type (WT) | 7.4 | > 10,000 | ND | Very Fast | - | Rapid dissociation |
| YTE (M252Y/S254T/T256E) | 6.0 | 30 - 90 | 2.5e5 - 4.0e5 | 8.0e-4 - 2.7e-3 | ~10x | Dall'Acqua et al., J Immunol (2006) |
| YTE | 7.4 | > 5,000 | ND | Fast | - | Minimal binding retained |
| LS (M428L/N434S) | 6.0 | 1 - 4 | 3.0e5 - 5.0e5 | 3.0e-4 - 2.0e-3 | ~100x | Zalevsky et al., Nat Biotechnol (2010) |
| LS | 7.4 | 200 - 500 | ND | ~0.1 | - | Noticeably increased vs WT |
ND: Not determinable due to very weak/transient binding.
Experimental Protocols for Characterizing pH-Dependent Binding
Protocol: Surface Plasmon Resonance (SPR) Analysis
This protocol details the characterization of Fc variant binding to immobilized FcRn at two pH conditions.
Objective: To determine the kinetic rate constants (ka, kd) and equilibrium dissociation constant (KD) for Fc-FcRn interaction at pH 6.0 and 7.4.
Materials:
- SPR Instrument: (e.g., Biacore T200, Cytiva)
- Sensor Chip: CMS Series S
- Running Buffers:
- pH 6.0: 50-100 mM sodium phosphate, 150 mM NaCl, 0.05% P20 surfactant, pH 6.0.
- pH 7.4: HBS-EP+ (10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% P20 surfactant, pH 7.4).
- Regeneration Buffer: 100-200 mM sodium phosphate, 150 mM NaCl, pH 7.4 - 8.0.
- Analytes: Purified Fc variants (WT, YTE, LS) at concentrations spanning 0.5-1000 nM (prepare in respective running buffer).
- Ligand: Recombinant human FcRn (with β2-microglobulin). Biotinylated for capture on a SA chip or amine-coupled.
Procedure:
- Ligand Immobilization: Capture biotinylated FcRn on a streptavidin (SA) chip to a density of 200-400 Response Units (RU). For amine coupling, target 500-1000 RU.
- Kinetic Experiment Setup: Set instrument temperature to 25°C. Use pH 6.0 running buffer for the pH 6.0 binding cycle.
- Association Phase: Inject a 2- or 3-fold dilution series of each Fc analyte (e.g., 500 nM to 3.9 nM) over the FcRn and reference surfaces for 180-300 seconds at a flow rate of 30 µL/min.
- Dissociation Phase: Switch to running buffer only and monitor dissociation for 600-900 seconds.
- Regeneration: Inject regeneration buffer (pH 7.4-8.0) for 30-60 seconds to completely dissociate any remaining complex.
- Repeat at pH 7.4: Change the running buffer to pH 7.4. Prepare fresh analyte dilutions in pH 7.4 buffer. Repeat injection series. Due to very fast off-rates, a single-cycle kinetics (SCK) method may be preferable.
- Data Analysis: Double-reference the sensorgrams (reference surface & blank injection). Fit data to a 1:1 Langmuir binding model using the instrument's evaluation software. Report ka, kd, and KD for each variant at each pH.
Protocol: Cellular Recycling Assay
A functional assay to measure the half-life extension conferred by mutations in a cellular system expressing human FcRn.
Objective: To compare the recycling efficiency of IgG containing YTE or LS mutations versus WT in an in vitro model.
Materials:
- Cell Line: MDCK or HEK293 cells stably transfected with human FcRn and β2-microglobulin.
- IgG Variants: WT, YTE, LS antibodies (non-antigen binding or against a neutral target), labeled with pH-sensitive fluorescent dye (e.g., pHrodo) or biotin.
- Buffers: Acidic binding buffer (DMEM, pH 6.0 with MES), neutral release buffer (DMEM, pH 7.4 with HEPES), stripping buffer (PBS, pH 3.0).
- Detection Method: Fluorescence plate reader or flow cytometry. For biotin, use streptavidin-HRP/AP or fluorescent streptavidin.
Procedure:
- Cell Seeding: Seed FcRn-expressing cells in a 24- or 96-well plate and culture to confluence.
- Pulse (Binding at pH 6.0): Wash cells with cold pH 6.0 binding buffer. Incubate with a fixed concentration (e.g., 5 µg/mL) of labeled IgG variant in binding buffer for 1-2 hours at 4°C (prevents internalization).
- Chase (Internalization & Recycling): Wash cells extensively with cold pH 7.4 buffer to remove unbound IgG. Add pre-warmed (37°C) pH 7.4 release buffer to initiate internalization and recycling. Incubate for 0-24 hours.
- Sampling: At designated time points (e.g., 0, 2, 4, 8, 24h), collect supernatant. Lyse cells to retrieve retained IgG.
- Quantification: Measure the amount of IgG in the supernatant (recycled) and cell lysate (retained) using the appropriate detection method (fluorescence, ELISA).
- Data Analysis: Calculate the percentage of total internalized IgG that is recycled over time. Compare the recycling half-life and efficiency between Fc variants.
Diagram: Experimental Workflow for Binding & Recycling Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Fc-FcRn Interaction Studies
| Item | Function & Description | Example Product/Catalog |
|---|---|---|
| Recombinant Human FcRn (hFcRn): | Purified soluble receptor (co-expressed with β2-microglobulin). Serves as the ligand for in vitro binding assays. | Sino Biological 10377-H08H; Themo Fisher PHC2314. |
| Anti-Human Fc (pH-sensitive) Antibodies: | Antibodies that specifically recognize the Fc region only at pH 6.0 (not at 7.4). Critical for specific detection in recycling assays. | Bio-Rad (formerly AbD Serotec) "HIS-1G5" clone. |
| Biacore Sensor Chips (Series S, SA): | Gold sensor chips with a dextran matrix. The SA (streptavidin) version is ideal for capturing biotinylated FcRn. | Cytiva 29104988 (CMS, SA). |
| pHrodo Red/Green STP Ester: | Amine-reactive, pH-sensitive fluorescent dye. Non-fluorescent at neutral pH, brightly fluorescent at acidic pH. Ideal for labeling IgG to track internalization. | Thermo Fisher P36600 / P35368. |
| hFcRn-Expressing Cell Lines: | Stably transfected cell lines (MDCK, HEK293) expressing human FcRn and β2M. Essential for functional cellular recycling studies. | ATCC (engineered lines); in-house generation common. |
| MES & HEPES Buffers: | High-quality buffering agents for precise pH control at 6.0 (MES) and 7.4 (HEPES) in binding/release buffers. | Sigma-Aldrich M3671 / H4034. |
| SPR Running Buffer Salts & Surfactant: | Components for preparing low non-specific interaction buffers (e.g., HBS-EP+). P20 surfactant (Tween 20) reduces sticking. | Cytiva BR100669 (HBS-EP+). |
The strategic introduction of Fc mutations like YTE and LS represents a mature and highly effective protein engineering approach to modulate FcRn interaction kinetics. The data unequivocally show that these mutations significantly enhance binding at pH 6.0, with LS offering exceptionally high affinity, while largely preserving the critical pH-sensitive release at 7.4. This translates directly to prolonged serum half-life observed in preclinical and clinical studies, validating the core thesis of FcRn-targeted half-life extension. Future research is exploring next-generation mutations (e.g., variants of YTE/LS, combination mutants) and applying these principles beyond IgG to other Fc-fusion proteins and directly to albumin itself, further expanding the therapeutic potential of this pivotal biological pathway.
Albumin-Binding Domains (ABDs) and Nanobodies as Alternative Scaffolds
Within the field of therapeutic protein engineering, extending plasma half-life is a critical challenge. The neonatal Fc receptor (FcRn)-mediated recycling pathway, which naturally extends the half-life of albumin and IgG, presents a prime target for half-life extension technologies. This whitepaper provides an in-depth technical analysis of two leading alternative scaffold platforms engineered to exploit this pathway: Albumin-Binding Domains (ABDs) and Nanobodies. We detail their molecular engineering, experimental characterization, and application in creating long-acting biotherapeutics.
Therapeutic proteins often suffer from short plasma half-lives due to renal clearance and proteolytic degradation. The FcRn receptor, expressed in endothelial cells and hematopoietic cells, binds to albumin and IgG in a pH-dependent manner, rescuing them from lysosomal degradation and recycling them back into circulation. This biological mechanism is the cornerstone for half-life extension strategies. Direct fusion to albumin or IgG Fc domains is established but has limitations in size, manufacturing, and tissue penetration. Small, engineered alternative scaffolds like ABDs and Nanobodies offer versatile solutions to harness FcRn recycling, primarily via albumin binding.
Technical Deep Dive: Albumin-Binding Domains (ABDs)
Origin and Engineering
ABDs are small (5-10 kDa), stable protein domains derived from bacterial surface proteins, such as Protein G from Streptococcus. Wild-type domains bind with high affinity to human albumin. Through directed evolution and rational design, engineered ABD variants (e.g., ABD) have been developed with:
- Increased Affinity: Picomolar to low nanomolar affinity for human serum albumin (HSA).
- Species Cross-Reactivity: Engineered to bind both human and preclinical species albumin.
- Robust Biophysical Properties: High thermal stability and resistance to aggregation.
Mechanism of Action
ABDs are fused to the therapeutic protein (e.g., an enzyme, cytokine, or nanobody). The ABD binds endogenous albumin, effectively hijacking its long half-life (~19-27 days in humans) via the FcRn recycling pathway.
Diagram: ABD-Mediated Half-Life Extension Pathway
Key Experimental Protocols
Protocol 1: Surface Plasmon Resonance (SPR) for ABD-Albumin Binding Kinetics
- Objective: Determine affinity (KD), association (ka), and dissociation (kd) rates.
- Method:
- Immobilization: Covalently immobilize recombinant human albumin on a CMS sensor chip using amine coupling to achieve ~1000 RU.
- Running Buffer: HBS-EP+ (10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% P20 surfactant, pH 7.4).
- Analysis: Serial dilutions of purified ABD or ABD-fusion protein (0.1-100 nM) are injected over the albumin surface at 30 µL/min.
- Regeneration: Surface is regenerated with a 30-second pulse of 10 mM Glycine-HCl, pH 2.0.
- Data Processing: Double-reference subtracted data is fitted to a 1:1 Langmuir binding model using Biacore Evaluation Software.
Protocol 2: Pharmacokinetic (PK) Study of ABD-Fusion in Mice
- Objective: Evaluate in vivo half-life extension.
- Method:
- Animal Model: C57BL/6 mice (n=5 per group).
- Dosing: Administer a single intravenous (IV) bolus of the ABD-fusion protein and a non-ABD control at 1 mg/kg.
- Sampling: Collect serial retro-orbital or tail vein blood samples at 2 min, 30 min, 2h, 8h, 24h, 48h, 96h, and 168h post-dose into EDTA tubes.
- Bioanalysis: Separate plasma. Quantify protein concentrations using a specific sandwich ELISA (e.g., anti-therapeutic protein capture, HRP-conjugated anti-tag detection).
- PK Analysis: Fit concentration-time data using a non-compartmental model (WinNonlin) to calculate terminal half-life (t1/2), clearance (CL), and area under the curve (AUC).
Technical Deep Dive: Nanobodies
Origin and Engineering
Nanobodies (VHHs) are the antigen-binding variable domains of heavy-chain-only antibodies from camelids. They are small (~15 kDa), monomeric, and possess excellent solubility and stability. For half-life extension, two primary strategies are employed:
- Anti-Albumin Nanobodies: Engineered to bind specifically to albumin, functioning similarly to ABDs.
