ESCRT Machinery: The Cellular Repair System for Lipid Nanoparticle Membrane Damage in Drug Delivery
This article explores the crucial role of the Endosomal Sorting Complexes Required for Transport (ESCRT) pathway in repairing lipid nanoparticle (LNP)-induced endosomal membrane damage, a key bottleneck in nucleic acid...
ESCRT Machinery: The Cellular Repair System for Lipid Nanoparticle Membrane Damage in Drug Delivery
Abstract
This article explores the crucial role of the Endosomal Sorting Complexes Required for Transport (ESCRT) pathway in repairing lipid nanoparticle (LNP)-induced endosomal membrane damage, a key bottleneck in nucleic acid delivery. Targeted at researchers and drug development professionals, it provides a foundational understanding of the mechanism, methodologies for studying ESCRT-LNP interactions, strategies to troubleshoot and optimize delivery efficiency by leveraging this pathway, and comparative analyses of ESCRT's role versus other cellular repair systems. The synthesis offers a roadmap for designing next-generation LNPs that actively engage cellular repair machinery to enhance therapeutic efficacy.
Unpacking the ESCRT Pathway: How Cells Detect and Repair LNP-Induced Membrane Damage
Lipid nanoparticles (LNPs) have emerged as the predominant non-viral delivery system for nucleic acid therapeutics, including mRNA vaccines and siRNA drugs. The primary intracellular barrier to efficient delivery is endosomal entrapment, where a significant majority of internalized LNPs fail to escape the endocytic pathway and are trafficked to lysosomes for degradation. Concurrently, the mechanism of endosomal escape—typically involving pH-dependent membrane disruption—poses a risk of damaging the limiting membrane of the endosome itself. This damage, if not properly managed by the cell, can compromise endosomal function and contribute to cytotoxicity. This guide frames the endosomal escape challenge within the context of cellular repair mechanisms, specifically the Endosomal Sorting Complex Required for Transport (ESCRT) pathway, which is recruited to repair membrane damage and may inadvertently limit LNP efficacy.
Mechanisms of Endosomal Entrapment and Escape
The Endocytic Journey
Following endocytosis, LNPs are encapsulated within early endosomes, which mature into late endosomes and ultimately fuse with lysosomes. The acidic pH of the endosomal lumen (progressing from ~6.5 in early endosomes to ~5.0 in late endosomes and lysosomes) is a critical trigger for LNP escape.
Membrane Disruption Strategies
Most ionizable cationic lipids used in LNPs are designed to become positively charged at low pH. This facilitates:
- Lipid Mixing: The ionizable lipid can adopt a cone-shaped morphology, promoting fusion between the LNP bilayer and the endosomal membrane.
- Pore Formation: The positive charge interacts with anionic phospholipids in the endosomal membrane, inducing hexagonal (HII) phase transitions or creating transient pores.
- Osmotic Lysis: Influx of protons and chloride ions increases osmotic pressure, swelling the endosome until it ruptures.
This disruptive activity creates discontinuities in the endosomal membrane, which the cell perceives as damage.
Quantitative Analysis of Delivery Efficiency
Despite optimization, escape efficiency remains low. Recent live-cell imaging and quantification studies reveal the following bottlenecks:
Table 1: Quantified Barriers in LNP-Mediated Nucleic Acid Delivery
| Delivery Stage | Typical Efficiency Range | Key Measurement Method |
|---|---|---|
| Cellular Uptake | 80-95% of applied dose | Flow cytometry (lipid dye quantification) |
| Endosomal Escape | 1-10% of internalized cargo | Gal8-mCherry assay / Split-GFP assays |
| Cytosolic Release | 0.5-5% of internalized cargo | Ribosome loading assays / Functional protein expression |
| Lysosomal Degradation | >90% of internalized LNPs | Co-localization analysis (LAMP1 staining) |
The Cellular Response: ESCRT-Mediated Membrane Repair
The ESCRT machinery is a conserved cellular system responsible for sealing small holes in lipid bilayers, including those on endosomes. When an LNP disrupts the endosomal membrane, calcium influx from the cytosol acts as a damage signal. This recruits early-acting ESCRT components (ALIX, TSG101) to the site of injury.
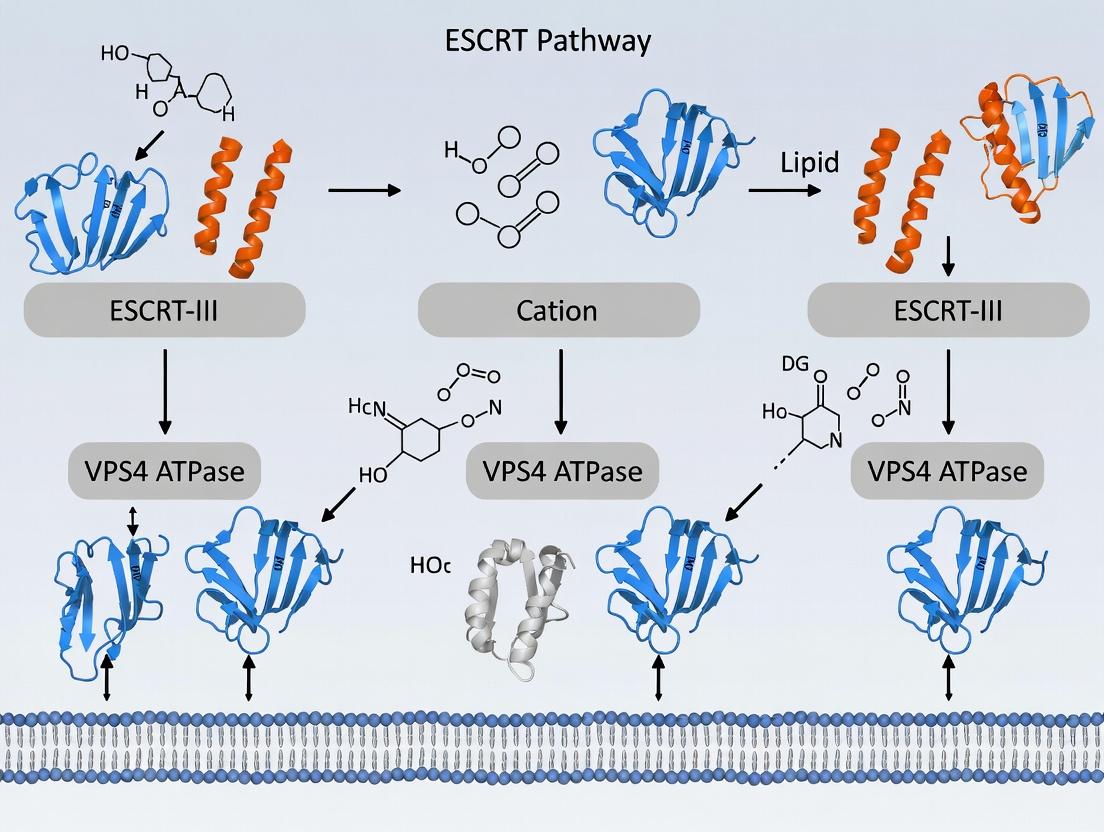
ESCRT Pathway for LNP-Induced Damage Repair:
Functional Consequences for LNP Delivery
The ESCRT-mediated repair response presents a fundamental trade-off:
- Cell Protection: Prevents endosomal leakage, lysosomal dysfunction, and cell death.
- Therapeutic Limitation: Rapidly reseals the membrane before the nucleic acid payload can fully escape, trapping it inside the endosome. This pathway is hypothesized to be a significant contributor to the low escape efficiencies quantified in Table 1.
Key Experimental Methodologies
Protocol: Galectin-8 (Gal8) Assay for Endosomal Damage
Gal8 is a cytosolic lectin that binds exposed glycans on damaged endosomes, serving as a high-resolution marker for LNP-induced membrane disruption.
Procedure:
- Cell Preparation: Seed HeLa or other relevant cells in an imaging-compatible 96-well plate.
- Transfection: Transfect cells with a plasmid encoding Gal8-mCherry 24h prior to LNP treatment.
- LNP Treatment: Treat cells with fluorescently labeled LNPs (e.g., DiO-labeled) at a standard dose.
- Fixation & Imaging: At defined time points (e.g., 1, 2, 4, 6h post-treatment), fix cells with 4% PFA. Perform immunofluorescence for LAMP1 (lysosomes) and nucleus (DAPI).
- Quantification: Acquire confocal images. Use image analysis software (e.g., CellProfiler) to calculate the Gal8 Punctuation Index: (Number of Gal8-positive puncta per cell) / (Number of LNP-positive puncta per cell). A higher index indicates greater membrane damage.
Protocol: Assessing ESCRT Dependence via siRNA Knockdown
This protocol tests the hypothesis that inhibiting ESCRT function will enhance LNP-mediated delivery by delaying membrane repair.
Procedure:
- Gene Knockdown: Transfect cells with siRNA targeting a critical ESCRT component (e.g., TSG101 or VPS4) or a non-targeting control (NTC) siRNA 72h before LNP treatment.
- Knockdown Validation: Harvest a subset of cells for western blot analysis to confirm protein knockdown.
- LNP Treatment & Readout: Treat remaining cells with LNPs encoding a luciferase or GFP reporter mRNA.
- Functional Assay:
- For luciferase: Lyse cells 24h post-treatment and measure luminescence. Normalize to total protein.
- For GFP: Analyze by flow cytometry to determine the percentage of GFP-positive cells and mean fluorescence intensity.
- Cytotoxicity Assay: Perform in parallel (e.g., LDH release) to ensure increased expression is not solely due to cell death from unrepaired damage.
Protocol: Co-localization Analysis for Endosomal Fate
Quantifies the proportion of LNPs that reach lysosomes versus those that cause damage/escape.
Procedure:
- Treat cells with fluorescent LNPs for a defined pulse (e.g., 30 min), then replace with fresh media.
- At chase time points (2, 4, 8, 24h), fix cells and immunostain for:
- EEA1 (Early Endosomes)
- Rab7 (Late Endosomes)
- LAMP1 (Lysosomes)
- Gal8 (Damage)
- Acquire high-resolution z-stack confocal images.
- Use co-localization plugins (e.g., JaCoP in ImageJ) to calculate Manders' overlap coefficient (MOC) between the LNP channel and each organelle/damage marker.
Table 2: Summary of Key Experimental Protocols
| Protocol Goal | Primary Readout | Key Controls | Technical Considerations |
|---|---|---|---|
| Measure Membrane Damage | Gal8 Punctuation Index | Untreated cells; cells treated with transfection reagent (e.g., Lipofectamine) as positive damage control. | Optimize Gal8-mCherry expression level to avoid background. |
| Test ESCRT Role | Reporter Expression (RLU or %GFP+) post-siRNA | Non-targeting siRNA; cytotoxicity assay. | Use a low, sub-optimal LNP dose to see enhancement clearly. |
| Track Intracellular Fate | Co-localization Coefficients (MOC) | Single-stained samples for spectral unmixing. | Use sequential scanning to avoid fluorophore bleed-through. |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for LNP Escape & ESCRT Research
| Reagent / Material | Supplier Examples | Primary Function in Experiments |
|---|---|---|
| Ionizable Cationic Lipids (e.g., DLin-MC3-DMA, SM-102) | MedChemExpress, Avanti Polar Lipids | Core component of LNP formulation; enables pH-dependent membrane disruption and endosomal escape. |
| Fluorescent Lipid Tracers (DiO, DiD, Rhodamine-PE) | Thermo Fisher, Avanti Polar Lipids | Incorporation into LNP bilayer allows visualization of cellular uptake and intracellular trafficking via microscopy/flow cytometry. |
| Gal8-mCherry Plasmid | Addgene (Plasmid #73359) | Expression construct for the definitive marker of endolysosomal membrane damage in live or fixed cells. |
| siRNA Pools (TSG101, VPS4A/B, ALIX, Non-Targeting) | Dharmacon, Sigma-Aldrich | Knockdown of specific ESCRT pathway components to probe their functional role in limiting LNP delivery. |
| Antibodies: LAMP1, EEA1, Rab7, TSG101 | Cell Signaling Technology, Abcam | Immunostaining to identify organelles and validate protein knockdown via western blot. |
| Luciferase Reporter mRNA | TriLink BioTechnologies | Encapsulated payload to quantitatively measure functional cytoplasmic delivery via luminescence assays. |
| Microfluidic Mixer (NanoAssemblr, HiPF) | Precision NanoSystems, Dolomite | Enables reproducible, scalable formulation of LNPs with precise size and encapsulation efficiency. |
Understanding endosomal entrapment requires moving beyond a passive view of the LNP and embracing the dynamic interplay between the nanoparticle and the cell's innate defense systems. The ESCRT pathway represents a sophisticated, rapid-response repair mechanism that actively counteracts LNP-mediated membrane disruption. Future research strategies aimed at enhancing LNP efficacy may involve:
- Temporal Control: Designing lipids that disrupt faster than ESCRT recruitment.
- Spatial Control: Targeting LNPs to endosomal sub-populations with lower ESCRT activity.
- Mechanistic Interference: Co-delivering transient ESCRT inhibitors alongside therapeutic payloads. A detailed understanding of the ESCRT-LNP interaction is therefore not merely biological curiosity but a critical path toward engineering the next generation of highly efficient delivery systems.
The Endosomal Sorting Complex Required for Transport (ESCRT) pathway is a universally conserved membrane remodeling machinery. In the context of lipid nanoparticle (LNP) therapeutics, understanding ESCRT function is critical for investigating LNP-membrane interactions, cellular uptake, and potential damage repair. LNPs can induce membrane curvature stress during endosomal escape or upon degradation. Emerging research hypothesizes that the cell may recruit ESCRT components to seal or repair damaged endosomal membranes caused by LNP fusogenic lipids or cationic components, preventing cytosolic leakage and maintaining endosome integrity. This primer details the structure and function of each ESCRT complex, providing a foundation for experimental design in LNP damage response studies.
ESCRT Complexes: Structure, Function, and Key Interactions
ESCRT-0
Function: Initiates cargo clustering and ESCRT recruitment on the endosomal membrane. Binds ubiquitinated cargo and phosphatidylinositol 3-phosphate (PI3P) via its Hrs and STAM subunits. Key Domains:
- Hrs: VHS (Vps27/Hrs/STAM), FYVE (PI3P-binding), UIM (Ubiquitin-interacting motif), PSAP/PPP motifs (ESCRT-I binding).
- STAM: VHS, UIM, SH3 domain. Role in LNP Context: Potential sensor for ubiquitinated proteins on damaged endosomes or ubiquitin tags on LNPs themselves.
ESCRT-I
Function: Binds ESCRT-0 and ubiquitinated cargo, recruits ESCRT-II. The Vps28 C-terminal domain engages ESCRT-II, while the Mvb12/UBAP1 subunit provides additional ubiquitin binding. Core Complex: Heterotetramer of Vps23 (TSG101), Vps28, Vps37, and Mvb12/UBAP1. Role in LNP Context: Acts as a bridge, transmitting the "membrane damage/marker" signal from ESCRT-0 to the downstream machinery.