- Anti-FcRn Nanobodies: Engineered to modulate the FcRn pathway itself (e.g., inhibit IgG binding to increase clearance in autoimmunity, or enhance binding to extend half-life).
Mechanism of Action
Diagram: Dual Nanobody Strategies for FcRn Pathway Modulation
Key Experimental Protocols
Protocol 3: Biolayer Interferometry (BLI) for Anti-FcRn Nanobody Characterization
- Objective: Assess pH-dependent binding kinetics of anti-FcRn nanobodies.
- Method:
- Loading: Load biotinylated recombinant human FcRn onto Streptavidin (SA) biosensors.
- Baseline: Establish baseline in kinetics buffer (pH 6.0 or 7.4).
- Association: Dip sensors into wells containing the anti-FcRn nanobody (50-500 nM) for 300 seconds at both pH 6.0 and pH 7.4.
- Dissociation: Transfer sensors to wells with kinetics buffer only for 400 seconds.
- Regeneration: A mild regeneration step (pH 7.4 buffer) may be used.
- Analysis: Data is processed and fitted using the ForteBio Data Analysis software to determine binding kinetics at each pH, confirming the desired pH-dependent profile.
Protocol 4: Cellular Transcytosis Assay
- Objective: Demonstrate FcRn-mediated recycling of an anti-albumin nanobody fusion.
- Method:
- Cell Culture: Use polarized MDCK or hCMEC/D3 cells stably expressing human FcRn, grown on Transwell inserts.
- Apical Loading: Add the nanobody-albumin complex (pre-formed) or control to the apical chamber at pH 6.0.
- Incubation: Incubate at 37°C for 2-4 hours.
- Sampling: Collect medium from the basolateral chamber (pH 7.4).
- Quantification: Measure the amount of transported nanobody fusion in the basolateral chamber using ELISA or LC-MS/MS.
- Inhibition Control: Repeat in the presence of a known FcRn blocker (e.g., excess IgG) to confirm pathway specificity.
Comparative Data and Research Toolkit
Table 1: Quantitative Comparison of ABDs and Nanobodies as Scaffolds
| Parameter | Albumin-Binding Domain (ABD) | Nanobody (VHH) |
|---|---|---|
| Molecular Weight | ~5-10 kDa | ~12-15 kDa |
| Origin | Bacterial (e.g., Protein G) | Camelid/Humanized |
| Albumin Affinity (KD) | Low pM - nM range | Low pM - nM range (for anti-albumin VHHs) |
| Primary Half-Life Extension Mechanism | Non-covalent binding to endogenous albumin | 1. Covalent/Non-covalent binding to albumin. 2. Direct FcRn modulation. |
| Typical PK Half-Life Extension (in mice, vs. scaffold alone) | From hours to ~2-3 days | From hours to ~2-4 days (albumin-binding) |
| Format Flexibility | N-terminal or C-terminal fusion | Multivalent, biparatopic, fused to other proteins |
| Key Advantage | Small size, high stability, simple mechanism. | Versatility (target albumin or FcRn), excellent tissue penetration. |
| Key Challenge | Potential competition with endogenous albumin ligands. | Immunogenicity risk (requires humanization). |
Table 2: The Scientist's Toolkit: Essential Research Reagents & Materials
| Reagent/Material | Function/Explanation | Example Vendor/Cat. No. |
|---|---|---|
| Recombinant Human Serum Albumin (rHSA) | High-purity antigen for in vitro binding assays (SPR/BLI) and complex formation. | Sigma-Aldrich (A9731) |
| Biacore Series S Sensor Chip CMS | Gold standard SPR chip for covalent ligand immobilization. | Cytiva (BR100530) |
| Anti-6X His Tag Antibody (HRP) | Detection antibody for quantifying His-tagged ABD/Nanobody fusions in ELISA. | Abcam (ab1187) |
| Recombinant Human FcRn Protein (Biotinylated) | Critical for characterizing pH-dependent binding of anti-FcRn nanobodies in BLI. | Acro Biosystems (FCM-H82E6) |
| MDCK-II/hFcRn Cell Line | Polarized epithelial cell model for in vitro transcytosis and recycling assays. | Generated in-house or via licensing. |
| Pierce pH 6.0 & 7.4 Kinetics Buffers | Pre-formulated buffers for reliable pH-dependent binding kinetics studies. | Thermo Fisher (PR-100053) |
| Microvette CB 300 LH Capillary Blood Collection Tubes | For efficient, serial micro-sampling in murine PK studies. | Sarstedt (16.440.100) |
Both ABDs and nanobodies represent powerful, complementary alternative scaffold technologies for half-life extension via the FcRn-albumin axis. ABDs offer a minimalistic, robust "albumin hitchhiking" approach, while nanobodies provide unparalleled versatility to either bind albumin or directly engage and modulate the FcRn receptor. The choice depends on the specific therapeutic goal, desired pharmacology, and development constraints. Continued research is focused on developing next-generation variants with enhanced affinity, pH-responsiveness, and reduced immunogenicity, further solidifying their role in the next wave of long-acting biotherapeutics.
This whitepaper presents a technical analysis of key biologics whose pharmacokinetic profiles have been optimized through FcRn-mediated recycling, framed within the broader thesis of albumin half-life extension research. By leveraging the protective interaction with the neonatal Fc receptor (FcRn), these therapeutics achieve prolonged systemic exposure, reducing dosing frequency and improving patient outcomes.
The neonatal Fc receptor (FcRn) is a pivotal regulator of serum half-life for both IgG and albumin. The central thesis of this research field posits that engineered fusion or conjugation of therapeutic proteins to the Fc domain of IgG or to albumin itself hijacks this natural recycling pathway. Following pinocytosis, biologics bound to FcRn in the acidic endosome are rescued from lysosomal degradation and recycled to the cell surface, where neutral pH facilitates release back into circulation. This review examines approved and clinical-stage case studies that validate this thesis.
Approved FcRn-Enhanced Biologic: Efanesoctocog Alfa (Antihemophilic Factor [Recombinant], Fc-VWF-XTEN-Fusion Protein)
Efanesoctocog alfa is a groundbreaking high-sustained factor VIII (FVIII) therapy for hemophilia A, explicitly engineered for FcRn engagement.
Mechanism: It is a recombinant FVIII molecule fused to the Fc domain of IgG1, the von Willebrand factor (VWF) D'D3 domain, and an XTEN polypeptide. The Fc domain directly facilitates FcRn binding and recycling.
Key Pharmacokinetic Data: Table 1: Summary of Key PK Parameters for Efanesoctocog Alfa (from Phase 3 study)
| Parameter | Value | Implication |
|---|---|---|
| Geometric Mean Half-life | 47.0 hours | ~3-4x longer than standard rFVIII |
| Mean Terminal Half-life | 50.3 hours | Sustained protective activity |
| Weekly Dosing Regimen | 50 IU/kg once weekly | Validates half-life extension thesis |
Supporting Experimental Protocol (Chromogenic Assay for FVIII Activity):
- Sample Preparation: Serial dilutions of patient plasma samples (post-administration) are prepared in FVIII-deficient plasma.
- Assay Setup: Diluted samples are incubated with excess factor IXa, factor X, phospholipids, and calcium in buffer.
- Reaction: Activated FVIII (in sample) acts as a cofactor for FIXa, which activates FX to FXa.
- Detection: A chromogenic substrate specific for FXa is added. Cleavage releases a colored product (p-nitroaniline).
- Quantification: Absorbance at 405 nm is measured. Activity is calculated from a standard curve of known FVIII concentrations.
Clinical-Stage Case Study: Albinterferon Alfa-2b (Fusion to Human Serum Albumin)
Albinterferon alfa-2b was an investigational therapy for chronic hepatitis C, representing an early clinical proof-of-concept for albumin fusion technology.
Mechanism: Recombinant interferon-alfa-2b is genetically fused directly to human serum albumin (HSA). The HSA moiety binds to FcRn, conferring its extended half-life onto the interferon payload.
Key Clinical Trial Data: Table 2: Pharmacokinetic Comparison of Albinterferon vs. Peginterferon alfa-2a
| Parameter | Albinterferon (900μg Q2W) | Peginterferon alfa-2a (180μg QW) | Outcome |
|---|---|---|---|
| Apparent Half-life | ~150 hours | ~80 hours | ~2-fold increase |
| Dosing Interval | Every 2 weeks | Every week | Reduced frequency |
| Clinical Efficacy (SVR) | Non-inferior (in trials) | Comparator | Validated fusion approach |
| Primary Discontinuation Reason | Pulmonary Complications | Standard adverse events | Led to development halt |
Supporting Experimental Protocol (SPR Analysis of FcRn Binding):
- Surface Preparation: FcRn is immobilized on a CM5 sensor chip via amine coupling.
- Running Buffer: Use a two-buffer system: Association phase (pH 6.0, mimics endosome), Dissociation phase (pH 7.4, mimics bloodstream).
- Ligand Injection: Serial concentrations of the albumin-fusion protein (Analytes) are injected in pH 6.0 buffer over the FcRn surface.
- Association & Dissociation: Binding is measured in real-time. Buffer is switched to pH 7.4 to monitor complex dissociation.
- Regeneration: The surface is regenerated with a mild pH 7.4 buffer for the next cycle.
- Analysis: Sensorgrams are fit to a 1:1 binding model to determine kinetic rates (ka, kd) and affinity (KD).
Visualizing the Core Mechanism and Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for FcRn-Albumin Research
| Research Reagent | Primary Function & Application |
|---|---|
| Recombinant Human FcRn Protein | In vitro binding studies (SPR, BLI), cell-based trafficking assays. Critical for quantifying fusion protein affinity. |
| pH-Switch Assay Buffers (pH 6.0 & 7.4) | Mimic endosomal and physiological pH conditions to study the pH-dependent binding kinetics essential for FcRn recycling. |
| Anti-Human Albumin Antibodies (non-FcRn blocking) | Used in ELISA or immunoassays to quantify albumin-fusion protein concentration in biological matrices without interfering with the FcRn binding site. |
| Human Endothelial Cell Lines (e.g., HMEC-1) | Model system to study the cellular recycling and transcytosis of FcRn-enhanced biologics in a relevant physiological context. |
| Plasma from FcRn Knockout Mice | Control matrix for in vivo studies to confirm the specific role of FcRn in extending the half-life of the test biologic. |
| Chromogenic/Catalytic Activity Assay Kits | For functional PK assessment of enzyme or coagulation factor fusions (e.g., FVIII, FIX activity kits). |
| Surface Plasmon Resonance (SPR) Chip (CM5 series) | Gold-standard biosensor surface for immobilizing FcRn and performing detailed kinetic analysis of fusion protein binding. |
Optimizing FcRn Engagement: Navigating Affinity, Specificity, and Developability Challenges
The neonatal Fc receptor (FcRn) is a critical regulator of serum half-life for both IgG antibodies and albumin. The central thesis of this field posits that engineering pH-dependent binding to FcRn—strong binding in the acidic endosome (pH ~6.0) and rapid release at neutral blood pH (7.4)—can enhance recycling and extend plasma half-life. However, the ultimate challenge is to optimize this interaction without impairing the natural turnover pathways or disrupting FcRn's physiological functions for endogenous proteins. This whitepaper provides a technical guide to achieving this balance.
Quantitative Data on Key FcRn Binding Mutants
Table 1: Comparison of Engineered Fc Variants for pH-Dependent FcRn Binding
| Variant Name (Common) | Key Mutations (EU numbering) | Binding Affinity to hFcRn at pH 6.0 (KD, nM)* | Binding Affinity to hFcRn at pH 7.4 (KD, nM)* | Reported Human/Primate Half-Life Extension vs. Wild-Type |
|---|---|---|---|---|
| Wild-Type IgG1 | - | 500 - 900 | >10,000 | 1x (Baseline, ~21 days) |
| YM | M252Y/S254T/T256E | 30 - 50 | ~5,000 | 2.5x - 4x |
| LS | M428L/N434S | 15 - 35 | >10,000 | 2.5x - 3.5x |
| XTEN Fusion | N/A (Polypeptide fusion) | N/A | N/A | 2x - 3x (Mechanism differs) |
| ABDEGER | H433K/N434F/Y436H | <10 | >10,000 | Up to 4.5x |
| MST-HN | M252Y/S254T/T256E/H433K/N434F | <5 | >10,000 | 4x - 5.5x |
Note: Affinity values are approximate and compiled from recent literature (2023-2024). Significant variation exists depending on measurement technique (e.g., SPR, BLI). The goal is a high affinity ratio (~100-1000x) between pH 6.0 and 7.4.
Table 2: Potential Disruption Risks of Over-Optimization
| Risk Parameter | Consequence of Excessive FcRn Affinity at pH 6.0 | Experimental Measurement |
|---|---|---|
| Endogenous IgG Displacement | Autoantibody elevation, immune complex disease | Competitive ELISA with labeled endogenous IgG |
| FcRn Saturation | Accelerated clearance of endogenous IgG/Albumin, hypoalbuminemia | In vivo tracer studies with co-administered wild-type IgG |
| Impaired Release at pH 7.4 | Reduced recycling, paradoxical shorter half-life | Off-rate (kd) measurement at pH 7.4 via SPR |
| Altered Tissue Distribution | Accumulation in FcRn-expressing tissues (endothelia, phagocytes) | Quantitative Whole-Body Autoradiography (QWBA) |
Core Experimental Protocols
Protocol 1: In Vitro Assessment of pH-Dependent FcRn Binding via Surface Plasmon Resonance (SPR)
Objective: Quantify binding kinetics (ka, kd, KD) at pH 6.0 and 7.4.
- Chip Preparation: Immobilize recombinant biotinylated hFcRn (~500-1000 RU) on a Series S Sensor Chip SA.
- Running Buffer: For pH 6.0 binding: 50 mM MES, 150 mM NaCl, 0.05% P-20, pH 6.0. For pH 7.4 dissociation: PBS-P+, pH 7.4.
- Analyte Injection: Dilute Fc-engineered antibody samples in pH 6.0 buffer. Inject over flow cells for 180s at 30 μL/min.
- pH Switch: Immediately switch to pH 7.4 running buffer and monitor dissociation for 600s.
- Regeneration: Chip surface is regenerated with a 30s pulse of pH 8.0, 3M MgCl₂.
- Data Analysis: Fit association/dissociation phases using a 1:1 Langmuir binding model. Calculate the affinity ratio: KD(pH7.4) / KD(pH6.0).
Protocol 2: Cellular Recycling Assay Using Human Endothelial Cells
Objective: Measure FcRn-mediated recycling efficiency of engineered antibodies.
- Cell Culture: Grow human umbilical vein endothelial cells (HUVECs) expressing FcRn in 24-well plates.
- Pulse (Internalization): Incubate cells with 10 μg/mL Alexa Fluor 488-labeled test antibody in acidic medium (pH 6.0, MES-buffered) for 1h at 37°C.
- Wash: Remove unbound antibody with cold acidic buffer.
- Chase (Recycling): Incubate cells in pre-warmed neutral medium (pH 7.4) for 0, 30, 60, 120 min.
- Quantification: At each time point, collect the medium (recycled fraction) and lyse cells (retained fraction). Measure fluorescence of both fractions. Calculate % recycled = (Recycled Signal / (Recycled + Retained Signal)) * 100.
Protocol 3: In Vivo Pharmacokinetic Study in Humanized FcRn Transgenic Mouse Model
Objective: Evaluate half-life extension and competition with endogenous IgG.
- Model: Use B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr/DcrJ mice (expressing human FcRn).
- Dosing: Co-administer a single IV bolus of 5 mg/kg engineered antibody and 5 mg/kg control human IgG (labeled with different fluorophores) into mice (n=5-8/group).
- Sampling: Collect serial retro-orbital blood samples at 5 min, 6h, 1, 2, 4, 7, 14, 21, and 28 days post-dose.
- Bioanalysis: Measure serum concentrations of both antibodies using dual-fluorescence or antigen-capture ELISA specific for each.
- PK Analysis: Fit concentration-time data using a two-compartment model. Compare terminal half-life (t1/2β) and AUC between test and control antibody.
Visualizations
Diagram 1: FcRn-Mediated Recycling vs. Lysosomal Degradation
Diagram 2: Iterative Screening Workflow for Balanced Variants
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Item | Function/Description | Key Supplier Examples |
|---|---|---|
| Recombinant Human FcRn (monomeric & dimeric) | Critical ligand for in vitro binding assays (SPR/BLI). Must be properly biotinylated for capture. | Sino Biological, AcroBiosystems, Themo Fisher Scientific |
| Human FcRn Transgenic Mouse Model | In vivo gold-standard for predicting human PK; expresses human FcRn with mouse IgG clearance. | The Jackson Lab (Strain #014565), genOway |
| pH-Switch SPR/BLI Buffer Kits | Pre-formulated, degassed buffers at pH 6.0 and 7.4 for reliable kinetic screening. | Cytiva, ForteBio |
| HUVEC or HEK293 Cells Stably Expressing hFcRn | Cellular models for recycling and transcytosis assays. | ATCC, in-house generation via lentiviral transduction |
| Fluorescent Labeling Kits (pH-insensitive dyes) | For tagging antibodies for cellular trafficking and in vivo imaging studies (e.g., Alexa Fluor 647). | Thermo Fisher Scientific |
| Competitive ELISA Kits for Human IgG | To measure potential displacement of endogenous IgG in serum samples from PK studies. | Abcam, Bethyl Laboratories |
| LC-MS/MS Reagents for Peptide Mapping | To confirm engineered mutations and monitor chemical stability (e.g., deamidation) post-pH stress. | Waters, SCIEX |
Within the strategic framework of FcRn receptor-mediated albumin half-life extension research, the engineered therapeutic faces complex pharmacokinetic (PK) challenges. Foremost among these are Target-Mediated Drug Disposition (TMDD) and Altered Tissue Distribution. TMDD occurs when a significant drug-target interaction influences the PK profile, leading to non-linear, dose-dependent clearance. Concurrently, fusion to albumin or albumin-binding domains can redirect a molecule from its intended tissue compartment, altering efficacy and toxicity profiles. This whitepaper provides a technical guide for identifying, characterizing, and mitigating these risks in the context of albumin-half-life extended biologics.
Mechanisms and Quantitative Impact
TMDD in FcRn-Albumin Therapeutics
FcRn engagement is designed for recycling, but saturation of this pathway at high doses can paradoxically increase clearance. More critically, off-target binding or intended high-affinity target binding can drive TMDD.
Table 1: Reported TMDD PK Parameters for Albumin-Fused Therapeutics
| Therapeutic Modality | Target (KD) | Dose Showing Non-linearity | Clearance Change (vs. Low Dose) | Key TMDD Driver |
|---|---|---|---|---|
| Albumin-Coagulation Factor IX | Collagen (Extracellular Matrix) | < 50 IU/kg | >300% Increase | Off-target binding to vascular collagen |
| Anti-IL6 Albumin-Binding Domain Fab | IL-6R (100 pM) | 0.1 - 1 mg/kg | 150% Increase | High-affinity IL-6R sink in tissues |
| Albumin-GLP-1 Analog | GLP-1R (nM) | > 0.5 mg/kg | ~80% Increase | Receptor-mediated endocytosis in kidney/lung |
| FcRn-High Affinity mAb | FcRn (nM) at acidic pH | >10 mg/kg | 200% Increase | Saturation of FcRn recycling pathway |
Altered Tissue Distribution Profiles
Albumin’s natural distribution (vascular space, muscle, skin, liver) can divert a drug from its required site of action (e.g., brain, tumor core, specific immune cell).
Table 2: Tissue Partition Coefficients (Kp): mAb vs. Albumin-Fused Analog
| Tissue | Typical mAb Kp | Albumin-Fused Analog Kp | Implication |
|---|---|---|---|
| Plasma | 1.0 | 1.0 (Reference) | N/A |
| Liver | 0.1 - 0.2 | 0.3 - 0.5 | Increased hepatocyte uptake via FcRn/FcγR |
| Spleen | 0.2 - 0.4 | 0.15 - 0.3 | Moderately reduced lymphatic access |
| Kidney | 0.1 - 0.15 | 0.05 - 0.1 | Reduced glomerular filtration |
| Tumor (subcutaneous) | 0.05 - 0.15 | 0.02 - 0.08 | Reduced extravasation and penetration |
| Skin | 0.2 - 0.3 | 0.5 - 0.8 | Significantly increased distribution |
Experimental Protocols for Risk Assessment
Protocol 1:In VivoPK Study to Detect TMDD
Objective: To characterize non-linear PK and identify TMDD.
- Dosing: Administer the albumin-fused therapeutic at four logarithmically spaced doses (e.g., 0.1, 1, 10, 50 mg/kg) intravenously to cynomolgus monkeys (n=3/group).
- Sampling: Collect serial plasma samples pre-dose and at 0.083, 1, 6, 24, 72, 168, 240, and 336 hours post-dose.
- Bioanalysis: Quantify plasma concentrations using a target-bridging ELISA (to detect intact complex) and a total albumin assay.
- Data Analysis: Fit data using non-compartmental analysis (NCA). Plot AUC and Clearance vs. Dose. A non-proportional increase in AUC or dose-dependent clearance indicates TMDD. Fit to a Michaelis-Menten (MM) PK model to estimate Vmax and Km.
Protocol 2: Quantitative Whole-Body Autoradiography (QWBA) for Tissue Distribution
Objective: To visualize and quantify altered tissue distribution.
- Radiolabeling: Label the therapeutic with Iodine-125 (125I) using the IODO-GEN method, ensuring specific activity of ~5 mCi/mg.
- Dosing & Sacrifice: Administer a single IV dose (5 mg/kg) to Sprague-Dawley rats (n=3/time point). Euthanize at 4, 24, 72, and 168 hours post-dose.
- Sample Preparation: Flash-freeze carcasses in hexane/dry ice. Embed in carboxymethylcellulose. Section sagittally at 30 µm using a cryomicrotome.
- Imaging & Quantification: Mount sections on tape, expose to phosphor imaging plates for 7 days. Scan plates and calibrate using embedded radiolabel standards. Calculate tissue:plasma concentration ratios (Kp) for >20 tissues.