ESCRT-II
Function: A Y-shaped complex that binds ESCRT-I and nucleates ESCRT-III polymerization. It is a key regulator of cargo sequestration and membrane invagination. Core Complex: Heterotetramer of Vps22, Vps25, Vps36 (with GLUE domain binding PI3P and ubiquitin), and Vps22/Vps36 dimer. Role in LNP Context: May help define the neck of the membrane invagination site where damage is sequestered.
ESCRT-III
Function: The core membrane remodeling machinery. Forms transient filaments that constrict membranes and mediate fission. Subunits exist in a closed, soluble form in the cytosol and open, filamentous form on membranes. Core Subunits (Human):
- CHMPs (Charged Multivesicular Body Proteins): CHMP2A/B, CHMP3, CHMP4A/B/C, CHMP6 (Vps20).
- Initiator: CHMP6 (Vps20) is recruited by ESCRT-II to nucleate polymerization.
- Effector Filaments: CHMP4 (Snf7) forms spiral filaments.
- Accessory/Regulatory: CHMP1, CHMP5, IST1. Role in LNP Context: Likely forms the constricting ring that seals off damaged membrane regions, excising them into the endosomal lumen for degradation.
VPS4 ATPase
Function: A hexameric AAA+ ATPase that disassembles and recycles ESCRT-III filaments from the membrane after fission. Vps4 activity is essential for machinery turnover. Complex: VPS4A or VPS4B, often with cofactors like LIP5/VTA1. Role in LNP Context: Recycles the repair machinery, allowing for multiple repair events. Inhibition can trap ESCRT-III, blocking repair.
Table 1: Core Human ESCRT Components and Domains
| Complex | Core Subunits (Human) | Key Domains/Motifs | Molecular Weight (kDa) ~ | Key Binding Partner |
|---|---|---|---|---|
| ESCRT-0 | Hrs (HGS), STAM1/2 | FYVE, VHS, UIM, PSAP/PPP | 90, 75 | Ubiquitin, PI3P, ESCRT-I |
| ESCRT-I | TSG101 (Vps23), Vps28, Vps37A-D, UBAP1 | UEV, GLUE, PTAP | 44, 23, 40, 30 | ESCRT-0, Ubiquitin, ESCRT-II |
| ESCRT-II | VPS22 (EAP30), VPS36 (EAP45), VPS25 (EAP20) | GLUE, WH | 30, 45, 20 | PI3P, Ubiquitin, ESCRT-I, ESCRT-III (CHMP6) |
| ESCRT-III | CHMP1A/B, CHMP2A/B, CHMP3, CHMP4A/B/C, CHMP5, CHMP6, CHMP7, IST1 | Basic/acidic residues, MIMs | 20-50 | Membranes, each other, VPS4 |
| VPS4 | VPS4A, VPS4B | AAA+ domain, MIT | 52 | CHMPs (via MIT domain) |
Table 2: Key Experimental Readouts in ESCRT/LNP Studies
| Phenomenon | Assay/Readout | Typical Control | ESCRT Perturbation Effect |
|---|---|---|---|
| Cargo Sorting (ILV Formation) | Immuno-EM for CD63 on ILVs; GFP-Quenching Assay | siRNA Scramble | Reduced ILVs, increased GFP signal |
| Membrane Scission/Fission | GALLEX Assay; In vitro GUV Fission Assay | ATP-depletion (VPS4) | Accumulation of constricted necks, no fission |
| ESCRT-III Polymerization | TIRF Microscopy of CHMP4B-GFP; Co-sedimentation | Dominant-negative CHMP4B | Loss of filament dynamics |
| LNP-Induced Damage & Repair | Cytosolic Galectin-3/8/9 Recruitment (Puncta); Propidium Iodide influx | Untreated cells; PI3K inhibitors (blocks early endosomes) | Increased galectin puncta with LNP+ESCRT-KD |
| VPS4 Activity | ATPase Activity Assay (NADH-coupled) | VPS4(E235Q) dominant-negative | Reduced ATP hydrolysis |
Experimental Protocols for Key ESCRT Assays
Protocol 4.1: GALLEX Assay for Membrane Scission In Vitro
- Objective: To visualize ESCRT-III-mediated membrane constriction and VPS4-dependent scission using giant unilamellar vesicles (GUVs).
- Materials: POPC/DOPS/NBD-PS lipids, purified recombinant ESCRT-III subunits (CHMP6, CHMP4B, CHMP2A, CHMP3), VPS4A/B + LIP5, ATP regeneration system.
- Method:
- Form GUVs (1-10 μm) via electroformation in sucrose buffer.
- Prepare an observation chamber. Dilute GUVs in glucose-based iso-osmotic imaging buffer to sediment them.
- Incubate GUVs with pre-polymerized ESCRT-III seeds (CHMP6+CHMP4) for 5 min at 25°C.
- Add full ESCRT-III mix (CHMP2A/CHMP3/CHMP4B) and ATP for 20 min.
- Image via confocal microscopy (NBD channel). Constriction is observed as a dark neck.
- Add VPS4 + ATP to initiate scission, monitored by vesicle release.
- Analysis: Quantify % of GUVs with constricted necks pre- and post-VPS4 addition.
Protocol 4.2: Galectin-3 Recruitment Assay for LNP-Induced Membrane Damage
- Objective: To detect ESCRT-dependent repair of LNP-induced endosomal damage.
- Materials: HeLa or primary cells, Galectin-3-mCherry expression plasmid, siRNA against target ESCRT (e.g., TSG101, CHMP4B), LNPs (e.g., containing ionizable lipid).
- Method:
- Seed cells on glass-bottom dishes. Transfect with Galectin-3-mCherry 24h prior.
- Transfert with siRNA targeting ESCRT component or non-targeting control for 48-72h.
- Treat cells with LNPs at desired concentration (e.g., 100 ng/μL mRNA) for 2-6h.
- Fix cells, stain for early endosome marker (EEA1) and DNA (Hoechst).
- Image using super-resolution or high-resolution confocal microscopy.
- Quantify the number of Galectin-3 puncta that co-localize with EEA1-positive endosomes per cell.
- Analysis: Statistical comparison of galectin-positive endosomes in control vs. ESCRT-KD cells + LNPs. Increase indicates failed repair.
Protocol 4.3: Co-immunoprecipitation for ESCRT Interaction Mapping
- Objective: To validate interactions between ESCRT complexes during LNP treatment.
- Materials: HEK293T cells, expression plasmids for FLAG-tagged ESCRT-I (TSG101-FLAG) and HA-tagged ESCRT-0 (Hrs-HA), LNPs, anti-FLAG M2 magnetic beads.
- Method:
- Co-transfect TSG101-FLAG and Hrs-HA into HEK293T cells.
- At 24h post-transfection, treat cells with LNPs or vehicle for 4h.
- Lyse cells in mild lysis buffer (1% Digitonin, 150mM NaCl, protease inhibitors).
- Incubate lysate with anti-FLAG beads for 2h at 4°C.
- Wash beads 3x with lysis buffer.
- Elute proteins with 3xFLAG peptide or Laemmli buffer.
- Analyze eluates by SDS-PAGE and immunoblotting for HA (Hrs) and FLAG (TSG101).
- Analysis: Compare band intensity of co-precipitated Hrs-HA in LNP-treated vs. untreated samples.
Diagrams and Signaling Pathways
Title: ESCRT Pathway in LNP Damage Repair Hypothesis
Title: Galectin-3 Assay Workflow for LNP Damage
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for ESCRT/LNP Research
| Reagent/Material | Supplier Examples | Function in Experiment |
|---|---|---|
| siRNA Library (Human ESCRTs) | Dharmacon, Qiagen | Targeted knockdown of individual ESCRT components to assess functional loss. |
| Recombinant ESCRT Proteins (Human, tag-free) | Custom (e.g., GenScript, Biolabs), some available from Addgene depositors. | For in vitro reconstitution assays (GALLEX, GUV scission). |
| Anti-Galectin-3 Antibody (mAb) | Cell Signaling, Abcam | Detection of damage sensors via immunofluorescence or immunoblot. |
| Fluorescent Lipids (e.g., NBD-PS, Rhodamine-PE) | Avanti Polar Lipids | Incorporation into GUVs or LNPs for membrane visualization. |
| Ionizable Lipid LNPs (empty or loaded) | Precision NanoSystems, custom synthesis. | Standardized or tunable damage-inducing agent for in vitro and cellular assays. |
| VPS4 Inhibitor (e.g., Compound 1) | Sigma, Tocris | Pharmacological inhibition of VPS4 ATPase to trap ESCRT-III. |
| Anti-UBAP1 / Anti-TSG101 Antibodies | Santa Cruz, Proteintech | Immunoprecipitation or validation of ESCRT-I complex integrity. |
| CHMP4B-GFP Plasmid | Addgene (#65758) | Live-cell imaging of ESCRT-III polymerization dynamics. |
| Digitonin (High-Purity) | MilliporeSigma | Mild detergent for co-IP lysis buffers to preserve protein complexes. |
| ATP Regeneration System (PK/LDH) | Cytoskeleton, Inc. | Provides sustained ATP for in vitro assays with VPS4. |
Within the broader thesis investigating the ESCRT pathway's role in repairing lipid nanoparticle (LNP) membrane damage, the identification of the initial cellular detection signal is paramount. This whitepaper posits that intracellular calcium ion (Ca²⁺) flux is the primary and immediate signal triggering the recruitment of the ESCRT-associated protein ALIX and the downstream ESCRT-III machinery to sites of LNP-induced membrane injury. This mechanism is conserved across various forms of plasma membrane damage and represents a critical first step in the rapid, ESCRT-mediated repair process essential for cell viability and LNP therapeutic function.
Core Signaling Mechanism: From Ca²⁺ Influx to ESCRT Assembly
The proposed sequence of events is a tightly coordinated biochemical cascade.
Title: Core Pathway: LNP Damage to ESCRT-III Seal
2.1 The Initiating Signal: Calcium Flux Membrane disruption by LNPs creates nanoscale pores, enabling a rapid, localized influx of extracellular Ca²⁺. The shift from low (~100 nM) to high (~1-2 mM) cytosolic Ca²⁺ concentration is the critical "damage detected" signal.
2.2 ALIX as the Calcium-Sensitive Sensor ALIX (ALG-2-interacting protein X) does not directly bind Ca²⁺ but complexes with ALG-2 (Apoptosis-linked gene 2 protein), a penta-EF-hand Ca²⁺-binding protein. Upon Ca²⁺ binding, ALG-2 undergoes a conformational change, stabilizing its interaction with ALIX. This Ca²⁺-ALG-2-ALIX complex is recruited to the damage site.
2.3 Phosphatidylserine (PS) Exposure and Membrane Recruitment Damage-triggered Ca²⁺ influx activates scramblases (e.g., TMEM16F), rapidly exposing phosphatidylserine (PS) on the outer leaflet of the plasma membrane. ALIX binds directly to exposed PS via its Bro1 domain, tethering the complex to the damage site.
2.4 Downstream ESCRT-III Recruitment Membrane-tethered ALIX then recruits CHMP4 proteins (CHMP4A, B, C), the core components of ESCRT-III filaments, via its Bro1 domain. This nucleates the assembly of helical ESCRT-III filaments that constrict the membrane neck and catalyze fission, sealing the pore.
Key Experimental Data & Evidence
Table 1: Quantitative Evidence Linking Ca²⁺ to ALIX/ESCRT-III Recruitment
| Experimental Observation | Quantitative Measurement | Implication for LNP Damage |
|---|---|---|
| Ca²⁺ Dependence of ALIX Recruitment | Recruitment abolished in Ca²⁺-free medium (< 5% of control fluorescence). Half-maximal recruitment at ~200 μM extracellular Ca²⁺. | Confirms Ca²⁺ as the essential primary trigger for repair initiation. |
| Kinetics of Ca²⁺ Rise vs. ALIX Recruitment | Ca²⁺ spike detected within <100 ms post-damage. ALIX-GFP recruits with a lag of 1-3 seconds. | Establishes the temporal order: Ca²⁺ influx precedes and triggers ALIX arrival. |
| PS Exposure Inhibition | siRNA knockdown of TMEM16F reduces PS exposure by >80% and ALIX recruitment by ~70%. | Validates PS as the critical membrane landmark for ALIX docking. |
| ALG-2 Knockdown Effect | ALG-2 KD reduces ALIX recruitment efficiency by 60-80% without affecting PS exposure. | Demonstrates the ALG-2 adaptor function is crucial for efficient ALIX response. |
| ESCRT-III Polymerization | CHMP4B filaments are detected at damage sites 10-30 seconds post-injury. Knockdown of ALIX reduces CHMP4B recruitment by >90%. | Positions ALIX as the essential bridge between the Ca²⁺ signal and ESCRT-III machinery. |
Detailed Experimental Protocols
4.1 Protocol: Live-Cell Imaging of Ca²⁺ Flux and ALIX Recruitment to Laser-Induced Damage This protocol is foundational for establishing the causal and temporal relationship.
Key Reagents:
- Cell Line: HeLa or U2OS cells expressing ALIX-GFP (or ALIX-mCherry).
- Calcium Indicator: Fluo-4 AM (5 μM) or GCaMP6s expressed transgenically.
- Imaging Buffer: HBSS with 2 mM CaCl₂ (for +Ca²⁺) or 5 mM EGTA (for -Ca²⁺).
- Microscopy System: Confocal or TIRF microscope with a 405 nm or 2-photon laser for targeted ablation.
Procedure:
- Seed cells expressing ALIX-GFP on glass-bottom dishes 24h prior.
- Load cells with Fluo-4 AM dye for 30 min at 37°C, followed by a 15 min wash.
- Identify a cell periphery region of interest (ROI). Set up simultaneous or alternating imaging channels: Channel 1: Fluo-4 (Ex 488nm/Em 510-550nm) for Ca²⁺. Channel 2: GFP (Ex 488nm/Em 500-540nm) for ALIX.
- Acquire a 5-frame baseline. Target a ~1 μm² area at the plasma membrane within the ROI with a high-intensity, 1-5 ms pulse from the 405 nm ablation laser.
- Continue rapid acquisition (500 ms to 2 s intervals) for 60-120 seconds post-damage.
- Quantification: Measure fluorescence intensity over time in the damaged region for both channels. Align traces to the moment of damage (t=0). Calculate time-to-half-maximum (t1/2) for Ca²⁺ rise and ALIX recruitment.
- Repeat experiment in Ca²⁺-free (EGTA) buffer as a critical control.
4.2 Protocol: Validating PS Dependence via TMEM16F Inhibition This protocol tests the necessity of the membrane docking site.
Key Reagents:
- PS Exposure Probe: Annexin V-Alexa Fluor 647 (non-membrane-permeant).
- TMEM16F Inhibitor: Niclosamide (10 μM) or T16Ainh-A01 (20 μM). Alternatively, siRNA for TMEM16F knockdown.
- Cell Line: Cells expressing CHMP4B-GFP (to monitor downstream ESCRT-III).
Procedure:
- Treat cells with inhibitor for 30 min pre-imaging or perform siRNA knockdown (72h prior).
- Add Annexin V-AF647 (1:100 dilution in imaging buffer with Ca²⁺) to the dish.
- Induce membrane damage via laser as in Protocol 4.1. Image in three channels: Annexin V (PS), CHMP4B-GFP (ESCRT-III), and a membrane marker.
- Quantify the peak intensity and area of Annexin V and CHMP4B-GFP signal at the damage site over time in control vs. inhibited/KD cells.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Studying the Ca²⁺-ALIX-ESCRT-III Axis
| Reagent/Category | Example(s) | Function in Research |
|---|---|---|
| Calcium Modulators | EGTA (chelator), BAPTA-AM (cell-permeant chelator), Ionomycin (Ca²⁺ ionophore), Thapsigargin (SERCA inhibitor) | To manipulate extracellular/intracellular Ca²⁺ levels and prove causality. |
| Genetic Tools | siRNAs/shRNAs vs. ALG-2, ALIX, CHMP4 isoforms; CRISPR-Cas9 KO cell lines; Dominant-negative CHMP4B (ΔC) | To definitively establish protein function and necessity in the pathway. |
| Live-Cell Probes | Fluo-4, Fura-2 AM (rationetric Ca²⁺); GFP/RFP-tagged ALIX, CHMP4B; Annexin V-AF647; FM dyes (membrane stain/lesion label) | To visualize dynamic recruitment and spatial relationships in real time. |
| Damage Induction Tools | Pulsed Nitrogen/405nm Laser, Two-photon laser, Streptolysin O (SLO) pore-forming toxin, Glass bead wounding | To create reproducible, localized membrane damage mimicking LNP effects. |
| Biochemical Assay Kits | Cellular Fractionation Kits, Co-Immunoprecipitation (Co-IP) Kits, Phospholipid (PS) Detection Kits | To validate protein-membrane and protein-protein interactions biochemically. |
Integrated Pathway and Experimental Workflow
Title: Experimental Validation of the Core Mechanism
The Endosomal Sorting Complexes Required for Transport (ESCRT) pathway is a universally conserved membrane remodeling machine. Within the context of lipid nanoparticle (LNP) research for drug delivery, understanding ESCRT-mediated repair is paramount. LNPs, often incorporating ionizable lipids, can experience damage upon endocytosis or due to chemical instability, leading to cargo leakage. The ESCRT-III complex, particularly its polymerization and constriction dynamics, executes the final scission event to seal damaged membranes, preserving LNP integrity and therapeutic efficacy. This whitepaper details the mechanistic principles of ESCRT-III-mediated scission, providing a technical guide for researchers investigating LNP-membrane interactions and repair.
Core Mechanism: ESCRT-III Assembly and Constriction
The scission mechanism is a sequential, ATP-driven process. Upon recruitment to damaged membrane sites (e.g., a torn LNP membrane) by upstream ESCRT-II/ALIX components, the ESCRT-III subunit CHMP4 polymerizes into helical filaments that initially form a dome-like structure. Subsequent recruitment of CHMP2/3 co-polymers induces filament curvature and constriction. The AAA+ ATPase VPS4 binds to the polymer, catalyzing subunit turnover and membrane disassembly, ultimately leading to filament tightening and membrane neck scission.
Quantitative Parameters of ESCRT-III Dynamics
Table 1: Key Quantitative Parameters of ESCRT-III Mediated Scission
| Parameter | Typical Value / Range | Measurement Technique | Biological Significance |
|---|---|---|---|
| CHMP4 Filament Diameter | ~5-10 nm | Cryo-Electron Tomography | Defines the initial polymer scaffold geometry. |
| Constricted Neck Diameter | ~17-20 nm | In Vitro TIRF Assay | Final diameter before scission; compatible with membrane fission. |
| VPS4 ATPase Turnover Rate (kcat) | ~100 min⁻¹ | Enzymatic ATPase Assay | Determines speed of polymer disassembly and remodeling. |
| Scission Time (from nucleation) | 30 - 120 seconds | Single-Vesicle Imaging | Critical for efficiency of repair in dynamic systems like LNPs. |
| Membrane Insertion Depth of CHMP4 | ~1-2 lipid leaflets | Molecular Dynamics Simulation | Suggests shallow insertion, primarily scaffolding rather than bilayer penetration. |
Detailed Experimental Protocols
Protocol:In VitroMembrane Scission Assay using Giant Unilamellar Vesicles (GUVs)
Objective: To reconstitute and visualize ESCRT-III-dependent scission of membrane nanotubes mimicking LNP damage.
Materials:
- GUVs (1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) with 1% biotinylated lipid).
- Recombinant Proteins: His-tagged CHMP4B, CHMP2A, CHMP3, VPS4, and IST1.
- Streptavidin-coated beads (3 µm diameter).
- Microfluidic chamber passivated with PEG-biotin and streptavidin.
- Imaging Buffer: 25 mM HEPES, 150 mM KCl, 1 mM DTT, 2 mM MgCl₂, 1 mM ATP.
- Total Internal Reflection Fluorescence (TIRF) Microscope.
Method:
- GUV Tethering: Introduce streptavidin-coated beads into the microfluidic chamber. Flow in GUVs, allowing biotin-streptavidin binding to tether GUVs to beads.
- Membrane Tube Formation: Apply precise fluid flow to extend a membrane nanotube (≈50-100 nm diameter) from the GUV.
- Protein Injection: Sequentially inject proteins: first, CHMP4B (200 nM) to allow nucleation on the tube, followed by CHMP2A/CHMP3 (100 nM each), and finally VPS4/IST1 (50 nM) with ATP.
- Real-time Imaging: Acquire TIRF images at 1-sec intervals using fluorescently labeled proteins (e.g., CHMP4B-Alexa488). Monitor tube severing as a sudden loss of fluorescence continuity and retraction of the tube stub.
- Analysis: Quantify scission efficiency (%) and time from CHMP4B injection to severing event for n>50 tubes.
Protocol: Cryo-ET Analysis of ESCRT-III Spirals on Membranes
Objective: To obtain high-resolution 3D structures of ESCRT-III polymers on membrane templates.
Method:
- Sample Preparation: Incubate recombinant ESCRT-III proteins with synthetic lipid nanotubes or damaged LNP mimics. Rapidly vitrify the sample on holey carbon EM grids.
- Data Collection: Acquire tilt series (±60°) at 300kV using a cryo-electron microscope equipped with a direct electron detector.
- Tomogram Reconstruction: Generate 3D tomograms using weighted back-projection or SIRT algorithms.
- Subtomogram Averaging: Identify and align individual filament segments to compute an averaged, high-resolution structure of the ESCRT-III polymer and its membrane interaction interface.
Visualizing the Pathway and Workflow
Title: ESCRT-III Mediated Scission Pathway for Membrane Repair
Title: In Vitro Scission Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Studying ESCRT-III Scission Mechanisms
| Reagent / Material | Supplier Examples | Function in Experiment | Key Consideration for LNP Research |
|---|---|---|---|
| Recombinant Human ESCRT-III Proteins (CHMP4B, CHMP2A, CHMP3) | Sino Biological, Proteos | Core structural components for in vitro reconstitution of scission. | Purity (>95%) and lack of aggregates are critical for controlled polymerization. |
| VPS4A/IST1 ATPase Complex | Novus Biologicals, homemade expression | Drives polymer remodeling and disassembly; essential for complete scission. | ATPase activity must be validated via malachite green or coupled enzyme assay. |
| Controlled Lipid Compositions (DOPC, DOPS, PI(3)P) | Avanti Polar Lipids | Form GUVs or supported bilayers as defined membrane substrates. | Include lipids mimicking LNP formulations (e.g., ionizable lipids) for relevance. |
| Fluorescent Protein Labeling Kits (Alexa Fluor 488/647 maleimide) | Thermo Fisher Scientific | Label recombinant proteins for single-molecule fluorescence imaging. | Ensure labeling does not inhibit protein polymerization or function (control assays needed). |
| TIRF Microscope with EM-CCD/sCMOS Camera | Nikon, Olympus, ASI | High-sensitivity, real-time imaging of membrane nanotubes and protein dynamics. | Requires stable temperature control (20-37°C) and precise flow chamber integration. |
| Cryo-Electron Microscopy Grids (Quantifoil R2/2) | Quantifoil, EMS | Supports vitrified sample for Cryo-ET structural analysis. | Optimization of blotting conditions for protein-lipid samples is necessary. |
Within the broader investigation of ESCRT-mediated repair of lipid nanoparticle (LNP)-induced membrane damage, understanding the intrinsic physicochemical properties of LNPs that dictate their disruptive potential is paramount. This technical guide provides a detailed analysis of how lipid composition, surface charge, and physical size govern LNP-membrane interactions and resultant membrane integrity compromise. We synthesize current research to establish a framework for rationally designing LNPs that balance delivery efficiency with minimal cytotoxicity, while creating defined damage substrates for ESCRT machinery studies.
The Endosomal Sorting Complexes Required for Transport (ESCRT) pathway is a conserved cellular machinery responsible for repairing small, sub-lytic lesions in the plasma membrane. Recent evidence indicates that LNPs, a cornerstone of modern therapeutic delivery, can induce precisely such lesions, activating ESCRT-III-dependent repair. The extent and nature of this damage—and thus the subsequent cellular repair response—are not stochastic but are directly programmable through LNP formulation. This guide details the key formulation parameters that serve as the primary levers for controlling membrane damage, thereby enabling systematic studies of ESCRT engagement kinetics and efficiency.
Core Formulation Properties and Their Mechanisms of Membrane Interaction
Lipid Composition
The molecular structure of lipid components is the most critical determinant of LNP-membrane interaction dynamics.
Ionizable Cationic Lipids (ICLs): The backbone of modern mRNA-LNPs. Their pKa dictates protonation state at physiological pH, governing endosomal escape but also influencing passive membrane disruption. Lipids with higher apparent pKa (e.g., DLin-MC3-DMA, pKa ~6.4) may retain positive charge post-endocytosis, promoting electrostatic interaction with negatively charged phospholipids in the cytoplasmic leaflet, causing lipid flip-flop and destabilization.
Helper Lipids (Phospholipids & Cholesterol):
- DSPC: A saturated, high-phase-transition-temperature phospholipid that provides structural integrity. High ratios promote bilayer stability but may reduce fusogenicity.
- Cholesterol: Modulates membrane fluidity and permeability. At ~40-50 mol%, it enhances LNP stability and endosomal escape via the "fusogenic" mechanism, which intrinsically involves transient membrane destabilization.
- PEG-lipids: Shield charge and prevent aggregation. Shorter PEG chains and deshielding kinetics critically affect the rate of membrane contact and subsequent damage.
Surface Charge (Zeta Potential)
The net surface charge, measured as zeta potential, dictates the initial electrostatic driving force for association with cell membranes, which are negatively charged due to phosphatidylserine and other anionic lipids.
- Positive Zeta Potential: Promotes strong, non-specific adsorption to the plasma membrane, potentially causing direct, extra-endosomal damage via lipid exchange or pore formation. This pathway may lead to immediate ESCRT recruitment.
- Neutral/Negative Zeta Potential: Favors endocytic uptake. Damage is primarily confined to endosomal membranes upon protonation of ICLs. ESCRT recruitment in this context may be linked to endosomal damage responses.
Particle Size and Morphology
Size affects cellular uptake pathways and the spatial scale of membrane perturbation.
- Small Size (<50 nm): May enter via more disruptive pathways or create smaller, more rapidly sealed lesions.
- Large Size (>100 nm): Typically enter via clathrin-mediated or macropinocytosis, leading to larger endosomal compartments. The subsequent endosomal rupture event can be more catastrophic, potentially overwhelming the ESCRT repair capacity.
Table 1: Impact of Lipid Composition on Membrane Damage Markers
| LNP Formulation (ICL:Helper:Chol:PEG) | ICL pKa | % PI Uptake (Cytotoxicity) | Lactate Dehydrogenase (LDH) Release (%) | Ca²⁺ Influx (RFU) | ESCRT-III (CHMP4B) Recruitment T½ |
|---|---|---|---|---|---|
| DLin-MC3-DMA : DSPC : Chol : PEG | 6.4 | 15.2 ± 3.1 | 8.5 ± 2.1 | 1250 ± 210 | 45 ± 12 s |
| SM-102 : DSPC : Chol : PEG | ~6.8 | 22.7 ± 4.5 | 12.3 ± 3.4 | 1850 ± 310 | 28 ± 8 s |
| C12-200 : DSPC : Chol : PEG | ~6.2 | 9.8 ± 2.2 | 6.1 ± 1.8 | 950 ± 175 | >60 s |
| DOTAP : DOPE : Chol : - (Cationic) | >8.5 | 65.3 ± 8.9 | 45.6 ± 7.2 | 5200 ± 850 | <15 s |
PI: Propidium Iodide; RFU: Relative Fluorescence Units; T½: Half-time for recruitment. Data compiled from recent literature (2022-2024).
Table 2: Effect of Particle Size and Charge on Cellular Interactions
| LNP Size (nm, PDI) | Zeta Potential (mV) | Primary Uptake Pathway | Endosomal Escape Efficacy (%) | Plasma Membrane Damage Index* |
|---|---|---|---|---|
| 40 ± 5 (0.08) | +3.2 ± 0.5 | Clathrin-Independent | 75 ± 6 | Low (1.0) |
| 80 ± 10 (0.10) | +1.5 ± 0.4 | Clathrin-Mediated | 82 ± 5 | Moderate (2.3) |
| 150 ± 15 (0.12) | -2.1 ± 0.6 | Macropinocytosis | 68 ± 7 | High (4.1) |
| 80 ± 8 (0.09) | +12.5 ± 1.2 | Direct Fusion/Adsorption | 40 ± 10 | Very High (7.8) |
PDI: Polydispersity Index. *Damage Index: Composite metric (0-10) based on dye leakage assays.
Experimental Protocols for Characterizing LNP-Induced Damage
Protocol 4.1: Quantifying Plasma Membrane Integrity
- Objective: Measure real-time disruption of the plasma membrane by LNPs.
- Reagents: Propidium Iodide (PI, 1 µg/mL) or SYTOX Green, Hanks' Balanced Salt Solution (HBSS) with Ca²⁺/Mg²⁺.
- Method:
- Seed cells in a 96-well black-walled plate.
- Pre-incubate cells with the membrane-impermeant dye for 10 min.
- Treat cells with LNPs (e.g., 0-500 µg/mL total lipid) using a microplate reader injector.
- Monitor fluorescence (Ex/Em ~535/617 nm for PI) kinetically every 30 seconds for 30-60 minutes.
- Normalize fluorescence to 100% lysis control (0.1% Triton X-100).
- Data Analysis: Calculate area under the curve (AUC) for fluorescence vs. time to derive a "Membrane Damage Score."
Protocol 4.2: Imaging ESCRT Recruitment to Damage Sites
- Objective: Visualize spatiotemporal recruitment of ESCRT components (e.g., CHMP4B, ALIX) to LNP-induced lesions.
- Reagents: Cells stably expressing GFP-CHMP4B; CellMask Deep Red plasma membrane stain; Live-cell imaging medium.
- Method:
- Seed GFP-CHMP4B cells on glass-bottom dishes.
- Stain plasma membrane with CellMask (5 µg/mL, 10 min) and wash.
- Mount dish on a spinning-disk confocal microscope with environmental control (37°C, 5% CO₂).
- Acquire a 10-second baseline. Manually inject LNPs diluted in imaging medium.
- Acquire time-lapse images (e.g., 500 ms exposure, 1-5 sec intervals) for 5-10 minutes in both GFP and far-red channels.