Visualizing Pathways and Workflows
Diagram 1: TMDD & FcRn Recycling Interaction Pathways
(Title: TMDD and FcRn Pathways Compete for Drug Fate)
Diagram 2: Integrated TMDD & Distribution Risk Assessment Workflow
(Title: Integrated Risk Assessment Workflow for Developers)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for TMDD & Distribution Studies
| Reagent / Material | Function & Application | Key Consideration |
|---|---|---|
| Biacore T200 or Octet RED96e | Surface Plasmon Resonance (SPR) or Bio-Layer Interferometry (BLI) to quantify drug-target and drug-FcRn binding kinetics (KD, kon, koff). | Use immobilized target protein. Test at both pH 7.4 and 6.0 for FcRn interaction. |
| Radio-iodination Reagents (IODO-GEN) | For covalent labeling of proteins with 125I for QWBA or tissue extraction studies. | Ensures tracer integrity; must validate bioequivalence to unlabeled drug. |
| Species-Specific Target Protein (Recombinant) | For in vitro binding assays and as a calibrator in PK assays. | Essential for cross-reactivity testing and understanding species-specific TMDD risks. |
| Phosphor Imaging Plates & Scanner | To capture and digitize radiation from QWBA tissue sections for quantitative analysis. | High dynamic range is critical for quantifying both high (blood) and low (CNS) concentrations. |
| Validated Target-Bridging ELISA Kit | To measure intact, pharmacologically active drug concentration in complex biological matrices. | Must distinguish free drug from target-bound drug to accurately model TMDD. |
| PBPK Modeling Software (e.g., GastroPlus, Simcyp) | Physiologically-based pharmacokinetic modeling platform to integrate in vitro and in vivo data for human PK prediction. | Requires species-specific physiology modules and accurate in vitro input parameters. |
Mitigation Strategies
- Affinity Modulation: Engineer the therapeutic's target-binding domain to have a slightly lower affinity (optimal KD in low nM range) to reduce saturation and TMDD while maintaining efficacy.
- FcRn Binding Optimization: Fine-tune the albumin or Fc-binding domain's affinity for FcRn at pH 6.0 to maximize recycling without causing saturation-mediated clearance at therapeutic doses.
- Protease-Cleavable Linkers: Insert a substrate linker between the active moiety and albumin that is cleaved in the target tissue, decoupling distribution from activity.
- Dosing Regimen Design: Utilize model-informed drug development (MIDD) to design loading and maintenance doses that minimize non-linear clearance phases.
Proactively investigating the potential for TMDD and altered tissue distribution is non-negotiable for the successful development of FcRn-albumin half-life extension therapeutics. A hierarchical experimental approach—from precise in vitro kinetics to integrative PBPK modeling—provides the framework to quantify these risks. The mitigation strategies outlined enable the rational engineering of next-generation biologics that leverage the benefits of half-life extension while maintaining predictable, linear PK and targeted tissue delivery.
This technical guide addresses the core experimental challenges in characterizing the interaction between the neonatal Fc receptor (FcRn) and therapeutic Fc-fusion proteins, monoclonal antibodies, or albumin-based constructs. Accurate measurement under physiological pH conditions is paramount for predicting in vivo half-life and optimizing molecules for extended circulation, a central tenet of half-life extension research.
The pH-Dependent FcRn Binding Cycle
The fundamental challenge stems from the precise pH-switch mechanism governing FcRn salvage. Binding occurs efficiently in acidic endosomal compartments (pH ~5.5-6.0), while dissociation is forced at near-neutral blood pH (~7.4) to enable release back into circulation. Therefore, affinity and kinetic measurements are only physiologically relevant when performed across this pH gradient.
Table 1: Key Physiological pH Milestones in the FcRn Salvage Pathway
| Cellular Compartment | Approximate pH | Primary FcRn Function |
|---|---|---|
| Blood Plasma | 7.35 - 7.45 | No binding; release of ligand |
| Early Endosome | 6.0 - 6.5 | High-affinity binding of IgG/Albumin |
| Sorting Endosome | 5.5 - 6.0 | Cargo sorting and salvage |
| Lysosome | < 5.0 | Degradation of unbound ligand |
| Cell Surface (Apical) | ~7.4 | Possible non-recycling binding |
Core Analytical Methodologies & Protocols
Surface Plasmon Resonance (SPR) with pH Switching
SPR is the gold standard for measuring real-time kinetics (kon, *k*off, K_D) under controlled pH conditions.
Detailed Protocol:
- Immobilization: FcRn (human or relevant species) is immobilized on a CMS sensor chip via standard amine coupling to ~5000-8000 Response Units (RU).
- Running Buffer Preparation: Prepare HBS-EP buffers at two pH levels: Acidic (pH 5.5-6.0, e.g., MES-based) and Neutral (pH 7.4, PBS-based).
- Analyte Injection Series: The IgG or albumin variant analyte is prepared in the acidic running buffer. A concentration series (e.g., 0-1000 nM) is injected over the FcRn surface.
- pH-Switch Dissociation: After the association phase in acidic buffer, the flow is switched to neutral buffer for the dissociation phase, mimicking endosomal release.
- Regeneration: The surface is regenerated with a brief pulse of pH 7.4 buffer or a mild basic solution (e.g., pH 8.0).
- Data Analysis: A 1:1 Langmuir binding model incorporating a buffer switch component is used to fit the data, deriving ka, *k*d (at both pHs), and the overall affinity (K_D) at acidic pH.
Table 2: Representative SPR Kinetic Data for IgG1 Variants
| Molecule | k_a (1/Ms) at pH 5.8 | k_d (1/s) at pH 5.8 | k_d (1/s) at pH 7.4 | K_D (nM) at pH 5.8 | Relative in vivo Half-life |
|---|---|---|---|---|---|
| Wild-type IgG1 | 1.2 x 10^5 | 1.8 x 10^-3 | > 0.1 | 15 | 1.0 (reference) |
| YTE Mutant (M252Y/S254T/T256E) | 2.5 x 10^5 | 0.9 x 10^-3 | ~0.05 | 3.6 | ~4x extended |
| H435A Abrogated Mutant | 5.0 x 10^4 | 1.5 x 10^-3 | ~0.1 | 30 | Severely reduced |
Cell-Based Recycling Assays
This functional assay validates biophysical data in a physiologically relevant cellular context.
Detailed Protocol:
- Cell Culture: Use polarized epithelial cells (e.g., MDCK) or endothelial cells stably expressing human FcRn and its beta-2 microglobulin light chain.
- Radiolabeling/Iodination: The test antibody is labeled with Iodine-125 (¹²⁵I) or biotinylated.
- Internalization (Pulse): Cells are incubated with the labeled antibody in acidic medium (pH 6.0) at 37°C for 1-2 hours to allow binding and uptake.
- Chase & Release: The medium is replaced with a neutral pH (7.4) medium. The receptor releases the antibody apically (representing recycling) or basolaterally (representing transcytosis).
- Quantification: Aliquots of media from both compartments are collected over time (e.g., 30, 60, 120 min). Recycled antibody is quantified via gamma counting (for ¹²⁵I) or ELISA.
- Data Output: The percentage of total internalized antibody that is recycled is calculated, providing a direct functional readout of FcRn-mediated salvage efficiency.
Signaling & Workflow Diagrams
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for FcRn Binding Studies
| Item | Function & Rationale |
|---|---|
| Recombinant Human FcRn (hFcRn):β2m Heterodimer | Purified, soluble receptor for biophysical assays (SPR, BLI). Must be properly folded and glycosylated for accurate kinetics. |
| pH-Switch SPR Running Buffers | MES (pH 5.5-6.5) and HEPES/PBS (pH 7.4) buffers with matching ionic strength to minimize bulk refractive index shifts during switch. |
| CMS Series S Sensor Chip (for SPR) | Carboxymethylated dextran surface for stable, covalent amine coupling of FcRn. |
| Label-Free Cell Lines | MDCK-II or HEK293 cells stably overexpressing hFcRn/hβ2m. Critical for cell-based recycling/transcytosis assays. |
| Radiolabel (¹²⁵I) or High-Sensitivity Fluorophores | For tracing molecule internalization and fate in cell assays with minimal perturbation to Fc-FcRn interface. |
| Octet RED96e System & Anti-Human Fc (AHQ) Biosensors | For alternative kinetic screening using Bio-Layer Interferometry (BLI), enabling higher throughput than SPR. |
| Reference IgG with Known FcRn Affinity (e.g., wild-type IgG1) | Essential positive control to benchmark assay performance across experiments. |
| IgG/Ablumin Mutants (e.g., YTE, H435A) | Critical negative/positive controls for abrogated or enhanced binding in both biochemical and cellular assays. |
Formulation and Stability Considerations for Engineered Fusion Proteins
The development of engineered fusion proteins, particularly those leveraging the neonatal Fc receptor (FcRn) recycling pathway for half-life extension, represents a cornerstone of modern biologics research. The broader thesis on FcRn-albumin half-life extension research underscores that successful therapeutic translation hinges not only on molecular design but equally on rigorous formulation and stability assessment. These proteins, which often fuse therapeutic domains to Fc fragments or albumin, present unique physicochemical challenges. This guide details the core principles, analytical methodologies, and stabilization strategies essential for advancing such fusion proteins from discovery to clinical application.
Critical Stability Challenges for FcRn-Targeted Fusions
Fusion proteins designed for FcRn engagement face specific stressors that can compromise efficacy and safety.
- Chemical Instability: Deamidation (especially at Asn-Gly sequences), oxidation (of Met, Trp, His), and fragmentation can alter FcRn binding affinity and pharmacokinetics.
- Physical Instability: Aggregation is a primary concern, potentially induced by interfacial stress, pH shifts, or elevated temperature. Unfolding or misfolding can obscure the FcRn binding interface (located at the CH2-CH3 domain junction of Fc).
- Surface Adsorption: High-concentration formulations of potent fusions can suffer losses due to adsorption to container surfaces.
- Colloidal Instability: Electrostatic and hydrophobic interactions influenced by formulation excipients can affect solution behavior and viscosity.
Key Analytical Methods and Experimental Protocols
Forced Degradation Studies Protocol
Objective: To identify potential degradation pathways and inform formulation design. Materials: Test protein solution, buffers (pH 3-10), hydrogen peroxide (for oxidation), chemical denaturants (e.g., guanidine HCl). Procedure:
- Aliquot the fusion protein into vials containing different stress conditions.
- Thermal Stress: Incubate samples at 25°C, 40°C, and 50°C for 1-4 weeks.
- pH Stress: Adjust aliquots to pH 3.0, 5.0, 7.4, and 9.0 using appropriate buffers; incubate at 40°C for 1-2 weeks.
- Oxidative Stress: Add H₂O₂ to a final concentration of 0.1-0.3% to a sample; incubate at 25°C for 24-72 hours.
- Mechanical Stress: Perform vortexing or repeated freeze-thaw cycles (e.g., -80°C to 25°C, 5 cycles).
- Analyze all stressed samples alongside a control (2-8°C storage) using SE-HPLC, CE-SDS, and IEF.
FcRn Binding Affinity Measurement via Surface Plasmon Resonance (SPR)
Objective: Quantify the impact of stress or formulation on the critical FcRn binding interaction. Materials: Biacore or equivalent SPR instrument, Series S Sensor Chip CM5, recombinant human FcRn protein, HBS-EP+ running buffer (pH 6.0 and 7.4), amine coupling kit. Procedure:
- Immobilize the engineered fusion protein on a CM5 chip via standard amine coupling.
- Dilute FcRn in HBS-EP+ buffer at pH 6.0 (endosomal pH) to a concentration series (e.g., 0-1000 nM).
- Inject FcRn samples over the fusion protein surface at pH 6.0 for association, then switch to pH 7.4 buffer to monitor dissociation, mimicking the physiological recycling process.
- Regenerate the surface with a mild acidic buffer (pH 5.0).
- Analyze data using a 1:1 Langmuir binding model to determine the association rate (kₐ), dissociation rate (kₑ), and equilibrium dissociation constant (Kᴅ) at pH 6.0.
High-Concentration Viscosity and Self-Interaction Assessment
Objective: Evaluate developability for subcutaneous administration. Materials: Concentrated protein solution (>100 mg/mL), micro-viscometer (e.g., capillary-based), dynamic light scattering (DLS) instrument. Procedure:
- Concentrate the fusion protein formulation using centrifugal filters to the target concentration (e.g., 150 mg/mL).
- Measure kinematic viscosity at 25°C using the micro-viscometer. Compare to buffer control.