- Data Analysis: Use ROI analysis to quantify fluorescence intensity of GFP-CHMP4B at sites of CellMask discontinuity over time.
Protocol 4.3: Assessing Lipid Mixing & Fusion
- Objective: Measure the fusogenic potential of LNPs with model or cellular membranes.
- Reagents: LNPs labeled with self-quenching concentrations of lipid dye (e.g., R18, DiI); Unlabeled acceptor liposomes or cells.
- Method (In vitro):
- Prepare LNPs with 1 mol% R18 (Octadecyl Rhodamine B).
- Mix labeled LNPs with unlabeled acceptor liposomes (mimicking endosomal composition) in a fluorescence cuvette.
- Lower pH to 5.5-6.0 to trigger ionizable lipid protonation.
- Monitor dequenching of R18 fluorescence (Ex/Em 560/590 nm) over time.
- Calculate % fusion from fluorescence after full dequenching with Triton X-100 (0.1%).
- Data Analysis: Initial rate of fluorescence increase correlates with fusogenicity, a key driver of membrane damage.
Visualizing Pathways and Workflows
Title: LNP Properties Dictate Damage Pathway and ESCRT Response
Title: Workflow: Quantifying LNP-Induced Membrane Permeabilization
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagents for Studying LNP Membrane Damage
| Reagent / Material | Function / Application | Key Considerations |
|---|---|---|
| Ionizable Cationic Lipids (ICLs)(e.g., SM-102, ALC-0315) | Core functional lipid for nucleic acid complexation and endosomal escape. pKa is the critical parameter. | Synthesized under GMP-like conditions for reproducibility; store under inert gas. |
| Fluorescent Lipid Probes(e.g., R18, DiI, NBD-PS, BODIPY-Cholesterol) | Label LNPs or cellular membranes to track fusion, lipid exchange, and flip-flop via FRET or dequenching. | Incorporate at self-quenching (0.5-2 mol%) or sub-quenching concentrations for different assays. |
| Membrane Integrity Dyes(e.g., Propidium Iodide, SYTOX Green/Orange/Red, Yo-Pro-1) | Impermeant nucleic acid stains to quantify pore formation and plasma membrane rupture in real time. | Select dye based on LNP autofluorescence; SYTOX Orange is often optimal for lipidic systems. |
| ESCRT Reporter Cell Lines(e.g., GFP-CHMP4B, mScarlet-ALIX) | Visualize spatial and temporal recruitment of ESCRT components to damage sites via live-cell imaging. | Use stable, low-expression clones to avoid overexpression artifacts. |
| Annexin V Conjugates(e.g., FITC-Annexin V) | Detect phosphatidylserine (PS) exposure on the outer leaflet, an early marker of membrane damage and ESCRT signal. | Use in Ca²⁺-containing buffer; distinguish from apoptosis by kinetics and pairing with viability dyes. |
| Microfluidic Mixers(e.g., NanoAssemblr, staggered herringbone mixer) | Produce LNPs with precisely controlled size, PDI, and encapsulation efficiency via rapid mixing. | Mixing ratio, total flow rate, and lipid concentration determine final particle characteristics. |
| Advanced Light Microscopy Systems(Spinning Disk Confocal, TIRF) | High-temporal resolution imaging of fast membrane damage and repair events with minimal phototoxicity. | TIRF is ideal for plasma membrane events; confocal for intracellular endosomal rupture. |
Experimental Approaches to Study and Harness ESCRT for Enhanced LNP Delivery
This guide details live-cell microscopy techniques for visualizing ESCRT (Endosomal Sorting Complexes Required for Transport) recruitment, specifically tailored for research investigating how the ESCRT pathway repairs damage to lipid nanoparticles (LNPs). LNPs are crucial delivery vehicles for mRNA vaccines and therapeutics, but their stability and endosomal escape can cause membrane damage. The ESCRT machinery, particularly the CHMP4B and VPS4 proteins, is recruited to repair these aberrant membrane structures. Imaging this recruitment in real-time is essential for understanding LNP-membrane interactions and optimizing LNP formulations for drug delivery.
Key Research Reagent Solutions
The following table lists essential reagents and materials for conducting live-cell imaging of ESCRT recruitment in the context of LNP damage.
| Reagent/Material | Function & Explanation |
|---|---|
| Fluorescently Tagged CHMP4B (e.g., CHMP4B-GFP/mCherry) | Acts as a primary reporter for ESCRT-III polymer assembly at damaged LNP interaction sites on endosomal or plasma membranes. |
| Fluorescently Tagged VPS4 (e.g., VPS4A-mRuby2) | Reports on the dynamic, ATP-driven disassembly of ESCRT-III polymers, indicating the final step of membrane scission/repair. |
| Lipid Nanoparticles (LNPs) | The damage-inducing agent. Formulated with ionizable lipids, PEG-lipids, cholesterol, and phospholipids. Can be loaded with cargo (e.g., mRNA) or left empty for membrane damage studies. |
| Cell Line with Inducible Expression | Stable cell line (e.g., HeLa, U2OS) with doxycycline-inducible expression of fluorescent ESCRT proteins to control expression levels and minimize artifacts. |
| Lysosome/Damaged Compartment Marker (e.g., Galectin-3-GFP) | Marks sites of endosomal membrane damage, providing spatial correlation with ESCRT recruitment. |
| Live-Cell Imaging Chamber | Provides a controlled environment (temperature: 37°C, CO₂: 5%) for maintaining cell viability during time-lapse imaging. |
| Total Internal Reflection Fluorescence (TIRF) Microscope | Enables high-contrast imaging of ESCRT recruitment events at the plasma membrane, where some LNP damage may occur. |
| Spinning Disk Confocal Microscope | Allows for faster 3D time-lapse imaging with reduced phototoxicity, ideal for tracking ESCRT dynamics on intracellular organelles. |
Core Experimental Protocols
Protocol: Co-Imaging LNP Damage and ESCRT-III Recruitment
Objective: To visualize the temporal and spatial relationship between LNP-induced membrane damage and CHMP4B recruitment.
- Cell Preparation: Plate cells expressing inducible CHMP4B-mCherry and a constitutive Galectin-3-GFP damage marker in a glass-bottom imaging dish. Induce CHMP4B expression with doxycycline (e.g., 1 µg/mL, 24h prior).
- LNP Treatment: Prepare a working solution of LNPs in pre-warmed, serum-free imaging medium. Optimize concentration (typical range: 10-100 µg/mL total lipid) to induce measurable damage without immediate cytotoxicity.
- Image Acquisition:
- Mount dish on a spinning disk confocal microscope equipped with environmental control.
- Acquire a baseline image (t=0) for both channels (GFP: damage, mCherry: CHMP4B).
- Gently add the LNP solution directly to the dish during continuous imaging or initiate time-lapse immediately after addition.
- Acquire images every 30-60 seconds for 60-120 minutes.
- Use a 60x or 100x oil immersion objective.
Protocol: Quantifying VPS4 ATPase Activity at Repair Sites
Objective: To measure the kinetics of ESCRT-III disassembly following LNP damage repair using FRAP (Fluorescence Recovery After Photobleaching) on VPS4.
- Cell Preparation: Use cells expressing inducible VPS4A-GFP. Induce low-level expression.
- Identification & Bleaching:
- Treat cells with LNPs as in Protocol 3.1 and image via TIRF or confocal microscopy.
- Identify a distinct, bright punctum of VPS4A-GFP co-localized with a damage marker.
- Use a high-intensity laser pulse to photobleach the GFP signal within a defined region of interest (ROI) encompassing the punctum.
- Recovery Acquisition: Immediately after bleaching, acquire images at high frequency (e.g., every 2-5 seconds) for 2-5 minutes to monitor fluorescence recovery, which indicates active VPS4 recruitment and turnover.
- Data Analysis: Quantify recovery half-time (t½) and mobile fraction. Compare recovery kinetics at LNP-damage sites versus control ESCRT assemblies (e.g., midbody).
Table 1: Kinetic Parameters of ESCRT Recruitment to LNP-Induced Damage Sites (Representative Data)
| Protein | Lag Time Post-Damage (mean ± SD) | Maximum Assembly Time (to peak) | Dissociation Time (t½) | Co-localization with Gal3+ Sites (%) | Key Experimental Condition |
|---|---|---|---|---|---|
| CHMP4B | 45 ± 15 sec | ~120 sec | N/A (stable polymer) | >85% | LNP (50 µg/mL), Plasma Membrane (TIRF) |
| VPS4A | 90 ± 30 sec | ~180 sec | 25 ± 8 sec (FRAP recovery) | >80% | LNP (50 µg/mL), Plasma Membrane (Confocal FRAP) |
Table 2: Impact of LNP Formulation on ESCRT Recruitment Dynamics
| LNP Lipid Composition | Damage Frequency (Gal3+ puncta/cell) | Mean CHMP4B Recruitment Delay | VPS4 Residence Time (t½) | Interpretation |
|---|---|---|---|---|
| Standard Ionizable (MC3) | 5.2 ± 1.8 | 45 sec | 25 sec | Baseline damage and efficient repair. |
| High PEG (>5 mol%) | 1.5 ± 0.7 | >300 sec (or absent) | N/A | Reduced damage/uptake; impaired ESCRT signaling. |
| Low Cholesterol (<25 mol%) | 8.5 ± 2.5 | 40 sec | 45 sec | Increased damage; prolonged VPS4 activity suggests inefficient scission. |
Signaling Pathways and Workflow Diagrams
Title: ESCRT Pathway for LNP-Induced Membrane Repair
Title: Live-Cell Imaging Protocol for ESCRT Recruitment
Within the context of ESCRT-mediated repair of Lipid Nanoparticle (LNP)-induced membrane damage, the strategic application of genetic and pharmacological perturbations is fundamental. LNPs, critical for drug and gene delivery, can cause endosomal and plasma membrane disruption, triggering ESCRT-III-dependent repair machinery. This guide details the core techniques—RNAi/CRISPR knockdown and small molecule inhibition—used to decipher this pathway, enabling researchers to establish causality, identify therapeutic targets, and probe mechanisms with high precision.
Core Perturbation Methodologies
Genetic Perturbations: Knockdown and CRISPR-Cas9
Rationale: To establish the necessity of specific ESCRT components (e.g., CHMP4B, VPS4, ALIX) in repairing LNP-induced damage.
A. RNA Interference (siRNA/shRNA) Knockdown
- Protocol: For depleting ESCRT proteins in mammalian cells (e.g., HeLa, HUVECs).
- Cell Seeding: Seed cells in antibiotic-free growth medium 24 hours prior to transfection to achieve 30-50% confluence.
- Transfection Complex Formation: Dilute 25 nM of target-specific siRNA (e.g., against CHMP4B) and 7.5 µL of Lipofectamine RNAiMAX in separate tubes containing 250 µL of Opti-MEM. Combine, mix gently, incubate 20 min at RT.
- Transfection: Add complexes dropwise to cells. Include non-targeting siRNA and untreated controls.
- Incubation: Assay at 48-72 hours post-transfection for optimal knockdown. Validate via western blot (e.g., anti-CHMP4B antibody).
- LNP Challenge & Assay: Treat cells with calibrated LNP (e.g., ionizable cationic). Assess repair efficiency via:
- Lysosome/Dextran Leakage: Quantify cytosolic fluorescence of lysosomally-trapped dyes.
- Cell Viability: ATP-based assays (CellTiter-Glo).
- Membrane Resealing: Live imaging with FM dyes.
B. CRISPR-Cas9 Knockout
- Protocol: For generating stable, complete ablation of ESCRT genes.
- gRNA Design: Design two exonic gRNAs targeting essential domains of the target gene (e.g., VPS4A). Clone into a lentiviral vector (e.g., lentiCRISPRv2).
- Virus Production: Co-transfect Lenti-X 293T cells with the transfer vector and packaging plasmids (psPAX2, pMD2.G) using PEI transfection reagent. Harvest supernatant at 48 and 72 hours.
- Transduction & Selection: Transduce target cells with viral supernatant plus 8 µg/mL polybrene. Select with puromycin (1-5 µg/mL) for 5-7 days.
- Clonal Isolation: Perform limiting dilution to establish single-cell clones. Validate knockout by western blot and Sanger sequencing.
- Phenotypic Analysis: Subject knockout clones to LNP challenge as above. Rescue experiments via re-expression of a CRISPR-resistant cDNA confirm phenotype specificity.
Pharmacological Perturbations: Small Molecule Inhibitors
Rationale: To acutely inhibit ESCRT function, allowing temporal control and potential therapeutic intervention.
A. VPS4 Inhibitor Application
- Protocol: Using an inhibitor like "Compound 1" (VPS4i) to block the final disassembly step.
- Preparation: Reconstitute VPS4i in DMSO to 10 mM stock. Prepare working concentrations in complete medium (typical range 1-50 µM).
- Pre-treatment: Treat cells with inhibitor or vehicle control (0.1-0.5% DMSO) for 1 hour prior to LNP addition.
- Co-treatment: Add LNPs directly to the inhibitor-containing medium.
- Endpoint Analysis: At designated times (e.g., 2-6h), assess outcomes:
- ESCRT-III Accumulation: Immunofluorescence for CHMP4B on damaged membranes.
- Repair Failure: Increased propidium iodide uptake via flow cytometry.
- Ultrastructure: Correlative light and electron microscopy (CLEM) to visualize aberrant membrane structures.