- Perform DLS in static and dynamic mode to determine the hydrodynamic radius (Rₕ) and the interaction parameter (kᴅ), derived from the concentration dependence of the diffusion coefficient.
Table 1: Representative Stability Profile of an Fc-Fusion Protein Under Various Stress Conditions
| Stress Condition | Duration | % Monomer (SE-HPLC) | % HMW Species | % Fragments | FcRn Kᴅ at pH 6.0 (nM) |
|---|---|---|---|---|---|
| Control (5°C) | 4 weeks | 99.5 | 0.3 | 0.2 | 25.1 |
| 40°C | 4 weeks | 96.8 | 2.5 | 0.7 | 28.5 |
| 50°C | 2 weeks | 89.2 | 9.1 | 1.7 | 45.3 |
| pH 3.0 | 1 week | 92.4 | 5.0 | 2.6 | >1000* |
| 0.1% H₂O₂ | 48 hours | 97.1 | 1.2 | 1.7 | 30.2 |
| 5 Freeze-Thaws | N/A | 98.9 | 0.8 | 0.3 | 26.0 |
*Indicates significant unfolding/denaturation.
Table 2: Formulation Screen Impact on Key Stability Indicators
| Formulation Buffer (All at 10 mg/mL) | Aggregation Rate at 40°C (%/month) | Viscosity at 150 mg/mL (cP) | Colloidal kᴅ Value | Recommended Use Case |
|---|---|---|---|---|
| Histidine, Sucrose, PS-20 (pH 6.0) | 0.5 | 12 | +15 | Long-term liquid storage |
| Succinate, NaCl (pH 5.0) | 0.8 | 8 | +5 | High-concentration SC |
| Phosphate, Arginine, PS-80 (pH 7.4) | 1.2 | 25 | -8 | Early-stage development |
| Acetate, Trehalose (pH 5.5) | 0.3 | 15 | +20 | Lyophilization candidate |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Fusion Protein Formulation & Stability Studies
| Item/Category | Example Product/Description | Primary Function in Research |
|---|---|---|
| Stabilizing Excipients | Sucrose, Trehalose, Sorbitol | Bulking agents and cryoprotectants; provide thermodynamic stabilization via preferential exclusion. |
| Surfactants | Polysorbate 20 (PS-20), Polysorbate 80 (PS-80) | Minimize aggregation induced by interfacial shear and surface adsorption. |
| Amino Acid Stabilizers | L-Arginine HCl, L-Histidine, Glycine | Modulate colloidal stability, reduce viscosity, and act as buffering agents. |
| Oxidation Inhibitors | Methionine, EDTA, Sodium Thiosulfate | Scavenge reactive oxygen species or chelate metal catalysts to prevent oxidation. |
| Analytical Standards | NISTmAb, reduced/alkylated mAb controls | System suitability standards for chromatography and electrophoresis assays. |
| FcRn Binding Reagents | Recombinant hFcRn (with β-2 microglobulin), SPR chips (CM5) | Essential for quantifying the critical pharmacokinetic driver interaction. |
| Forced Degradation Kits | Commercial peroxide, light stress kits | Provide standardized reagents for controlled comparative degradation studies. |
Pathways and Workflows
Title: Stability Testing Workflow for Fc-Fusion Proteins
Title: FcRn Recycling Pathway and Stability Impact
Addressing Immunogenicity Risks of Novel Fc or Albumin-Binding Motifs
Therapeutic proteins, peptides, and oligonucleotides often suffer from rapid clearance, necessitating frequent administration. Leveraging the neonatal Fc receptor (FcRn) recycling pathway—by engineering molecules to bind either directly to FcRn or to endogenous albumin which itself binds FcRn—has become a cornerstone strategy for extending plasma half-life. While successful, the introduction of de novo Fc or albumin-binding motifs (ABMs) presents a significant immunogenicity risk. These novel protein sequences or chemical linkers can be recognized as "non-self" by the host immune system, potentially leading to the development of anti-drug antibodies (ADAs). ADA formation can neutralize drug efficacy, alter pharmacokinetics (PK), and, in severe cases, cause adverse events. This technical guide addresses the systematic evaluation and mitigation of immunogenicity risks associated with novel binding motifs within the context of FcRn-mediated half-life extension research.
Immunogenicity Risk Factors for Novel Binding Domains
The immunogenic potential of an engineered therapeutic is influenced by multiple factors intrinsic to the novel motif and the final construct. Key risk factors include:
- Sequence Foreignness: The degree of deviation from human germline sequences. Fully non-human sequences (e.g., bacterial domains) carry higher risk than engineered human variants.
- Aggregation Propensity: Novel motifs can destabilize the protein or create interfacial regions prone to aggregation. Protein aggregates are potent immunogenic triggers.
- Post-Translational Modifications (PTMs): Introduction of new glycosylation sites or chemical modifications (e.g., from site-specific conjugation) can create neoepitopes.
- Processing & Presentation: The ability of antigen-presenting cells (APCs) to process the motif and present novel T-cell epitopes on MHC class II molecules is a critical determinant.
- Structural Context & Flexibility: Epitopes presented in flexible, solvent-exposed loops are more immunogenic than those in rigid, buried structures.
Integrated Pre-Clinical Immunogenicity Risk Assessment: Protocols & Data
A tiered experimental approach is required to assess risk prior to clinical studies. Key methodologies are outlined below.
In SilicoT-cell Epitope Prediction
Protocol: Utilize robust algorithms (e.g., NetMHCIIpan, IEDB tools) to scan the amino acid sequence of the novel binding motif and its junctions with the parent molecule.
- Input the FASTA sequence of the full construct.
- Select a panel of common human MHC class II alleles (DRB1*01:01, *03:01, *04:01, *07:01, *15:01).
- Run prediction for peptides of 15-mer length, sliding window of 1.
- Identify core 9-mer binding motifs with predicted IC50 < 1000 nM or percentile rank < 10%.
- Flag sequences containing multiple overlapping or promiscuous predicted epitopes.
Ex VivoT-cell Activation Assays (Human Peripheral Blood Mononuclear Cell [PBMC] Assay)
Protocol: This assay measures the potential of a protein to activate naïve T-cells from a diverse human donor pool.
- Isolate PBMCs from 50+ healthy human donors (cryopreserved).
- Plate cells in 96-well U-bottom plates.
- Treat with the following, in triplicate:
- Test article (novel binding motif construct, 10 µg/mL).
- Negative control (wild-type protein without motif, 10 µg/mL).
- Positive control (anti-CD3/CD28 beads or keyhole limpet hemocyanin [KLH]).
- Vehicle control.
- Culture for 7 days.
- Measure T-cell activation by IFN-γ ELISpot or flow cytometry for activation markers (CD25+, CD134+).
- A response is considered positive if stimulation index (SI = test counts / vehicle counts) > 2 and statistically significant (p < 0.05) in >5% of donors.
ADA Assessment in Transgenic Mouse Models
Protocol: Use mice engineered to express human MHC class II alleles or human immune system components (e.g., HLA-DR transgenic mice).
- Formulate test and control articles in appropriate buffer.
- Administer 3-5 subcutaneous or intravenous doses to mice (n=8-10/group) over 4 weeks (e.g., Days 1, 15, 29).
- Collect serum samples pre-dose and 7-10 days after the final dose.
- Screen for total ADA using a validated bridging electrochemiluminescence (ECL) assay:
- Biotinylate and sulfo-tag the drug.
- Incubate diluted serum samples with a mixture of both labeled reagents.
- Capture on a streptavidin plate and read on an MSD instrument.
- Report signal as ECL units. Positivity is determined by a cut point based on negative control serum + 3 standard deviations.
Table 1: Example Immunogenicity Assessment Data for Candidate ABM-Fusion Proteins
| Construct | Predicted Promiscuous T-cell Epitopes | PBMC Assay (% Responders, Mean SI) | HLA-DR3 Tg Mouse (% ADA Positive) | PK Impact in ADA+ Mice (AUC reduction) |
|---|---|---|---|---|
| WT Protein | 0 | 2%, 1.5 | 0% | N/A |
| ABM-Variant A | 3 | 18%, 4.2 | 75% | 92% |
| ABM-Variant B | 1 | 5%, 2.1 | 12% | 15% |
| Fc-Mutant X | 2 | 12%, 3.8 | 50% | 70% |
Mitigation Strategies: Deimmunizing Novel Binding Motifs
When a risk is identified, several engineering strategies can be employed.
- Epitope Deletion/Substitution: Replace key residues in predicted T-cell epitopes while maintaining binding affinity. Alanine scanning or homology grafting from human proteins can guide this.
- Humanization: If the motif is derived from a non-human source, graft its functional paratope onto a closest human scaffold.
- Masking: Conjugate the motif via site-specific chemistry to polysialic acid (PSA) or polyethylene glycol (PEG) to sterically shield potential epitopes.
- Tolerization: Co-administer with immunosuppressive regimens (e.g., rapamycin) or fuse with immune-tolerant domains (e.g., human serum albumin itself). This is higher-risk and less predictable.
Table 2: Key Research Reagent Solutions for Immunogenicity Assessment
| Reagent / Material | Function in Assessment | Example Vendor / Source |
|---|---|---|
| Cryopreserved Human PBMCs (Multi-Donor) | Provides a diverse human immune repertoire for ex vivo T-cell assays. | STEMCELL Technologies, AllCells |
| HLA-Transgenic Mice (e.g., HLA-DR3) | In vivo model with human-relevant T-cell epitope presentation. | Taconic Biosciences, Jackson Labs |
| MSD ECL Assay Plates & SULFO-TAG NHS-Ester | High-sensitivity platform for developing ADA and PK assays. | Meso Scale Discovery |
| IFN-γ ELISpot Kit (Human) | Quantifies antigen-specific T-cell responses from PBMC assays. | Mabtech, R&D Systems |
| NetMHCIIpan Prediction Server | In silico prediction of peptide binding to a wide range of HLA alleles. | DTU Health Tech (www.cbs.dtu.dk/services/NetMHCIIpan) |
| Size-Exclusion Chromatography (SEC) Columns | Critical for analyzing aggregation propensity (HMW species) of novel constructs. | Cytiva (Superdex series), Waters (BEH series) |
| Surface Plasmon Resonance (SPR) Chip with Captured FcRn | Validates that deimmunization mutations do not abrogate FcRn or albumin binding affinity. | Cytiva (Series S CMS chip, human FcRn protein) |
Signaling Pathways in Immunogenicity: From Motif to ADA Response
Diagram 1: Cellular Pathway from Immunogenic Motif to ADA Impact
Integrated Immunogenicity Risk Assessment Workflow
Diagram 2: Sequential Immunogenicity Risk Assessment Workflow
The successful development of next-generation half-life extension technologies based on novel Fc or albumin-binding motifs is critically dependent on proactive immunogenicity risk management. An integrated strategy—combining predictive in silico tools, sensitive ex vivo human immune assays, and relevant in vivo models—enables early identification and mitigation of immunogenic sequences. By systematically applying these principles within the framework of FcRn biology, researchers can engineer safer, more effective biotherapeutics with optimized pharmacokinetic profiles and a reduced likelihood of clinical immunogenicity.
Comparative Analysis and Validation of FcRn Technologies: Bench to Bedside
This whitepaper provides a detailed technical comparison of major half-life extension platforms, framed within the ongoing research on neonatal Fc receptor (FcRn)-mediated recycling. The central thesis posits that understanding and exploiting the FcRn-albumin salvage pathway is the cornerstone of modern biologics engineering, with each platform representing a distinct strategy to hijack this endogenous mechanism to reduce clearance and improve therapeutic exposure.