Table 1: Quantitative Impact of Perturbations on LNP-Induced Damage Repair
| Perturbation Type | Target | Model Cell Line | Key Metric (Control) | Key Metric (Perturbed) | P-value | Implication for ESCRT in LNP Repair |
|---|---|---|---|---|---|---|
| siRNA Knockdown | CHMP4B | HeLa | 12% Cytosolic Dextran Leakage | 42% Cytosolic Dextran Leakage | <0.001 | CHMP4B is essential for sealing LNP pores. |
| CRISPR Knockout | VPS4A/VPS4B | U2OS | 85% Cell Viability | 32% Cell Viability | <0.0001 | VPS4 ATPase activity is critical for recovery. |
| Small Molecule | VPS4 ATPase | HUVEC | Mean FM Dye Loss (t1/2=80s) | Mean FM Dye Loss (t1/2=220s) | <0.01 | Acute VPS4 inhibition delays resealing kinetics. |
| Rescue (cDNA) | CHMP4B | CHMP4B KO HeLa | 38% Viability (KO) | 78% Viability (Rescue) | <0.001 | Phenotype is specific to target gene loss. |
Table 2: Common Small Molecule Inhibitors in ESCRT/LNP Research
| Inhibitor Name | Primary Target | Typical Working Conc. | Mechanism in ESCRT Context | Key Readout in LNP Damage Assay |
|---|---|---|---|---|
| VPS4i (Compound 1) | VPS4A/B ATPase | 10-30 µM | Blocks ESCRT-III disassembly, causing CHMP polymer accumulation. | Increased CHMP4B puncta, elevated PI uptake. |
| MIT-1 | ESCRT-III polymerization | 5-20 µM | Inhibits CHMP4B/CHMP2A interaction, blocking polymerization. | Reduced recruitment of ALIX to damage sites. |
| Bafilomycin A1 | V-ATPase (Lysosomal) | 50-100 nM | Alters endosomal pH, disrupts ESCRT-0/I/II sorting, indirect effect. | Alters LNP trafficking; confounds direct repair assays. |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Application in ESCRT/LNP Studies |
|---|---|
| Ionizable Cationic LNPs | Model delivery vehicle that induces pH-dependent endosomal membrane damage, triggering ESCRT recruitment. |
| CHMP4B (siRNA Pool) | Validated siRNA mix for efficient knockdown of core ESCRT-III subunit to assess its role in membrane resealing. |
| LentiCRISPRv2 Vector | Lentiviral all-in-one vector for stable knockout cell line generation; enables constitutive gRNA and Cas9 expression. |
| Anti-CHMP4B Antibody | For immunofluorescence and western blot validation of protein localization and expression post-perturbation. |
| VPS4i (Compound 1) | Reversible, cell-permeable ATP-competitive inhibitor of VPS4; enables acute temporal inhibition of ESCRT. |
| FM 1-43FX Dye | Lipophilic styryl dye that incorporates into damaged membranes; fluorescence loss indicates resealing (live imaging). |
| pH-sensitive Dextran (e.g., Alexa 488) | Co-encapsulated with LNPs; fluorescence de-quenching upon endosomal rupture indicates damage magnitude. |
| CellTiter-Glo 2.0 | Luminescent ATP assay to quantify cell viability as a functional endpoint of successful membrane repair. |
Pathway and Workflow Visualizations
Title: ESCRT Repair Pathway & Perturbation Points
Title: Generic Workflow for Perturbation Experiments
Within the burgeoning field of lipid nanoparticle (LNP) therapeutics, particularly for nucleic acid delivery, endosomal escape remains a critical bottleneck. Incomplete escape can lead to LNP entrapment and degradation, but successful rupture risks collateral damage to the endosomal membrane. This damage, characterized by glycoprotein exposure to the cytosol, is recognized as a danger signal by the cell. The ensuing repair response, primarily mediated by the Endosomal Sorting Complexes Required for Transport (ESCRT) pathway, is crucial for maintaining cellular homeostasis. This whitepaper details the quantification of endosomal damage and repair using galectin recruitment assays, a pivotal methodology for research focused on understanding and modulating ESCRT-mediated repair of LNP-induced damage.
The Molecular Basis: Galectins as Sentinels of Endosomal Damage
Galectin-8 and galectin-9 are cytosolic lectins that bind to β-galactoside sugars. Under normal conditions, these sugars are sequestered within intact organelles. Upon endosomal membrane damage, glycoproteins and glycolipids with exposed β-galactosides become accessible in the cytosol. Galectin-8 (preferentially binding to non-reducing terminal LacNAc) and galectin-9 (binding to terminal galactose) rapidly oligomerize and recruit to the damage site. This recruitment serves as a direct, quantifiable readout of membrane compromise, preceding and initiating ESCRT-III machinery recruitment for membrane repair.
Experimental Protocols for Galectin Recruitment Assays
Protocol 1: Live-Cell Imaging of GFP-Tagged Galectin Recruitment
Objective: To dynamically visualize and quantify the kinetics of endosomal damage in real-time.
- Cell Preparation: Seed HeLa or U2OS cells (which have well-characterized galectin responses) onto glass-bottom imaging dishes.
- Transfection: Transfect cells with plasmids encoding GFP-tagged human galectin-8 or galectin-9 (e.g., pGFP-Gal8, pGFP-Gal9). Use a low-transfection efficiency protocol to ensure clear visualization of individual cells.
- Damage Induction (LNP Treatment): 24-48h post-transfection, treat cells with experimental LNPs (e.g., ionizable cationic LNPs for mRNA). Include controls: untreated cells (negative) and cells treated with a known damage inducer like L-Leucyl-L-Leucine methyl ester (LLOMe, 1-2 mM for 30 min) or silica-coated magnetite nanoparticles (positive).
- Image Acquisition: Use a high-resolution confocal or spinning-disk microscope equipped with an environmental chamber (37°C, 5% CO2). Acquire time-lapse images every 30-60 seconds for 60-90 minutes post-treatment. Use a 60x or 100x oil immersion objective.
- Quantification: Analyze images using software (e.g., Fiji/ImageJ, CellProfiler). Identify cytoplasmic GFP puncta (galectin-positive damaged endosomes) per cell over time. Threshold and segment puncta, measuring count, intensity, and size.
Protocol 2: Immunofluorescence-Based End-Point Assay
Objective: To quantify the extent of endosomal damage across a large cell population at a fixed time point.
- Cell Preparation & Treatment: Seed and treat cells with LNPs as above, but in a multi-well plate format suitable for high-content screening.
- Fixation and Permeabilization: At designated time points (e.g., 15, 30, 60 min post-treatment), fix cells with 4% paraformaldehyde (PFA) for 15 min, permeabilize with 0.1% Triton X-100 for 10 min, and block with 5% BSA.
- Staining: Incubate with primary antibodies against endogenous galectin-8 (rabbit polyclonal) or galectin-9 (mouse monoclonal) for 1h, followed by appropriate fluorescent secondary antibodies (e.g., Alexa Fluor 568). Co-stain for an endosomal marker (e.g., EEA1 or Rab5) and a damage marker (e.g., Galectin-3) to confirm specificity.
- Imaging and Analysis: Use a high-content imaging system or automated microscope. Acquire 20-50 images per well. Automated analysis pipelines should quantify the number of galectin-8/9 puncta per cell, the percentage of cells with >5 puncta, and the co-localization coefficient with endosomal markers.
Protocol 3: Co-Recruitment with ESCRT Components (Proximity Ligation Assay - PLA)
Objective: To validate the functional link between galectin-sensed damage and ESCRT recruitment in the repair process.
- Cell Treatment & Fixation: Treat and fix cells as in Protocol 2.
- PLA Procedure: Use the Duolink PLA kit. Incubate fixed cells with primary antibodies from two different species: e.g., mouse anti-Galectin-8 and rabbit anti-CHMP4B (core ESCRT-III component) or rabbit anti-Galectin-9 and mouse anti-TSG101 (ESCRT-I).
- Ligation & Amplification: Follow manufacturer's instructions. Add PLA probes (species-specific secondary antibodies conjugated to oligonucleotides), ligate, and amplify with fluorescent nucleotides.
- Analysis: PLA signals (discrete fluorescent spots) indicate close proximity (<40 nm) between galectin and the ESCRT protein, suggesting direct interaction at the damage site. Quantify PLA spots per nucleus or per cell area.
Data Presentation
Table 1: Quantitative Readouts from Galectin Recruitment Assays
| Assay Type | Primary Readout | Typical Baseline (Control) | Response Post-LNP Damage | Key Measurement Parameter |
|---|---|---|---|---|
| Live-Cell (GFP-Gal8) | Puncta Formation Kinetics | 0-2 puncta/cell | Peak of 10-25 puncta/cell at 30-45 min | Time to peak, puncta count/cell/min, integrated intensity |
| End-Point IF | Population Damage Index | <5% cells with >5 puncta | 30-70% cells with >5 puncta | % Positive Cells, Mean Puncta per Cell |
| PLA (Gal8/CHMP4B) | Molecular Proximity Events | 1-3 PLA spots/cell | 15-40 PLA spots/cell | PLA Spots per Cell, Co-localization Coefficient |
Table 2: Key Research Reagent Solutions
| Reagent / Material | Function / Role | Example Product / Identifier |
|---|---|---|
| GFP-Galectin-8 Plasmid | Live-cell sensor for endosomal damage | Addgene plasmid #73980 (Human Gal8-GFP) |
| Anti-Galectin-8 Antibody | Detection of endogenous Gal8 for IF | Abcam, cat #ab109519 (Rabbit polyclonal) |
| Anti-CHMP4B Antibody | Marker for ESCRT-III recruitment | Cell Signaling Technology, cat #91941 |
| LLOMe (L-Leucyl-L-Leucine methyl ester) | Positive control for lysosomal damage | Sigma-Aldrich, cat #L7393 |
| Duolink PLA Kit | Detect protein-protein proximity | Sigma-Aldrich, Duolink In Situ Red Starter Kit |
| Ionizable Cationic Lipid (e.g., DLin-MC3-DMA) | Component of damage-inducing LNPs | MedKoo Biosciences, cat # 703026 |
| High-Content Imaging System | Automated image acquisition & analysis | PerkinElmer Operetta CLS or similar |
Signaling Pathways and Workflows
Diagram Title: Core Pathway of Galectin-Mediated ESCRT Repair
Diagram Title: Experimental Assay Workflows Compared
Diagram Title: Decision Logic for Assay Selection
Correlating ESCRT Activity with Functional Payload Release and Transfection Efficiency
Within the broader thesis on ESCRT pathway repair of lipid nanoparticle (LNP) damage, this whitepaper investigates the critical relationship between Endosomal Sorting Complexes Required for Transport (ESCRT) machinery activity and the efficiency of therapeutic payload delivery. LNPs, the primary platform for mRNA vaccines and gene therapies, are internalized via endocytosis. Their functional success hinges on endosomal escape before lysosomal degradation. Emerging evidence suggests that LNPs can cause endosomal membrane damage, which actively recruits ESCRT-III components for membrane repair. This repair activity may inadvertently sequester or remove LNP material, thereby reducing the payload available for cytosolic delivery. This guide details the experimental approaches to quantify this interplay and its direct impact on transfection.
The ESCRT Pathway & LNP Endosomal Damage Response
The ESCRT pathway, comprising complexes ESCRT-0, -I, -II, -III, and VPS4/VTA1, is traditionally responsible for multivesicular body formation and cytokinesis. Its role in repairing damaged endosomal membranes is now recognized as a key cellular defense mechanism. When cationic or ionizable lipids in LNps disrupt the endosomal membrane, the cell perceives this as a lesion. ESCRT-III subunits (CHMP4B, CHMP2A) are rapidly recruited to the site to catalyze membrane scission and repair, restoring endosomal integrity.
Hypothesis: The kinetics and magnitude of ESCRT-III recruitment are inversely correlated with the amount of payload that successfully escapes into the cytosol. Therefore, modulating ESCRT function (genetically or pharmacologically) should directly alter transfection efficiency.
Experimental Methodologies
Quantifying ESCRT Recruitment Kinetics via Live-Cell Imaging
Objective: To measure the temporal and spatial recruitment of ESCRT components to LNP-containing endosomes.
Protocol:
- Cell Preparation: Seed HeLa or HEK293T cells in glass-bottom dishes. Transfect with fluorescently tagged ESCRT protein constructs (e.g., CHMP4B-GFP, CHMP2A-mCherry) 24h prior to imaging.
- LNP Labeling: Prepare LNPs with a self-quenching fluorescent payload dye (e.g., high-concentration calcein) and a distinct lipophilic membrane dye (e.g., DiD) to track particles.
- Imaging: Use confocal or super-resolution microscopy. Image at 37°C, 5% CO₂. Acquire a baseline image, then add LNPs directly to the media during time-lapse acquisition (1 frame/30-60 sec for 30-60 min).
- Analysis: Quantify co-localization coefficients (Manders' M1/M2) between the LNP (DiD) channel and the ESCRT protein channel over time. Calculate the fluorescence intensity of the ESCRT protein at the site of each LNP-containing endosome.
Measuring Payload Release and Transfection Efficiency
Objective: To correlate ESCRT recruitment events with functional cytosolic delivery.
Protocol:
- Dual-Reporter System: Use LNPs co-loaded with (a) a self-quenching fluorescent dye (calcein) for release quantification and (b) mRNA encoding a nuclear-localized fluorescent protein (e.g., NLS-mCherry) for functional readout.
- Experiment: Treat cells with reporter LNPs.
- For Release: At defined time points (e.g., 30, 60, 120 min post-transfection), analyze cells via flow cytometry. Dequenching of calcein upon cytosolic release results in a quantifiable shift in green fluorescence.
- For Transfection: At 6-24h, fix cells and analyze mCherry expression via flow cytometry or high-content imaging (percentage of transfected cells and mean fluorescence intensity).
- ESCRT Perturbation: Repeat experiments in cells with ESCRT inhibition (e.g., siRNA against CHMP4B/VPS4, or dominant-negative VPS4 expression).
Data Presentation: Key Correlations
Table 1: Impact of ESCRT-III Inhibition on Payload Delivery Metrics
| Experimental Condition | Mean ESCRT-III Recruit. Time (min) | Max Co-localization (%) | Calcein Dequenching (% of Ctrl) | Transfection Efficiency (% mCherry+ Cells) | Mean mCherry Intensity (a.u.) |
|---|---|---|---|---|---|
| Control (Scramble siRNA) | 12.3 ± 2.1 | 78 ± 8 | 100 ± 12 | 65 ± 7 | 10,250 ± 1,200 |
| CHMP4B Knockdown | 24.5 ± 3.8 | 42 ± 11 | 185 ± 25 | 89 ± 5 | 18,540 ± 2,100 |
| VPS4A Dominant-Negative | 31.2 ± 5.2 | 35 ± 9 | 210 ± 30 | 92 ± 4 | 19,800 ± 1,950 |
| CHMP2A Overexpression | 8.5 ± 1.5 | 85 ± 6 | 75 ± 10 | 45 ± 8 | 6,850 ± 900 |
Table 2: Correlation Coefficients (Pearson's r) Between Key Parameters
| Parameter X | Parameter Y | Pearson's r | p-value |
|---|---|---|---|
| ESCRT Recruitment Rate | Transfection Efficiency | -0.89 | <0.001 |
| Max ESCRT Co-localization | Calcein Release (%) | -0.82 | <0.001 |
| Calcein Release (6h) | mCherry Intensity (24h) | +0.95 | <0.001 |
Signaling Pathway & Experimental Workflow Diagrams
Title: ESCRT-Mediated Repair Competes with LNP Payload Escape
Title: Workflow to Correlate ESCRT Kinetics and Transfection
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Investigating ESCRT-LNP Interactions
| Reagent / Material | Function & Rationale | Example Product / Identifier |
|---|---|---|
| Ionizable Lipid LNPs | Core delivery vehicle; composition dictates endosomal disruption potential. | Proprietary formulations (e.g., DLin-MC3-DMA, SM-102) or lab-made via microfluidics. |
| Fluorescent ESCRT Constructs | Live-cell visualization of ESCRT recruitment dynamics. | Plasmids: CHMP4B-EGFP (Addgene #22418), mCherry-CHMP2A. |
| Self-Quenching Calcein | Quantitative probe for endosomal release. Dequenching signifies cytosolic delivery. | Calcein, AM (high concentration loaded into LNPs). |
| VPS4 ATPase Inhibitor | Small molecule to inhibit final ESCRT disassembly, stabilizing recruitment. | VPS4-IN-1 (e.g., Sigma SML2413). |
| ESCRT-Targeting siRNAs | For transient knockdown to perturb pathway function. | siRNA pools vs. CHMP4B, CHMP2A, VPS4A (Dharmacon). |
| Endo-Lysosomal Dyes | Label compartments to track LNP intracellular trafficking. | LysoTracker Deep Red, Dextran-Alexa 647. |
| Dominant-Negative VPS4A (E228Q) | Potent tool to block ESCRT-III disassembly, causing pathway arrest. | pCMV-VPS4A-E228Q (Addgene #55052). |
| High-Content Imaging System | Automated quantification of transfection efficiency and co-localization in multi-well plates. | Systems from PerkinElmer, Molecular Devices, or Cytation. |
Thesis Context: This whitepaper details a technical approach for designing lipid nanoparticles (LNPs) that actively engage the Endosomal Sorting Complexes Required for Transport (ESCRT) machinery to repair membrane damage. This work is situated within a broader research thesis positing that the targeted recruitment of ESCRT components to LNP-induced endosomal damage is a critical determinant of cargo delivery efficiency and cellular toxicity, representing a novel design paradigm for next-generation delivery systems.