Fc-Fusion
Proteins of interest are genetically fused to the Fc domain of an IgG. The Fc domain binds to FcRn in acidic endosomes, diverting the fusion protein from lysosomal degradation and facilitating its recycling to the cell surface and release back into circulation at neutral pH.
Albumin Fusion
The therapeutic protein is fused directly to human serum albumin (HSA) or an albumin-binding domain. HSA is a natural, high-affinity ligand for FcRn, leveraging the same protective recycling pathway as Fc fusions but with a distinct binding site on the receptor.
PEGylation
Covalent attachment of polyethylene glycol (PEG) polymers to a protein. PEGylation primarily extends half-life by increasing hydrodynamic radius (reducing renal filtration) and shielding the protein from proteolytic enzymes and immune recognition. It is largely independent of FcRn recycling.
Other/Next-Generation Platforms
Includes XTEN and other unstructured polypeptide fusions, Fc-mimetic peptides (e.g., Abdegs), and engineered albumin variants. These often aim to fine-tune FcRn affinity or create alternative stealth modalities.
Quantitative Comparison of Key Parameters
Table 1: Platform Characteristics & Performance Data
| Parameter | Fc-Fusion | Albumin Fusion | PEGylation | Other (e.g., XTEN) |
|---|---|---|---|---|
| Typical Half-life Extension | ~2-3 weeks (IgG-like) | ~2-3 weeks (Albumin-like) | Highly variable (hours to weeks) | ~1-2 weeks |
| Primary Mechanism | FcRn-mediated recycling | FcRn-mediated recycling | Increased size, reduced clearance | Mixed (size, stability, sometimes FcRn) |
| Molecular Weight Increase | ~55 kDa (Fc dimer) | ~67 kDa (HSA) | 5-60 kDa per PEG chain | 20-100 kDa (polypeptide) |
| Potential Immunogenicity | Low (human sequence) | Low (human sequence) | Moderate (anti-PEG antibodies) | Variable (depends on sequence) |
| Impact on Bioactivity | Can be significant (sterics, FcγR binding) | Can be significant (large carrier) | Often reduced (steric shielding) | Designed to minimize impact |
| Production Complexity | High (complex protein expression/purification) | High (complex protein expression/purification) | Moderate (conjugation chemistry) | Moderate to High |
| Key Commercial Example | Etanercept (Enbrel) | Albiglutide (Tanzeum) | Pegfilgrastim (Neulasta) | Efpeglenatide (GLP-1-XTEN) |
Table 2: FcRn Binding Affinity & Pharmacokinetic Data
| Platform / Molecule | FcRn Binding Affinity (KD, nM) | Reported Terminal Half-life (Human) | Dosing Frequency |
|---|---|---|---|
| Native IgG1 | 50-500 (pH 6.0) | ~21 days | Weekly to monthly |
| Engineered Fc (YTE mutant) | ~10 (pH 6.0) | ~85-100 days | Extended interval |
| Human Serum Albumin (HSA) | 300-800 (pH 6.0) | ~19 days | Weekly to biweekly |
| PEGylated IFN-α2a | No binding | ~80 hours | Weekly |
| XTEN-GLP-1 Fusion | No binding | ~6-7 days | Weekly |
Detailed Experimental Protocols
Protocol: Surface Plasmon Resonance (SPR) for FcRn Binding Affinity
Objective: Quantify the binding affinity of an Fc-fusion or albumin-fusion protein to human FcRn at endosomal pH. Materials: Biacore T200/8K series SPR instrument, CMS sensor chip, recombinant human FcRn, sodium acetate (pH 5.5), HBS-EP+ buffer, amine coupling reagents (EDC/NHS), glycine-HCl (pH 2.0). Method:
- Immobilization: Dilute FcRn in 10 mM sodium acetate (pH 5.5) to 10 µg/mL. Activate the CMS chip surface with a 1:1 mixture of 0.4 M EDC and 0.1 M NHS for 7 minutes. Inject FcRn solution for 7 minutes to achieve ~2000 RU. Deactivate with 1 M ethanolamine-HCl (pH 8.5) for 7 minutes.
- Kinetic Analysis: Dilute Fc-fusion/albumin-fusion analytes in HBS-EP+ buffer, pH 6.0. Use a multi-cycle kinetics approach. Inject analyte concentrations (e.g., 0, 6.25, 12.5, 25, 50, 100 nM) at 30 µL/min for 180s association, followed by 600s dissociation in pH 6.0 buffer.
- Regeneration: After each cycle, regenerate the FcRn surface with two 30-second pulses of HBS-EP+ buffer, pH 7.4.
- Data Processing: Double-reference the data (reference surface & buffer blank). Fit the sensorgrams to a 1:1 Langmuir binding model using Biacore Evaluation Software to derive ka (association rate), kd (dissociation rate), and KD (kd/ka).
Protocol: Pharmacokinetic Study in Humanized FcRn Mouse Model
Objective: Evaluate the serum half-life of a novel half-life extension construct in vivo. Materials: C57BL/6-FcRn-/-hFcRn transgenic mice (B6.mFcRn-/-.hFcRn Tg32), test article, control IgG, PBS, EDTA microtainers, ELISA kits for detection. Method:
- Dosing: Randomly assign mice (n=5/group) to receive a single 5 mg/kg intravenous bolus of the test article or control via tail vein.
- Serial Bleeding: Collect blood samples (50-100 µL) via submandibular or retro-orbital puncture at time points: 5 minutes (Cmax), 6h, 24h, 48h, 96h, 168h, 240h, and 336h post-dose.
- Sample Processing: Immediately centrifuge blood samples at 5,000xg for 10 minutes. Collect plasma and store at -80°C until analysis.
- Bioanalysis: Quantify test article concentrations in plasma using a validated sandwich ELISA (e.g., anti-therapeutic capture, species-specific HRP-conjugated detection).
- PK Analysis: Use non-compartmental analysis (NCA) in software like Phoenix WinNonlin to calculate key parameters: AUC0-inf, Cmax, clearance (CL), volume of distribution (Vd), and terminal half-life (t1/2).
Signaling & Workflow Visualizations
Diagram Title: FcRn Recycling Pathway for Half-Life Extension
Diagram Title: Primary Mechanisms of Action by Platform
Diagram Title: PK Study Workflow in hFcRn Mice
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for FcRn-Albumin Half-Life Research
| Item/Catalog (Example) | Function & Application |
|---|---|
| Recombinant Human FcRn Protein (R&D Systems, 8638-FC) | Key reagent for in vitro binding studies (SPR, ELISA). Provides the target for affinity measurements at endosomal pH. |
| hFcRn Transgenic Mouse Model (Taconic, B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr) | Gold-standard in vivo model for predicting human PK of FcRn-dependent therapeutics. Expresses human FcRn with human tissue expression pattern. |
| Anti-Human IgG Fc-HRP Conjugate (Jackson ImmunoResearch, 109-035-098) | Critical detection antibody for quantifying Fc-fusion or IgG concentrations in bioanalytical ELISAs from plasma/serum samples. |
| Biacore Series S CM5 Sensor Chip (Cytiva, BR100530) | Gold standard SPR chip for immobilizing FcRn to perform real-time, label-free kinetic binding analysis of test molecules. |
| pH 6.0 Phosphate Assay Buffer (Thermo Fisher, 28372) | Mimics the acidic environment of the endosome for conducting FcRn binding assays in vitro. |
| Human Serum Albumin, Fatty Acid Free (Sigma, A3782) | Used as a standard, carrier, or competitor in assays involving albumin fusion proteins or studying albumin-FcRn interactions. |
| Site-Specific PEGylation Kits (Thermo Fisher, 22341) | Enable controlled, reproducible conjugation of PEG polymers to amines or thiols on proteins for generating PEGylated comparators. |
| Protein A/G Purification Resin (Thermo Fisher, 20399) | Standard affinity resin for purifying Fc-fusion proteins and antibodies from cell culture supernatants during construct development. |
| Meso Scale Discovery (MSD) Streptavidin Plates (Meso Scale Diagnostics, L15SA-1) | Used for developing sensitive immunoassays with low sample volumes for PK/TK studies, often offering broader dynamic range than ELISA. |
This guide explores preclinical models critical for predicting the pharmacokinetics (PK) of therapeutic proteins, with a specific focus on strategies for extending half-life via the neonatal Fc receptor (FcRn) salvage pathway. The research thesis centers on engineering biologics—such as antibodies, Fc-fusion proteins, and albumin-based therapeutics—to have enhanced affinity for FcRn at acidic pH, thereby evading lysosomal degradation and recycling back into systemic circulation. Accurate prediction of human PK from preclinical data is paramount, requiring robust in vitro assays and sophisticated in vivo transgenic mouse systems that recapitulate human FcRn biology.
In Vitro Assays for FcRn Binding and Prediction
In vitro assays provide high-throughput, mechanistic data on FcRn interaction, forming the foundation for candidate selection.
Key Experimental Protocols
Protocol: Surface Plasmon Resonance (SPR) for pH-Dependent FcRn Binding Kinetics
- Immobilization: Human or mouse FcRn is immobilized on a CMS sensor chip via amine coupling.
- Running Buffer: Use buffers at two pH conditions: pH 6.0 (mimicking endosomal environment) and pH 7.4 (mimicking bloodstream).
- Analyte Injection: Serial dilutions of the therapeutic candidate (e.g., monoclonal antibody) are injected over the chip surface at pH 6.0.
- Dissociation & Regeneration: Dissociation is monitored in pH 6.0 buffer, followed by a switch to pH 7.4 buffer to monitor the release. The surface is regenerated with a mild acidic or basic pulse (e.g., 10 mM Glycine pH 1.5).
- Data Analysis: Kinetic rates (ka, kd) and affinity (KD) at pH 6.0 are calculated using a 1:1 Langmuir binding model. The critical parameter is a high KD at pH 7.4, indicating release.
Protocol: Cell-Based Transcytosis Assay
- Cell Culture: Grow a polarized monolayer of human endothelial or epithelial cells (e.g., MDCK or HUVEC) expressing human FcRn on a transwell insert.
- Apical Loading: Add the test therapeutic (fluorophore- or radio-label) to the apical compartment at pH 6.0.
- Incubation: Allow transport for 2-4 hours.
- Quantification: Measure the amount of therapeutic that appears in the basolateral compartment (pH 7.4) via fluorescence or scintillation counting. An isotype control with no FcRn binding is used for comparison.
- Data Expression: Calculate the apparent permeability (Papp) and fold-increase over control.
Quantitative Data from In Vitro Assays
Table 1: Representative In Vitro FcRn Binding and Transcytosis Data for Engineered Antibodies
| Therapeutic Candidate | SPR KD at pH 6.0 (nM) | SPR KD at pH 7.4 (μM) | In Vitro Transcytosis (Fold over WT) | Predicted Human Half-life Extension |
|---|---|---|---|---|
| Wild-type IgG1 | 500 - 800 | > 50 | 1.0 (baseline) | 1x (~21 days) |
| YTE Mutant (M252Y/S254T/T256E) | 50 - 100 | > 100 | 1.8 - 2.5 | ~2.5x |
| LS Mutant (M428L/N434S) | 10 - 30 | > 100 | 3.0 - 4.0 | ~3x - 4x |
| Albumin | 300 - 500 | > 50 | 2.0 - 3.0 | ~19 days |
The Scientist's Toolkit: Key Reagents for In Vitro FcRn Studies
| Item | Function & Relevance |
|---|---|
| Recombinant Human FcRn (soluble) | Essential for SPR, ELISA, and affinity measurements. |
| Biacore Series S Sensor Chip CM5 | Gold-standard for label-free kinetic analysis via SPR. |
| pH-Specific Assay Buffers (pH 6.0 & 7.4) | Crucial for mimicking physiological conditions of endosome and blood. |
| MDCK-II cells stably expressing hFcRn | Preferred polarized cell line for transcytosis and recycling assays. |
| Anti-human Fc HRP Conjugate | For detection of human IgG in ELISA and cell-based assay supernatants. |
| 96-well HTS Transwell Plates | Enable high-throughput screening of permeability and transport. |
Diagram Title: FcRn-Mediated Recycling Pathway for Half-life Extension
Transgenic Mouse Systems for In Vivo PK Prediction
Wild-type mice are poor predictors for human PK of Fc-engineered proteins due to species specificity in FcRn interaction. Transgenic mouse models expressing human FcRn (hFcRn) are indispensable.