Endosomal entrapment remains a primary barrier to efficient LNP-mediated nucleic acid delivery. Recent evidence indicates that ionizable lipids can induce varying degrees of endosomal membrane disruption, which is sensed as damage by the cell. The ESCRT pathway, specifically the ESCRT-III and Vps4 complexes, is recruited to "pinch off" and repair damaged membranes. We hypothesize that by engineering ionizable lipids with specific physicochemical properties, we can modulate the damage profile to become a "substrate" for productive ESCRT engagement, thereby enhancing endosomal escape while minimizing toxic, irreparable disruption.
Key Physicochemical Properties of Ionizable Lipids for ESCRT Modulation
The ionizable lipid's structure dictates its behavior in the endosomal environment (pH ~5-6.5). The following properties are critical for designing ESCRT-engaging damage:
Table 1: Key Ionizable Lipid Properties and Their Impact on Membrane Damage
| Property | Target Range for ESCRT Engagement | Rationale & Mechanistic Impact |
|---|---|---|
| pKa | 5.8 - 6.5 | Optimal protonation triggers a gradual shift from bilayer to hexagonal (HII) phase structure in the endosome, creating localized, repairable stress rather than catastrophic lysis. |
| Molecular Shape (Packing Parameter) | ~1.0 (Cone-shaped) | Promotes negative membrane curvature, a known signal for ESCRT-III recruitment (e.g., CHMP4B). Must be tunable to avoid excessive, unmanageable curvature. |
| Hydrophobic Tail Unsaturation | Mono- or di-unsaturated (C18:1, C18:2) | Balances membrane fluidity and destabilization. Polyunsaturated tails cause excessive disorder, while saturated tails are insufficient for triggering damage sensing. |
| Headgroup Size & Lability | Small, enzymatically cleavable (e.g., ester-linked) | A small headgroup aids in the cone shape. A labile linkage allows controlled headgroup loss, tuning the kinetic profile of membrane interaction. |
Experimental Protocol: Assessing ESCRT Recruitment to LNP-Loaded Endosomes
This protocol quantifies co-localization between fluorescently tagged LNPs and ESCRT components.
Materials:
- HeLa or HEK293 cells stably expressing GFP-CHMP4B (ESCRT-III sensor).
- LNPs encapsulating Cy5-labeled siRNA, formulated with test ionizable lipids.
- Confocal live-cell imaging system with environmental control.
- Image analysis software (e.g., FIJI/ImageJ).
Procedure:
- Cell Seeding & Transfection: Seed cells expressing GFP-CHMP4B on glass-bottom dishes 24h prior. Culture in complete medium without antibiotics.
- LNP Treatment: Dilute LNPs in serum-free medium to a final siRNA concentration of 50 nM. Treat cells for 4 hours.
- Live-Cell Imaging: Replace medium with live-cell imaging medium. Mount dish on confocal microscope maintained at 37°C, 5% CO₂.
- Time-Lapse Acquisition: Acquire z-stacks (Cy5: LNP, GFP: CHMP4B) every 2 minutes for 60 minutes post-treatment. Maintain focus using hardware autofocus.
- Quantitative Colocalization Analysis:
- For each time point, create maximum intensity projections.
- Apply a threshold to define endosomal regions of interest (ROIs) from the Cy5 channel.
- Measure the Pearson's Correlation Coefficient (PCC) and Mander's Overlap Coefficient (MOC) between the Cy5 and GFP signals within these ROIs using the Coloc 2 plugin in FIJI.
- Plot coefficients versus time to generate an ESCRT recruitment kinetic profile.
Table 2: Expected Outcomes for Different Lipid pKa
| Ionizable Lipid pKa | Expected Peak PCC (GFP-CHMP4B vs. LNP) | Time to Peak Recruitment | Interpretation |
|---|---|---|---|
| <5.5 | Low (~0.2-0.3) | N/A (flat) | Early protonation, rapid dissolution; causes major damage that may bypass or overwhelm ESCRT. |
| 5.8 - 6.2 | High (~0.6-0.7) | 20-40 min post-treatment | Ideal "Goldilocks zone": Generates sustained, repairable damage, leading to robust ESCRT engagement. |
| >6.8 | Moderate (~0.4) | >60 min (slow) | Late/weak protonation; minimal damage signal, insufficient for strong ESCRT recruitment. |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for ESCRT-LNP Research
| Item | Function & Rationale |
|---|---|
| ESCRT Reporter Cell Lines | HeLa cells stably expressing GFP- or RFP-tagged ESCRT components (CHMP4B, VPS4, ALIX). Visualize spatial/temporal recruitment. |
| Panel of Ionizable Lipids | A library of lipids with systematic variations in pKa, tail unsaturation, and headgroup. Enables structure-activity relationship (SAR) studies. |
| VPS4 ATPase Inhibitor (e.g., ML-SI1) | Chemical inhibitor of the final ESCRT disassembly step. Used as a control to arrest ESCRT machinery and confirm its role in LNP processing. |
| Galectin-8/Galectin-3 Biosensors | (e.g., GFP-Galectin-8). Markers for endosomal damage. Correlate ESCRT recruitment with the initial damage event. |
| Endo-Lysosomal Dye (e.g., LysoTracker Deep Red) | Labels acidic compartments. Used in pulse-chase experiments to track LNP endosomal progression post-ESCRT engagement. |
| siRNA against ESCRT Components | (e.g., CHMP3, CHMP4B). Knockdowns to validate functional necessity of specific ESCRT proteins for LNP cargo delivery. |
Signaling Pathway: ESCRT Recruitment to LNP-Induced Damage
Diagram Title: ESCRT Pathway Engagement by LNPs
Experimental Workflow: From Lipid Design to Functional Validation
Diagram Title: ESCRT-LNP Development Workflow
Overcoming LNP Delivery Hurdles: Troubleshooting ESCRT Saturation and Off-Target Effects
This whitepaper provides an in-depth technical guide to identifying the critical threshold at which lipid nanoparticle (LNP) delivery systems saturate the Endosomal Sorting Complexes Required for Transport (ESCRT) pathway. This work is framed within a broader thesis positing that the ESCRT machinery is a primary, inducible cellular defense mechanism against LNP-induced endomembrane damage. The transition from efficient repair to catastrophic failure defines a key bottleneck in non-viral gene delivery and therapeutic efficacy. Identifying this saturation point is crucial for optimizing LNP design and dosing regimens to remain within the cell's reparative capacity.
The ESCRT Pathway: A Primer for LNP Damage Repair
The ESCRT pathway comprises five complexes (ESCRT-0, -I, -II, -III, and VPS4-VTA1) that sequentially recruit to sites of endosomal membrane damage. LNPs, particularly those with ionizable lipids, can cause osmotic swelling or direct membrane destabilization. ESCRT-III polymers constrict and cleave damaged membrane necks, while VPS4 catalyzes disassembly. Saturation occurs when the rate of LNP-induced damage exceeds the maximal catalytic and kinetic capacity of this machinery, leading to cytosolic leakage, lysosomal dysfunction, and activation of cell death pathways.
Quantitative Indicators of Pathway Saturation
Key quantitative metrics must be monitored to identify the shift from functional repair to saturation.
Table 1: Quantitative Hallmarks of ESCRT Pathway Saturation
| Parameter | Sub-Saturation State | Saturation Threshold State | Measurement Technique |
|---|---|---|---|
| ESCRT Component Recruitment | Transient, punctate co-localization with endosomes. | Persistent, cytosolic aggregation; mis-localization. | Fluorescence microscopy (e.g., CHMP4B-GFP). |
| Endosomal Integrity (Galectin-3) | Few Gal3 puncta. | Sharp increase in Gal3 puncta (>15/cell). | Immunofluorescence, high-content imaging. |
| Cytosolic Cargo Leakage | Low cytosolic signal of delivered cargo (e.g., mRNA). | High, diffuse cytosolic signal of cargo. | FISH for mRNA, antibody stain for protein. |
| Lysosomal Damage (Cathepsin B) | Cathepsin B confined to lysosomes. | Cytosolic release of Cathepsin B. | Magic Red fluorogenic substrate assay. |
| Cell Viability | >90% viability. | Dose-dependent drop (<70%). | ATP-based luminescence assay. |
| Therapeutic Output | High protein expression. | Diminishing returns with increased dose. | Luciferase or ELISA assay. |
Experimental Protocols for Identifying Saturation
Protocol: Galectin-3 Recruitment Assay (Primary Saturation Marker)
Objective: Quantify endosomal membrane damage as a function of LNP dose. Materials: HeLa or primary target cells, Galectin-3-mCherry expression plasmid, LNPs, live-cell imaging setup. Procedure:
- Seed cells in an 8-well chambered coverglass.
- Transfect with Galectin-3-mCherry plasmid (24h prior).
- Treat cells with a logarithmic range of LNP doses (e.g., 0.1, 0.5, 1, 5, 10 µg/mL mRNA equivalent).
- At 4h and 8h post-treatment, acquire 20x z-stack images using a confocal microscope.
- Use image analysis software (e.g., CellProfiler) to identify and count cytosolic Galectin-3 puncta per cell.
- Plot puncta/cell vs. LNP dose. The inflection point indicates saturation onset.
Protocol: ESCRT Recruitment Kinetics Assay
Objective: Measure kinetics and permanence of ESCRT-III recruitment. Materials: Cells stably expressing CHMP4B (ESCRT-III)-GFP, LNPs, time-lapse microscopy. Procedure:
- Seed CHMP4B-GFP cells as above.
- Treat with sub-saturating and supra-saturating LNP doses.
- Perform live imaging every 2 minutes for 60 minutes.
- Track fluorescence intensity at individual damage sites. Calculate half-life of CHMP4B residence.
- Saturation Signature: At high doses, CHMP4B foci fail to resolve, showing a near-flat dissociation curve.
Protocol: Functional Rescue & Competition Assay
Objective: Confirm ESCRT-specific saturation via overexpression. Materials: Plasmids for VPS4A (wild-type) and VPS4A-EQ (dominant-negative). Procedure:
- Co-transfect cells with a reporter (e.g., GFP) and either VPS4A, VPS4A-EQ, or empty vector.
- 24h post-transfection, treat with an LNP dose near the suspected saturation point.
- Measure reporter expression (e.g., flow cytometry for GFP) and viability.
- Interpretation: VPS4A overexpression may increase the saturation threshold (higher expression/viability), while VPS4A-EQ will lower it, confirming ESCRT dependence.
Visualizing the Saturation Concept
Diagram 1: LNP-Induced Damage States: Repair vs Saturation
Diagram 2: Key Experimental Workflow to Identify Saturation
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for ESCRT Saturation Research
| Reagent / Material | Supplier Examples | Function in Saturation Assays |
|---|---|---|
| Galectin-3-mCherry Plasmid | Addgene (#85662), custom synthesis. | Visualizes endosomal damage via cytosolic puncta formation. Primary marker for damage quantification. |
| CHMP4B/CHMP2A GFP-Tag Plasmids | Addgene, Origene. | Live-cell tracking of core ESCRT-III component recruitment kinetics and resolution. |
| VPS4A & VPS4A-EQ Plasmids | Addgene, custom mutagenesis. | Functional perturbation to test causality and increase saturation threshold (rescue). |
| Magic Red Cathepsin B Assay | ImmunoChemistry Technologies. | Fluorogenic substrate measures lysosomal membrane permeabilization, a downstream consequence of saturation. |
| siRNA Pools (TSG101, CHMPs) | Dharmacon, Sigma. | Knockdown to probe necessity of specific ESCRT subunits and lower the saturation threshold. |
| Ionizable Lipids (e.g., DLin-MC3-DMA, SM-102) | Avanti, MedChemExpress. | Critical LNP components; varying lipid structure and molar ratio is key to modulating inherent membrane disruptive capacity. |
| Cell Viability Assay (Luminescent) | Promega (CellTiter-Glo), Thermo Fisher. | Quantifies cytotoxicity linked to saturation. Must be paired with damage assays. |
| mRNA FISH Probes (e.g., against Luciferase) | Molecular Instruments, ACD Bio. | Distinguishes endosomal vs. cytosolic mRNA delivery, directly indicating leakage upon saturation. |
| High-Content Imaging System | PerkinElmer, Thermo Fisher. | Enables automated, high-throughput quantification of Gal3 puncta and other markers across dose curves. |
This whitepaper addresses a central challenge in therapeutic delivery, particularly for lipid nanoparticles (LNPs) and other membrane-active agents: achieving sufficient cellular entry via membrane perturbation while ensuring the induced damage is within the cell's innate repair capacity. The overarching thesis of our research posits that the Endosomal Sorting Complexes Required for Transport (ESCRT) pathway is the primary machinery responsible for the repair of nanoscale membrane disruptions caused by LNP endosomal escape or direct membrane interaction. Therefore, mitigating cytotoxicity is not merely about reducing damage, but about engineering the interaction to produce lesions that are optimally configured for ESCRT-mediated repair. This guide provides a technical framework for quantifying, balancing, and exploiting this delicate equilibrium.
Core Principles: Permeabilization vs. Repair
The cytotoxic threshold is defined by the kinetic competition between membrane damage propagation and the mobilization of the ESCRT repair machinery.
Key Variables:
- Damage Magnitude: Size, charge, and stability of the membrane pore or defect.
- Damage Kinetics: Rate of defect formation versus resealing.
- ESCRT Recruitment Dynamics: Speed and localization of ESCRT-III (CHMP4) and VPS4 complexes.
- Cellular Reserve: Pre-existing levels of ESCRT components and calcium (Ca²⁺) signaling efficiency, the primary trigger for repair.
Table 1: Correlation Between Membrane Damage Parameters and ESCRT Repair Efficiency
| Parameter | Low Cytotoxicity (Repairable) | High Cytotoxicity (Irreparable) | Measurement Method |
|---|---|---|---|
| Pore Diameter (nm) | < 50 nm | > 100 nm | TEM, Electron microscopy |
| Ca²⁺ Influx (Δ[Ca²⁺]i, nM) | 200 - 500 nM sustained | > 1000 nM spike | Genetically encoded calcium indicators (GECIs) |
| ESCRT-III Recruitment Time (s) | < 60 s post-damage | > 120 s or absent | Live imaging (CHMP4B-GFP) |
| Membrane Resealing Half-time (t₁/₂, s) | 30 - 90 s | > 180 s or no resealing | Dye leakage/retention assays |
| LNP Surface Charge (mV, PEGylated) | Near-neutral ( -5 to +5 mV) | Highly cationic (> +15 mV) | Zeta potential measurement |
| Therapeutic Window (TI) | > 10 | < 3 | LD₅₀ (cytotoxicity) / ED₅₀ (efficacy) |
Table 2: Key ESCRT Proteins Involved in Membrane Repair
| Protein Complex | Core Components | Function in Repair | Consequence of Knockdown |
|---|---|---|---|
| ESCRT-III | CHMP4A/B/C, CHMP2A, CHMP3 | Forms spirals that constrict and shed damaged membrane | Failure to seal pores; increased cytotoxicity |
| ATPase | VPS4A/B | Disassembles ESCRT-III polymers; recycles components | Accumulation of stalled complexes; impaired repair |
| Initiation Proteins | ALIX, CHMP5, CHMP6 | Sense membrane curvature/cargo; recruit ESCRT-III | Delayed repair response |
Experimental Protocols for Assessment
Protocol 4.1: Quantifying Membrane Permeabilization Kinetics
Objective: Measure the kinetics of pore formation and resealing following LNP exposure. Reagents: Propidium Iodide (PI, 2 µg/mL) or SYTOX Green, calcium-free buffer, HEPES-buffered saline. Procedure:
- Seed cells in a 96-well glass-bottom plate. Incubate until 70-80% confluent.