Key Mouse Models and Generation Protocol
Protocol: Generating and Validating a Human FcRn Transgenic Mouse Model (e.g., B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr/DcrJ)
- Model Design: The mouse Fcgr gene is knocked out to eliminate mouse FcRn. A transgene containing the human FCGRT gene (and often human β2-microglobulin) is inserted.
- Breeding & Genotyping: Maintain on an immunodeficient (e.g., NSG) or immunocompetent background. Regular genotyping via PCR for the hFcRn transgene and absence of mFcRn is required.
- Validation: Validate by measuring the clearance of a known human IgG versus mouse IgG. Human IgG should exhibit prolonged half-life comparable to humans, while mouse IgG clears rapidly.
In Vivo PK Study Protocol
Protocol: Single-Dose PK Study in hFcRn Tg Mice
- Dosing: Administer a single intravenous (IV) bolus of the test therapeutic (e.g., 5 mg/kg) to groups of hFcRn Tg mice (n=3-5).
- Serial Bleeding: Collect blood samples at multiple time points (e.g., 5 min, 6h, 24h, 3d, 7d, 14d, 21d).
- Bioanalysis: Quantify serum concentrations using a specific ELISA (e.g., anti-human Fc capture).
- PK Analysis: Fit concentration-time data using non-compartmental analysis (NCA) to determine key parameters: terminal half-life (t1/2), clearance (CL), and area under the curve (AUC).
Table 2: Comparative PK Parameters in Different Mouse Models
| Mouse Model | FcRn Expression | Human IgG1 t1/2 (days) | Engineered (LS) IgG t1/2 (days) | Predictive Value for Human PK |
|---|---|---|---|---|
| Wild-type (C57BL/6) | Mouse only | 6 - 8 | 6 - 9 | Low |
| Fcgr -/- (KO) | None | 2 - 3 | 2 - 3 | Low (Negative control) |
| hFcRn Tg32 (Hemizygous) | Human only | 18 - 22 | 25 - 32 | High |
| hFcRn Tg276 (Homozygous) | Human only | 20 - 24 | 28 - 35 | High |
Diagram Title: Integrated Preclinical PK Prediction Workflow
Integration and Correlation to Human PK
The most predictive approach integrates data from both in vitro and in vivo models. A strong correlation between in vitro FcRn affinity at pH 6.0, in vitro transcytosis efficiency, and in vivo half-life extension in hFcRn Tg mice has been established. Multi-parameter regression models using this integrated data set can provide quantitative predictions of human CL and t1/2 with significantly greater accuracy than any single model.
Within the thesis of FcRn-targeted half-life extension, the synergistic use of mechanistic in vitro assays and humanized in vivo transgenic models forms a robust, predictive framework for selecting and advancing next-generation biologics with optimized pharmacokinetic profiles. This guide provides the essential methodologies and data frameworks necessary for researchers to implement this critical path in drug development.
Within the strategic framework of advancing FcRn (neonatal Fc receptor) antagonist therapies for albumin half-life extension, translational pharmacokinetic/pharmacodynamic (PK/PD) modeling is the critical quantitative discipline that converts in vitro and preclinical in vivo data into rational First-in-Human (FIH) dose selections. This guide details the systematic methodology for building robust translational PK/PD models, ensuring efficient and de-risked transition from bench to bedside.
The PK/PD Modeling Workflow: From Mechanism to Human Projection
The core translational workflow integrates data across biological scales, anchored by the mechanistic understanding of FcRn-mediated IgG/albumin recycling.
Diagram 1: Translational PK/PD Modeling Workflow
Core Mechanistic Model: FcRn-Albumin Interaction
The model structure is founded on the established physiology of FcRn-mediated protection of albumin from lysosomal degradation.
Diagram 2: FcRn-Mediated Albumin Recycling & Antagonist Action
Key Quantitative Parameters & Data Tables
Translational models require precise parameter estimation from preclinical studies.
Table 1: CriticalIn Vitro& Preclinical PK Parameters for FcRn Antagonists
| Parameter | Symbol (Typical) | Experimental Source | Relevance to Human Projection |
|---|---|---|---|
| Target Affinity (Human FcRn) | KD | Surface Plasmon Resonance (SPR) | Primary driver of in vivo potency. |
| Plasma Clearance (CL) | CLrat, CLNHP | IV PK in rodents & NHPs | Allometrically scaled to predict human CL. |
| Volume of Distribution (Vd) | Vss | IV PK in rodents & NHPs | Predicts human Vss and loading dose. |
| In Vivo Potency (EC50) | EC50 | Albumin turnover study in transgenic mice (hFcRn) | Links antagonist concentration to PD effect. |
| Endogenous Albumin Half-Life | t1/2,alb | Radio-labeled albumin study | Defines baseline for % extension calculation. |
Table 2: Allometric Scaling Factors for Human Dose Projection
| Species | Body Weight (kg) | Scaling Exponent (CL) | Scaling Exponent (Vd) | Typical Prediction Outcome |
|---|---|---|---|---|
| Mouse | 0.025 | 0.75 | 1.0 | Often under-predicts human CL; used qualitatively. |
| Rat | 0.25 | 0.75 | 1.0 | Improved prediction; requires protein binding correction. |
| NHP (Cynomolgus) | 3.0 | 0.75-0.85 | ~1.0 | Most reliable for mAbs/Fc-fusions; often used as primary. |
Detailed Experimental Protocols
Protocol 1:In VivoPharmacodynamic Assessment in Humanized FcRn Mouse Model
Objective: Quantify the relationship between antagonist plasma concentration and the increase in endogenous albumin half-life.
Materials:
- Human FcRn transgenic mouse model (e.g., B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr/DcrJ).
- Test FcRn antagonist (formulated for subcutaneous/IV dosing).
- BrdU or stable isotope label (e.g., ^13^C6-Phe) for albumin turnover measurement.
- LC-MS/MS platform for pharmacokinetic and endogenous albumin analysis.
Method:
- Dosing & Sampling: Administer a single SC dose of antagonist at multiple levels (e.g., 0, 1, 3, 10, 30 mg/kg). Collect serial plasma samples over 21-28 days.
- Albumin Turnover Measurement: At Day 1 post-dosing, inject BrdU or administer a ^13^C6-Phe-labeled diet. Track the incorporation and decay of labeled albumin in plasma via LC-MS/MS.
- Data Analysis: Fit labeled albumin decay curves to a one-compartment model to estimate apparent albumin half-life (t1/2,alb) for each dose group. Plot t1/2,alb against the corresponding average antagonist concentration (Cavg) or trough level.
- Model Fitting: Fit the exposure-response data to an Emax model:
Effect = E0 + (Emax * C) / (EC50 + C), where Effect is t1/2,alb or fold-change from baseline.
Protocol 2: Population PK/PD Simulation for FIH Dose Selection
Objective: To predict the clinical dose required to achieve a target albumin half-life extension in a human population.
Method:
- Base Model Development: Integrate in vitro KD, preclinical PK, and the in vivo EC50 into a mechanistic PK/PD model using software (e.g., NONMEM, Monolix, or Phoenix).
- Allometric Scaling: Scale animal CL and Vss to human using the allometric equation:
Human Parameter = Animal Parameter * (Human Weight / Animal Weight)^Exponent. Typically, use NHP data with an exponent of 0.85 for CL and 1.0 for Vss. - Incorporating Variability: Introduce inter-individual variability (IIV) on key parameters (e.g., CL, V, EC50) based on preclinical variability or typical clinical values (~30% CV).
- Virtual Trial Simulation: Simulate 1000 virtual subjects receiving various dose regimens (e.g., 5 mg/kg QW, 10 mg/kg Q2W). For each regimen, predict the steady-state distribution of albumin half-life extension.
- Dose Determination: Select the FIH dose range that achieves a target effect (e.g., >1.5-fold increase in albumin t1/2) in >80% of the virtual population while remaining below exposures associated with preclinical adverse effects.
The Scientist's Toolkit: Essential Research Reagents & Materials
| Item | Function & Relevance |
|---|---|
| Recombinant Human FcRn Protein | Critical for in vitro binding assays (SPR, ELISA) to determine antagonist affinity (KD) and pH dependency. |
| Human FcRn Transgenic Mouse Model | The gold-standard in vivo model for PD studies, as it expresses the human FcRn pathway, enabling relevant assessment of albumin half-life extension. |
| Stable Isotope-Labeled Amino Acids (^13^C6-Phenylalanine) | Enables precise, non-radioactive measurement of endogenous albumin synthesis and catabolism rates in vivo (kinetic PD readout). |
| Anti-Human Fc Domain (FcγRII/III null) Antibodies | Used in PK assays to specifically quantify therapeutic FcRn antagonist levels in complex biological matrices without interference. |
| Validated LC-MS/MS Platform | For simultaneous, high-sensitivity quantification of both the therapeutic antagonist and endogenous albumin kinetics in preclinical and clinical samples. |
| Industry-Standard PK/PD Modeling Software (e.g., NONMEM, Phoenix NLME) | Provides the computational environment for developing, validating, and simulating mechanistic translational models. |
Within the broader thesis of FcRn receptor-mediated albumin and IgG half-life extension research, a critical translational question persists: How predictive are preclinical in vitro and in vivo FcRn models of human pharmacokinetic (PK) outcomes? This in-depth review assesses the correlation between preclinical data and clinical results for therapeutics engineered for FcRn binding, focusing on quantitative PK parameters and the methodologies that generate them.
Key Preclinical Models and Their Clinical Correlation
In VitroBinding and Cellular Assays
These assays measure the pH-dependent binding kinetics of engineered proteins to FcRn, a fundamental prerequisite for extended half-life.
Experimental Protocol: Surface Plasmon Resonance (SPR) for FcRn Binding
- Immobilization: Recombinant human FcRn is immobilized on a CMS sensor chip via amine coupling.
- Running Buffer: HBS-EP buffer (pH 7.4) is used for association and initial dissociation.
- pH-Switch Dissociation: The analyte (e.g., Fc-fusion protein or albumin variant) is injected at pH 6.0 to mimic endosomal conditions, allowing binding. Dissociation is monitored first at pH 6.0, then switched to pH 7.4 to mimic recycling to the bloodstream, triggering release.
- Data Analysis: Kinetic rates (ka, kd) and affinity (KD) at both pH values are calculated using a 1:1 Langmuir binding model. The critical metric is the KD ratio (pH 7.4 / pH 6.0), where a higher ratio indicates stronger preferential binding at acidic pH.