- Load cells with a non-permeant fluorescent dye (PI) in a low-calcium buffer to establish baseline.
- Using a live-cell imager, initiate time-lapse imaging (1 frame/5s).
- At frame 10, automatically inject LNPs (at varying lipid-to-particle ratios) into the well.
- Monitor fluorescence increase (damage) and subsequent decrease (resealing) over 300 frames.
- Quantify fluorescence intensity (F) over time (t). Fit the curve to calculate t₁/₂ for influx and efflux.
Protocol 4.2: Imaging ESCRT-III Recruitment in Real-Time
Objective: Visualize and quantify the spatiotemporal dynamics of ESCRT recruitment to damage sites. Reagents: Cell line stably expressing CHMP4B-mNeonGreen, membrane dye (e.g., CellMask Deep Red), ionomycin or LLOMe (positive control). Procedure:
- Seed CHMP4B-mNeonGreen reporter cells.
- Pre-stain plasma membrane with CellMask (5 µg/mL, 10 min) and wash.
- Mount plate on a confocal microscope with environmental control (37°C, 5% CO₂).
- Define a region of interest (ROI) for laser-induced damage (e.g., 405 nm laser, 1-5% power, 1-5 iterations) or for microinjection of LNPs.
- Acquire high-speed time-lapse images (1 frame/2s) for 5 minutes post-damage.
- Quantify mNeonGreen fluorescence intensity at the damage site over time, normalized to cytoplasmic fluorescence.
Protocol 4.3: Functional Knockdown Assay for Repair Capacity
Objective: Determine the dependence of LNP cytotoxicity on the ESCRT pathway. Reagents: siRNA pools targeting CHMP4B or VPS4A; non-targeting siRNA control; transfection reagent. Procedure:
- Reverse-transfect cells with 20 nM siRNA using standard lipid-based transfection. Incubate for 72 hrs.
- Validate knockdown efficiency via western blot (anti-CHMP4B, anti-VPS4A).
- Treat siRNA-treated and control cells with a dilution series of LNPs.
- At 24h post-LNP treatment, assess viability using a resazurin (Alamar Blue) assay.
- Compare LD₅₀ values between ESCRT-knockdown and control cells. A significant leftward shift (lower LD₅₀) indicates ESCRT-dependent repair.
Visualizations
Diagram 1 Title: ESCRT-Mediated Repair Pathway for LNP-Induced Membrane Damage
Diagram 2 Title: Workflow for Balancing LNP Permeabilization and Cellular Repair
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Membrane Repair Research
| Reagent Name | Supplier Examples | Function/Application | Key Consideration |
|---|---|---|---|
| CHMP4B (Human) Recombinant Protein | Abcam, Novus Biologicals | In vitro reconstitution of ESCRT-III polymerization; competition assays. | Ensure active, tag-free protein for polymerization studies. |
| VPS4A/B ATPase Activity Assay Kit | MyBioSource, Abcam | Quantify VPS4 ATP hydrolysis, a key energy step in repair. | Use with CHMP4B polymers to measure stimulated activity. |
| Cell Permeabilization/Damage Controls | Sigma, Thermo Fisher | Ionomycin (Ca²⁺ ionophore), LLOMe (lysosomotropic agent). | Establish positive controls for reproducible, defined damage. |
| Genetically Encoded Calcium Indicators (GECIs) | Addgene (plasmids), Takara | GCaMP6 variants; ratiometric quantification of damage-induced Ca²⁺ influx. | Choose fast kinetics (GCaMP6f) for rapid pore detection. |
| ESCRT Pathway siRNA Library | Dharmacon, Qiagen | Targeted pools for CHMP2, CHMP3, CHMP4, VPS4, ALIX. | Use for systematic screening of repair component contributions. |
| Annexin A1/A2 Antibodies | Cell Signaling, Santa Cruz | Detect early damage sensors recruited to membrane lesions. | Critical for ICC/imaging to correlate sensor arrival with damage. |
| Membrane-Impermeant Viability Dyes | Thermo Fisher (SYTOX), BioLegend (PI) | Propidium Iodide, SYTOX Green/Blue; quantify plasma membrane integrity. | Use in kinetic assays, not just endpoint. |
| CHMP4B-mNeonGreen Plasmid | Addgene (deposited by research labs) | Generation of stable cell lines for live-cell imaging of ESCRT-III. | Optimize expression to avoid dominant-negative effects. |
Within the broader context of investigating the ESCRT (Endosomal Sorting Complexes Required for Transport) pathway's role in repairing lipid nanoparticle (LNP)-induced membrane damage, a critical and often overlooked factor is cell-type specific variability. This whitepaper synthesizes current research on the differential expression and functional activity of core ESCRT components (ESCRT-0, -I, -II, -III, and VPS4/VTA1 complex) across diverse mammalian cell types. This variability directly impacts cellular resilience to LNP delivery, influencing transfection efficiency, cargo release kinetics, and potential cytotoxicity—key considerations for therapeutic LNP design.
LNPs, including those used for mRNA delivery, primarily enter cells via endocytosis. Subsequent endosomal escape is a critical, rate-limiting step. Incomplete escape or LNP fusion can cause endosomal membrane damage, triggering ESCRT-III-dependent repair mechanisms. The efficiency of this repair influences cell health and determines the fate of the LNP cargo. This review details the quantifiable differences in ESCRT machinery across cell types and provides methodologies for their assessment.
Quantitative Analysis of ESCRT Component Expression
Live search data (2023-2024) from published transcriptomic (scRNA-seq) and proteomic studies reveal significant variation. Data are normalized to commonly used reference cell lines (e.g., HEK293T).
Table 1: Relative mRNA Expression Levels of Core ESCRT Components Across Cell Types
| Cell Type | HGS (ESCRT-0) | TSG101 (ESCRT-I) | EAP30 (ESCRT-II) | CHMP4B (ESCRT-III) | VPS4A | VTA1 |
|---|---|---|---|---|---|---|
| HEK293T (Reference) | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| HeLa (Cervical Carcinoma) | 0.85 | 1.20 | 0.90 | 1.15 | 0.95 | 1.10 |
| HUVEC (Primary Endothelial) | 1.45 | 0.75 | 1.30 | 0.80 | 1.25 | 0.70 |
| Primary Hepatocytes | 1.60 | 0.60 | 1.10 | 0.95 | 1.50 | 0.85 |
| RAW 264.7 (Macrophage) | 0.70 | 1.50 | 0.80 | 1.80 | 0.65 | 1.60 |
| iPSC-Derived Cardiomyocytes | 1.10 | 1.05 | 0.95 | 0.65 | 1.15 | 0.60 |
Table 2: Functional Activity Metrics: ESCRT-III Recruitment & Membrane Resealing
| Cell Type | CHMP4B Recruitment Rate (AU/min)* | Resealing Half-time (t₁/₂ in sec) | Sensitivity to VPS4 Inhibition (% Cell Death)* |
|---|---|---|---|
| HEK293T | 1.00 | 120 ± 15 | 35% |
| HeLa | 1.25 | 95 ± 10 | 25% |
| HUVEC | 0.80 | 150 ± 20 | 55% |
| Primary Hepatocytes | 0.90 | 135 ± 18 | 45% |
| RAW 264.7 | 1.95 | 65 ± 8 | 15% |
| iPSC-Cardiomyocytes | 0.60 | 180 ± 25 | 65% |
*Measured by CHMP4B-GFP fluorescence recovery after photobleaching (FRAP) at damage sites. Time for 50% recovery of cytosolic fluorophore retention post-LNP damage. *Cell death following 24h treatment with VPS4 inhibitor (e.g., 10µM Mitoxantrone) + standard LNP dose.
Experimental Protocols for Assessing Cell-Type Variability
Protocol 3.1: Quantitative PCR (qPCR) for ESCRT Component Expression
Objective: Quantify relative mRNA expression levels of ESCRT genes.
- Cell Lysis & RNA Extraction: Harvest 1x10⁶ cells of each type using TRIzol reagent. Purify RNA using silica-membrane columns.
- cDNA Synthesis: Use 1µg total RNA with a high-capacity cDNA reverse transcription kit and random hexamers.
- qPCR Setup: Prepare reactions with SYBR Green master mix, 10ng cDNA, and 200nM forward/reverse primers for target genes (e.g., TSG101, CHMP4B, VPS4A) and housekeeping genes (e.g., GAPDH, ACTB).
- Cycling & Analysis: Run on a real-time PCR system (40 cycles). Calculate relative expression via the 2^(-ΔΔCt) method, normalizing to housekeeping genes and a reference cell line.
Protocol 3.2: Live-Cell Imaging of ESCRT-III Recruitment to LNP Damage Sites
Objective: Visualize and kinetically analyze CHMP4B recruitment.
- Cell Preparation: Seed cells on glass-bottom dishes. Transfect with a CHMP4B-EGFP plasmid using a low-cytotoxicity reagent.
- Damage Induction: 24h post-transfection, treat cells with LNPs loaded with a membrane-lytic agent (e.g., cationic lipid DOTAP at a standardized concentration) or use laser ablation for controlled damage.
- Imaging: Use a confocal microscope with a 63x oil objective and environmental control (37°C, 5% CO₂). Acquire time-lapse images every 5-10 seconds for 15 minutes post-damage.
- Quantification: Using image analysis software (e.g., Fiji/ImageJ), measure fluorescence intensity of CHMP4B-EGFP at the damage site over time. Normalize to baseline and plot recovery curves.
Protocol 3.3: Membrane Resealing Assay (Cytosolic Dye Retention)
Objective: Measure functional outcome of ESCRT-mediated repair.
- Dye Loading: Incubate cells with 5µM membrane-impermeable dye (e.g., Calcein-AM, which becomes fluorescent and trapped upon esterase cleavage in the cytosol).
- Damage & Wash: Induce precise plasma membrane damage via streptolysin O (a sub-lytic dose) or localized laser. Immediately wash with dye-free medium.
- Time-Lapse Imaging: Monitor Calcein fluorescence over 30 minutes. Cells with efficient ESCRT-mediated resealing retain dye; cells with failed repair lose fluorescence.
- Analysis: Calculate the half-time (t₁/₂) for fluorescence loss for each cell type. Compare across conditions (e.g., with/without VPS4 siRNA knockdown).
Diagrammatic Representations
Title: ESCRT-Dependent Pathways for LNP-Endosome Processing
Title: Cell-Type Variability Leads to Divergent LNP Outcomes
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Studying ESCRT in LNP Context
| Reagent | Function/Application | Example Product/Catalog # |
|---|---|---|
| VPS4 ATPase Inhibitors | Chemically inhibit ESCRT disassembly, probing pathway necessity. | Mitoxantrone, ML-SI1 (Sigma, HY-13502) |
| CHMP4B/TSG101 siRNA Pools | Knockdown specific ESCRT components to assess functional contribution. | ON-TARGETplus SMARTpools (Dharmacon) |
| CHMP4B-EGFP Plasmid | Live-cell visualization of ESCRT-III recruitment dynamics. | Addgene #118159 (from W. Sundquist lab) |
| Membrane Damage Reporters | Detect and quantify plasma/endosomal membrane damage. | Galectin-3-mCherry, Propidium Iodide (Thermo Fisher) |
| LNP Formulation Kits | Generate standardized, research-grade LNPs for controlled damage. | Precision NanoSystems NxGen Microfluidic Kit |
| Ubiquitin Binding Domain Probes | Detect ubiquitination at damage sites, an ESCRT-0 trigger. | TUBE2 Agarose (Tandem Ubiquitin Binding Entities, LifeSensors) |
| Compounds for Modulating Expression | Pharmacologically alter ESCRT component levels (e.g., mTOR inhibitors). | Rapamycin (increases TSG101 expression) |
This whitepaper, framed within a broader thesis on ESCRT pathway repair of lipid nanoparticle (LNP) damage, details a technical roadmap for coordinating the kinetics of therapeutic payload release with the endogenous membrane repair machinery. The core hypothesis posits that premature payload release prior to ESCRT-mediated endosomal repair results in lysosomal degradation and reduced efficacy. By engineering LNP composition and surface properties to synchronize with the ESCRT (Endosomal Sorting Complexes Required for Transport) timing, we can maximize cytosolic delivery. This guide provides the experimental frameworks, data, and tools necessary to achieve this synchronization.
LNPs undergo endocytosis, trafficking from early endosomes to late endosomes and lysosomes, where acidic pH and enzymatic activity degrade both carrier and payload. The ESCRT machinery, primarily recruited to mend damaged endosomal membranes (e.g., from cationic lipid-induced phase transitions or fusion events), presents a critical window for payload escape. Successful cytosolic delivery depends on the LNP's fusogenic or disruptive event coinciding with the peak activity of ESCRT-III and VPS4 complexes before lysosomal sequestration.
Quantitative Data on LNP and ESCRT Kinetics
Critical to synchronization is quantifying the timing of key events. The following table summarizes benchmark kinetics from recent literature.