Table 1: Correlation of In Vitro FcRn Affinity Ratios with Human Half-Life Extension
| Therapeutic Modality | Example Molecule | In Vitro KD Ratio (pH7.4/pH6.0) | Predicted Δ vs. Wild-Type | Observed Human t1/2 (days) | Observed Δ vs. Wild-Type |
|---|---|---|---|---|---|
| IgG1 Fc Variant | Efgartigimod* | ~0.8 (weak pH-dependent binding) | Moderate Increase | ~15-18 (IgG) | ~2-3x increase |
| Albumin Variant | ALB-1 (hypothetical) | >100 (highly pH-dependent) | Significant Increase | ~28-35 | ~4-5x increase |
| Wild-Type IgG | Natalizumab | ~1.0 (minimal pH-dependency) | Baseline | ~11 | Baseline |
| Wild-Type HSA | Human Serum Albumin | ~50 | Baseline | ~19 | Baseline |
Note: Efgartigimod is an Fc fragment engineered for increased FcRn affinity at pH 6.0 and reduced affinity at pH 7.4.
In VivoPharmacokinetic Models
Transgenic mice expressing human FcRn (hFcRn Tg) are the standard model, as mouse FcRn does not bind human IgG/Fc effectively.
Experimental Protocol: PK Study in hFcRn Transgenic Mice
- Animal Model: Use homozygous hFcRn Tg mice (e.g., B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr/DcrJ).
- Dosing: Administer a single intravenous (IV) bolus of the test article and a reference control (e.g., wild-type IgG) at 5-10 mg/kg.
- Sampling: Collect serial plasma/serum samples (e.g., 5 min, 6h, 24h, 3d, 7d, 14d, 21d post-dose).
- Bioanalysis: Quantify concentrations using a target-specific ELISA or LC-MS/MS method.
- PK Analysis: Perform non-compartmental analysis (NCA) to estimate terminal half-life (t1/2), clearance (CL), and area under the curve (AUC).
Table 2: Predictive Performance of hFcRn Mouse PK for Human Outcomes
| PK Parameter in Model | Typical Correlation with Human PK (R²) | Strengths | Key Limitations |
|---|---|---|---|
| Terminal Half-life (t1/2) | 0.65 - 0.85 | Good for rank-ordering variants. | Mouse t1/2 is shorter; absolute values don't translate directly. |
| Clearance (CL) | 0.70 - 0.90 | Strong correlation for FcRn-dependent CL. | Does not capture target-mediated drug disposition (TMDD) in humans. |
| Relative Exposure (AUCtest/AUCref) | 0.75 - 0.95 | Excellent for predicting the fold-improvement over wild-type. | Dependent on choice of appropriate reference molecule. |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for FcRn-PK Research
| Item | Function & Rationale |
|---|---|
| Recombinant hFcRn Protein | Purified soluble FcRn (often with β-2-microglobulin) is essential for in vitro binding assays (SPR, ELISA). Species-specific variants (cyno, mouse) are needed for cross-reactivity studies. |
| hFcRn Transgenic Mice | The definitive in vivo model for predicting human PK of Fc- or albumin-based therapeutics due to humanized FcRn expression patterns. |
| pH-Switch SPR Buffer Kits | Pre-formulated running buffers at precise pH 6.0 and 7.4 are critical for replicating endosomal and serum conditions in binding kinetics experiments. |
| Isotype Control Human IgG | Wild-type human IgG1 (e.g., from pooled sera) serves as the essential benchmark control in both in vitro affinity and in vivo PK studies. |
| Anti-Human Fc / Albumin ELISA Kits | For specific, sensitive quantification of therapeutic concentrations in complex biological matrices during PK studies. |
| PK Analysis Software | Tools like Phoenix WinNonlin are industry standard for performing non-compartmental analysis (NCA) to calculate critical PK parameters from concentration-time data. |
Visualizing Key Concepts and Workflows
Diagram 1: Translational PK Prediction Pathway
Diagram 2: FcRn-Mediated Recycling Mechanism
Preclinical FcRn models, particularly integrated in vitro affinity profiling and in vivo studies in hFcRn transgenic mice, provide a robust framework for predicting the rank-order and relative magnitude of human half-life extension. While they are highly effective for optimizing engineering strategies and selecting clinical candidates, absolute human PK parameters (e.g., exact clearance rate) require additional scaling factors. Future model refinement will integrate quantitative systems pharmacology (QSP) to account for patient-specific variables like FcRn expression levels and competitive endogenous IgG/albumin loads, further closing the translational gap in albumin half-life extension research.
The neonatal Fc receptor (FcRn) has been a cornerstone of half-life extension (HLE) strategies, recycling IgG and albumin to prolong therapeutic serum persistence. However, reliance on traditional FcRn engagement presents limitations, including target competition, dependency on specific pH cycles, and finite extension windows. This has catalyzed a paradigm shift towards novel, orthogonal HLE modalities that operate independently of FcRn. This guide details the core technologies defining this new competitive landscape.
Core Technology Classes & Quantitative Comparison
The following technologies represent the primary emerging classes. Their key characteristics are summarized in Table 1.
Table 1: Comparative Analysis of Next-Generation HLE Technologies
| Technology Class | Mechanistic Basis | Key Structural Motif | Reported Half-Life Extension (vs. Unmodified) | Primary Advantages | Key Challenges |
|---|---|---|---|---|---|
| XTEN Polymers | Unstructured, hydrophilic polypeptide chain increasing hydrodynamic radius. | Recombinantly fused polypeptide (e.g., 864 AA). | ~100-200 fold increase in rodents (~6 days terminal t½). | Protease-resistant, biodegradable, low immunogenicity. | Manufacturing scale-up, potential viscosity. |
| ELP Polymers | Thermally responsive polypeptide forming reversible aggregates above Tt. | Recombinantly fused (VPGXG)n repeats. | Up to 20-40 fold increase in mice (~2 days terminal t½). | Tunable pharmacokinetics via Tt, simple purification. | Requires careful Tt engineering for in vivo behavior. |
| HESylation | Conjugation to Hydroxyethyl Starch (HES). | Covalent conjugation to HES polymer (various MW). | 5-15 fold increase in preclinical models. | Well-established, safe excipient history, scalable. | Potential for starch accumulation, polydispersity. |
| Albu-Binding DARPin | High-affinity, non-covalent binding to endogenous albumin. | Engineered DARPin domain (14-19 kDa). | Matches albumin half-life (~19 days in humans). | Modular, non-covalent, does not alter albumin function. | Risk of displacement by endogenous ligands. |
| Protease-Resistant Backbones | De novo design of proteins resilient to proteolytic cleavage. | Engineered sequences with substituted cleavage sites. | Variable; 5-50 fold increases demonstrated. | Directly addresses a primary clearance pathway. | Requires extensive screening and validation per scaffold. |
Detailed Experimental Protocols
Protocol 1: Evaluating HLE of XTEN-Fusion ProteinsIn Vivo
Objective: Determine pharmacokinetic (PK) parameters of an XTEN-modified therapeutic in a murine model.
Materials:
- Purified XTEN-fusion protein and unmodified control.
- C57BL/6 mice (n=5-6 per group).
- Microtainer serum separator tubes.
- ELISA kit specific for the therapeutic domain.
Methodology:
- Dosing & Sampling: Administer a single 2 mg/kg intravenous (IV) bolus to mice. Collect blood samples (e.g., via retro-orbital or submandibular bleed) at pre-dose, 5 min, 30 min, 2h, 8h, 24h, 48h, 96h, 144h, and 192h post-dose.
- Sample Processing: Allow blood to clot, centrifuge at 10,000 x g for 10 min, collect serum, and store at -80°C.
- Bioanalysis: Quantify serum concentrations using a validated sandwich ELISA. Coat plates with a capture antibody against the therapeutic domain. Use a biotinylated detection antibody against a different epitope, followed by streptavidin-HRP.
- PK Analysis: Fit serum concentration-time data using non-compartmental analysis (NCA) software (e.g., Phoenix WinNonlin) to calculate key parameters: terminal half-life (t½), area under the curve (AUC), and clearance (CL).
Protocol 2: Assessing Albumin-Binding Affinity of DARPin Agents via Surface Plasmon Resonance (SPR)
Objective: Measure the binding kinetics (KD, ka, kd) of an Albumin-Binding DARPin to human serum albumin (HSA).
Materials:
- Biacore T200 SPR instrument.
- CMS Series S sensor chip.
- HSA (fatty acid-free).
- DARPin analyte in series of concentrations (0.5 nM to 100 nM).
- Running Buffer: HBS-EP+ (10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% v/v Surfactant P20, pH 7.4).
- Regeneration Solution: 10 mM Glycine-HCl, pH 2.0.
Methodology:
- Immobilization: Dilute HSA to 20 µg/mL in 10 mM sodium acetate, pH 4.5. Activate the CM5 chip surface with a 1:1 mixture of 0.4 M EDC and 0.1 M NHS. Inject HSA solution over one flow cell to achieve ~5000 RU immobilization. Deactivate remaining esters with 1 M ethanolamine-HCl, pH 8.5.
- Kinetic Run: Prime system with HBS-EP+. Inject DARPin analytes over the HSA and reference flow cells at a flow rate of 30 µL/min for 120s association, followed by 600s dissociation.
- Data Analysis: Subtract reference cell data. Fit the resulting sensorgrams to a 1:1 Langmuir binding model using the Biacore Evaluation Software to calculate association rate (ka), dissociation rate (kd), and equilibrium dissociation constant (KD = kd/ka).
Visualizing Key Pathways & Workflows
Diagram 1: HLE Tech Mechanisms vs FcRn
Diagram 2: SPR Workflow for Albumin Binders
The Scientist's Toolkit: Key Research Reagents & Materials
Table 2: Essential Reagents for Next-Gen HLE Research
| Reagent / Material | Supplier Examples | Primary Function in Research |
|---|---|---|
| Fatty Acid-Free Human Serum Albumin (HSA) | Sigma-Aldrich, Equitech-Bio | Critical for in vitro binding studies (SPR, ELISA) and as a carrier protein in assays to mimic physiological conditions. |
| XTEN or ELP Encoding Plasmids | Biotech vendors (e.g., Amunix), academic repositories. | Backbone vectors for the recombinant fusion and expression of polymer-modified biologics. |
| Hydroxyethyl Starch (HES) Conjugation Kits | Nanocs, Creative PEGWorks. | Provide activated HES polymers (e.g., HES-succinimidyl carbonate) for controlled, site-specific conjugation to protein therapeutics. |
| Anti-PEG / Anti-XTEN Antibodies | Academia-derived, custom from Abcam. | Essential for quantifying or detecting polymer-conjugated therapeutics in complex biological matrices during PK studies. |
| Protease Panels (e.g., Cathepsins B, D, L) | R&D Systems, BioVision. | Used to screen and validate the protease resistance of engineered protein scaffolds in high-throughput assays. |
| Surface Plasmon Resonance (SPR) Instrumentation | Cytiva (Biacore), Bruker. | Gold-standard for label-free, real-time analysis of binding kinetics between HLE agents and their targets (e.g., albumin). |
| Size-Exclusion Chromatography with MALS (SEC-MALS) | Wyatt Technology, Agilent. | Determines the absolute molecular weight and hydrodynamic radius of polymer-conjugated proteins, confirming conjugation and aggregation state. |
Conclusion
The strategic exploitation of the FcRn recycling pathway has revolutionized the design of long-acting biologics, moving from foundational understanding to a diverse engineering toolkit. Successful implementation requires a nuanced balance of optimized pH-dependent binding, thorough preclinical validation in relevant models, and careful navigation of developability hurdles. As the field advances, future directions include engineering greater FcRn specificity to avoid interference with endogenous IgG homeostasis, developing novel non-Fc/non-albumin scaffolds with tuned recycling, and applying these principles to modalities beyond proteins, such as oligonucleotides and peptides. This knowledge empowers researchers to develop next-generation therapeutics with improved patient compliance and clinical efficacy through optimized pharmacokinetic profiles.