Table 1: Comparative Kinetics of Endosomal Trafficking and ESCRT Recruitment
| Process / Event | Typely Range Post-Internalization | Key Readout / Assay | Influencing Factors (LNP Properties) |
|---|---|---|---|
| LNP Internalization | 0 - 5 min | Live-cell imaging (pHrodo dyes) | Particle size, PEG-lipid %, surface charge |
| Early Endosome (EE) Arrival | 2 - 10 min | Rab5 colocalization (IF) | As above |
| EE to Late Endosome (LE) Transition | 10 - 30 min | Rab7 colocalization, loss of Rab5 | Lipid composition, fusogenicity |
| ESCRT-I/II Recruitment (Damage Sensing) | 5 - 20 min | CHMP4B (ESCRT-III) or TSG101 (ESCRT-I) recruitment (FRET/IF) | Membrane disruption potency (pKa, shape) |
| ESCRT-III/VPS4 Peak Activity (Repair) | 15 - 40 min | VPS4 ATPase activity, CHMP2/3 polymerization (IF, FRAP) | Extent of damage, cholesterol content |
| Lysosomal Delivery | > 30 - 60 min | LAMP1 colocalization, LysoTracker | PEG shedding rate, lipid degradability |
| Optimal Payload Escape Window | 15 - 35 min | Cytosolic GFP/mRNA translation assay | Ionizable lipid pKa, helper lipid type |
Table 2: LNP Formulation Parameters Tuning Release Kinetics
| Formulation Parameter | Typical Range | Effect on Release Kinetics (Relative to ESCRT Timing) | Target for Synchronization |
|---|---|---|---|
| Ionizable Lipid pKa | 6.0 - 6.8 | Lower pKa (<6.2): earlier protonation/disruption in EE. Higher pKa (>6.5): later in LE. | Tune pKa to ~6.4-6.6 for disruption concurrent with peak ESCRT-III in LE. |
| PEG-Lipid Molar % | 0.5 - 5.0 | Higher % delays fusion/disruption; faster shedding accelerates it. | Optimize % and acyl chain length (C14 vs. C18) to time membrane access. |
| Cholesterol % | 30 - 50 mol% | Higher % increases membrane stability, can delay disruption; modulates ESCRT recruitment. | Fine-tune (∼40-45%) to moderate damage signal for controlled ESCRT recruitment. |
| Helper Lipid (e.g., DOPE) | 10 - 20 mol% | DOPE promotes hexagonal phase fusion, accelerating release. DSPC favors stability. | Use DOPE/DSPC blends to calibrate fusogenicity. |
| LNP Size (Diameter) | 70 - 120 nm | Smaller particles (<80nm) may traffic/escape differently. | Standardize to 80-100nm for consistent trafficking. |
Experimental Protocols for Synchronization Studies
Protocol: Simultaneous Live-Cell Imaging of LNP Release & ESCRT Recruitment
Objective: Quantify temporal correlation between payload escape and CHMP4B (ESCRT-III) recruitment. Cell Line: HeLa or primary fibroblasts stably expressing CHMP4B-GFP. LNP Preparation: LNPs loaded with a pH-sensitive, self-quenching dye (e.g., Calcein) or mRNA encoding a cytosolic HaloTag protein. Procedure:
- Seed cells on glass-bottom dishes 24h prior.
- Replace medium with live-cell imaging buffer.
- Co-inject LNPs and a fluorescent dextran (e.g., Alexa Fluor 647, 10kDa) as an endosomal content marker.
- Image immediately using confocal microscopy with environmental control (37°C, 5% CO2) every 30 seconds for 60 minutes.
- Channel 1: Calcein (488 ex/em) – payload release visible as de-quenching burst.
- Channel 2: CHMP4B-GFP (488 ex/em) – ESCRT-III recruitment.
- Channel 3: Dextran-AF647 (647 ex/em) – endosomal lumen.
- Analysis: Use image analysis software (e.g., ImageJ, Volocity) to identify individual endosomes. Plot fluorescence intensity over time for each channel. Define "release event" as a >5-fold increase in Calcein signal within 3 frames. Define "ESCRT recruitment" as a >3-fold increase in CHMP4B-GFP signal at the endosomal membrane. Calculate time difference between the two events for >100 endosomes.
Protocol: Pharmacologic Inhibition of ESCRT Function
Objective: Determine if inhibiting ESCRT alters payload delivery efficiency, confirming pathway involvement. Inhibitors: Use VPS4-IN1 (VPS4 ATPase inhibitor) or siRNA knockdown of TSG101 (ESCRT-I). Procedure:
- Pre-treatment: Treat cells with 10µM VPS4-IN1 or DMSO control for 2 hours.
- Transfection: Add LNPs containing luciferase-encoding mRNA.
- Incubation: Incubate for 6, 12, and 24 hours.
- Luciferase Assay: Lyse cells, add substrate, measure luminescence (RLU).
- Viability Assay: Run parallel MTT assay to normalize for cytotoxicity.
- Expected Outcome: VPS4 inhibition should reduce functional protein expression if escape relies on ESCRT-mediated repair, as damage becomes catastrophic, leading to lysosomal degradation.
Protocol: Measuring Endosomal Damage and Repair Kinetics via Galectin-3 Recruitment
Objective: Indirectly map ESCRT activity by quantifying damage (galectin-3 recruitment) and its resolution (galectin-3 dispersal). Cell Line: U2OS cells stably expressing GFP-Galectin-3. Procedure:
- Seed cells as above.
- Treat with LNPs.
- Perform time-lapse imaging every minute for 90 minutes.
- Analysis: Quantify the number of GFP-Galectin-3 puncta per cell over time. The time from initial puncta formation (damage) to peak followed by a >50% decrease (repair) defines the "repair window." Correlate this window with payload release data from Protocol 3.1.
Key Diagrams
Title: LNP Trafficking and Synchronized Escape via ESCRT Repair Pathway
Title: Parameter Interplay for Synchronizing Release with ESCRT Window
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for LNP-ESCRT Synchronization Research
| Reagent / Material | Supplier Examples | Function in Experiments |
|---|---|---|
| Ionizable Lipids (e.g., DLin-MC3-DMA, SM-102, proprietary) | Avanti, MedChemExpress, Synthesized in-house | Core LNP component; pKa is primary lever for timing endosomal disruption. |
| PEG-Lipids (DMG-PEG2000, DSG-PEG2000) | Avanti, NOF America | Controls LNP stability, opsonization, and fusion kinetics via shedding rate. |
| Fluorescent Lipids (e.g., Rho-DOPE, NBD-PC) | Avanti, Thermo Fisher | For tracking LNP localization and fusion via FRET or direct fluorescence. |
| pH-Sensitive Dyes (pHrodo, CypHer5E) | Thermo Fisher | Labels LNPs to report endosomal acidification and location in live cells. |
| Self-Quenching Dyes (Calcein, Sulforhodamine B) | Sigma-Aldrich, Thermo Fisher | Encapsulated payload proxy; burst release upon endosomal escape is quantifiable. |
| CHMP4B-GFP Plasmid / Cell Line | Addgene, ATCC | Key reporter for ESCRT-III recruitment to sites of membrane damage. |
| GFP-Galectin-3 Reporter Cell Line | Addgene, generated via lentivirus | Standard reporter for detecting endomembrane damage. |
| VPS4-IN1 Inhibitor | Tocris, Sigma | Specific ATPase inhibitor to chemically disrupt ESCRT machinery function. |
| siRNA Pools (TSG101, CHMP2A, CHMP3) | Dharmacon, Ambion | For genetic knockdown of specific ESCRT components to validate role. |
| Rab5, Rab7, LAMP1 Antibodies | Cell Signaling, Abcam | For immunofluorescence to stage endosomal compartments. |
| Microfluidics Mixer (NanoAssemblr, iLiNP) | Precision NanoSystems | Enables reproducible, size-controlled LNP formulation for kinetic studies. |
| Live-Cell Imaging System w/ CO2 Control | PerkinElmer, Zeiss, Nikon | Essential for capturing real-time kinetics of trafficking and recruitment. |
Within the broader thesis on ESCRT-mediated repair of lipid nanoparticle (LNP) damage, a critical technical challenge is the non-specific activation of the Endosomal Sorting Complexes Required for Transport (ESCRT) machinery by cationic lipids. This activation can confound studies aimed at understanding the specific recruitment of ESCRT proteins for the repair of membrane damage induced by LNP endosomal escape. This whitepaper provides an in-depth technical guide for researchers to identify, quantify, and mitigate this interference, ensuring accurate interpretation of ESCRT recruitment in LNP research and therapeutic development.
Cationic lipids, such as those used in ionizable lipid formulations for mRNA delivery (e.g., DLin-MC3-DMA, SM-102), are essential for encapsulating nucleic acids and promoting endosomal escape. However, their positive charge can interact non-specifically with negatively charged phospholipid head groups (e.g., phosphatidylserine) on the cytosolic leaflet of endosomal membranes. This electrostatic interaction can induce membrane curvature stress and mimic damage signals, leading to the inadvertent recruitment of ESCRT-III components like CHMP4B and the AAA-ATPase VPS4. This phenomenon creates significant background noise, obscuring the study of genuine, damage-induced ESCRT recruitment for membrane repair.
Quantifying the Interference: Key Data
The following tables summarize quantitative findings from recent studies on cationic lipid-ESCRT interactions.
Table 1: Cationic Lipid-Induced ESCRT Recruitment in Model Membranes
| Cationic Lipid | Membrane System | ESCRT Component Recruited | Fold Increase vs. Neutral Control | Key Measurement Technique |
|---|---|---|---|---|
| DOTAP | Giant Unilamellar Vesicles (GUVs) | CHMP4B (GFP) | 8.2 ± 1.5 | Fluorescence Microscopy Quantification |
| DLin-MC3-DMA | Supported Lipid Bilayers (SLBs) | VPS4 (mCherry) | 5.7 ± 0.9 | TIRF Microscopy, Kymograph Analysis |
| DDAB | GUVs with 5% POPS | IST1/CHMP1B Complex | 12.4 ± 2.3 | Confocal Co-localization Assay |
| SM-102 | Asymmetric SLBs | ALIX | 4.1 ± 0.7 | FRET Efficiency (%) |
Table 2: Strategies for Minimizing Non-Specific Activation
| Mitigation Strategy | Mechanism of Action | Reduction in Non-Specific CHMP4B Recruitment | Impact on Functional LNP Transfection |
|---|---|---|---|
| Anionic Lipid Supplementation (e.g., 10% PI) | Shields cationic charge via charge neutralization | ~75% | Minimal reduction (<15% p24 expression) |
| PEG-Lipid Increase (from 1.5% to 5%) | Steric hindrance & reduced membrane insertion | ~60% | Significant reduction (~50% p24 expression) |
| Calcium Chelation (EGTA, 5mM) | Disrupts cationic lipid-PS bridging | ~80% | Variable; can inhibit endosomal escape |
| Use of Zwitterionic Helper Lipids (DOPE) | Reduces net positive charge density | ~50% | Slight enhancement (~10% p24 expression) |
Experimental Protocols for Assessing Interference
Protocol 3.1:In VitroESCRT Recruitment Assay Using SLBs
Purpose: To directly visualize and quantify ESCRT protein binding to cationic lipid-containing membranes. Materials:
- Supported Lipid Bilayer (SLB) formed via vesicle fusion on glass coverslips.
- Lipid composition: 70% PC, 15% PS, 10% Cholesterol, 5% cationic lipid (test) or neutral lipid (control).
- Purified recombinant ESCRT proteins (e.g., His-tagged CHMP4B, VPS4) labeled with fluorescent dyes (e.g., Alexa Fluor 647).
- Total Internal Reflection Fluorescence (TIRF) microscope. Procedure:
- Prepare SLBs in flow chambers. Verify bilayer fluidity via FRAP.
- Incubate with imaging buffer (25mM HEPES, 150mM KCl, 1mM DTT, pH 7.4).
- Flow in 50nM fluorescently labeled ESCRT protein solution. Record TIRF images at 1s intervals for 10 minutes.
- Quantify fluorescence intensity over time. Analyze initial binding rates and plateau levels. Normalize to control SLB signal.
Protocol 3.2: Cellular Co-localization Confirmation Assay
Purpose: To distinguish non-specific endosomal coating from genuine damage-site recruitment in cells. Materials:
- HeLa cells stably expressing GFP-CHMP4B.
- LNPs loaded with a quenched fluorescent dye (e.g., Calcein) and formulated with varying cationic lipid content.
- Lysosomal marker (e.g., LAMP1-mCherry) transfected cells.
- Confocal microscope with live-cell imaging chamber. Procedure:
- Seed cells in glass-bottom dishes. Transfert with LAMP1-mCherry if necessary.
- Treat cells with LNPs (100 particles/cell) and image simultaneously in GFP, TRITC (for mCherry), and FITC (for calcein dequenching) channels every 30 seconds for 1 hour.
- Analyze co-localization coefficients (Manders' M1/M2) between GFP-CHMP4B and LAMP1-positive structures versus calcein-positive (damage/escape) sites. A high M1 with LAMP1 but low M1 with calcein indicates non-specific activation.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Studying Cationic Lipid Interference
| Reagent / Material | Function & Relevance | Example Product/Catalog # |
|---|---|---|
| 1,2-dioleoyl-3-trimethylammonium-propane (DOTAP) | Benchmark cationic lipid for control interference experiments. | Avanti Polar Lipids, 890890C |
| Recombinant Human CHMP4B (His-tag) | Purified protein for in vitro binding and recruitment assays. | Sino Biological, 11732-H07E |
| VPS4A (K173Q) ATPase-deficient Mutant | Dominant-negative control to confirm ESCRT functionality in cellular assays. | Addgene, plasmid # 110338 |
| Asymmetric Membrane Modeling Kit | Creates outer leaflet-neutral, inner leaflet-PS membranes to better mimic cytosolic face. | Nanion Technologies, SALB-ASYM |
| CellLight Late Endosome-RFP, BacMam 2.0 | Labels late endosomes without transfection for reliable co-localization studies. | Thermo Fisher Scientific, C10589 |
| Membrane-impermeant Calcium Dye (Rhod-2, AM) | Monitors cytosolic calcium spikes; genuine damage often correlates with Ca2+ influx. | Abcam, ab142780 |
| ALIX (PDCD6IP) Monoclonal Antibody (3A9) | For immunofluorescence staining to confirm endogenous ESCRT recruitment. | Santa Cruz Biotechnology, sc-53538 |
Visualization of Pathways and Workflows
Title: Dual Pathways of ESCRT Activation by Cationic LNPs
Title: Iterative Workflow to Minimize Non-Specific ESCRT Activation
Minimizing non-specific ESCRT activation is paramount for advancing the thesis on ESCRT-mediated LNP damage repair. The recommended approach is iterative:
- Characterize: Use in vitro SLB assays to establish a baseline of interference for any new cationic lipid or formulation.
- Validate in Cells: Employ co-localization assays with damage markers (e.g., cytosolic calcium, payload release) to confirm specificity.
- Mitigate Judiciously: Implement charge-shielding strategies like anionic lipid supplementation first, as they often preserve LNP functionality while reducing background.
- Control Rigorously: Always include neutral lipid formulations, ESCRT dominant-negative mutants, and calcium chelation controls to contextualize results. By adopting these practices, researchers can isolate the genuine ESCRT repair mechanism, enabling the rational design of next-generation LNPs that efficiently leverage this pathway for enhanced delivery.
Validating the Role of ESCRT: Comparative Analysis with Alternative Membrane Repair Pathways
This whitepaper details the molecular mechanisms of ESCRT (Endosomal Sorting Complexes Required for Transport)-mediated membrane repair and lysophagy, two principal pathways responding to endolysosomal damage. This discussion is framed within a broader research thesis investigating the ESCRT pathway's role in repairing damage induced by lipid nanoparticles (LNPs), a critical consideration for therapeutic delivery and nanomedicine. Lysosomal membrane permeabilization (LMP) is a key event in cell stress, aging, and disease, and its precise repair is essential for cellular homeostasis.
Core Mechanisms: ESCRT and Lysophagy
The ESCRT Machinery for Membrane Repair
The ESCRT pathway facilitates the scission of membrane necks away from the cytoplasm. During endolysosomal damage, ESCRT-I and ESCRT-II components recognize ubiquitinated cargo and recruit ESCRT-III, which polymerizes to form spirals that constrict and cleave damaged membrane regions. The AAA-ATPase VPS4 catalyzes complex disassembly. This process seals small, selective holes without degrading the entire organelle.
Lysophagy for Organelle Clearance
Lysophagy is a selective autophagy pathway triggered by severe or irreparable damage. It involves the recognition of exposed lumenal glycoproteins (e.g., LAMP1, LAMP2) by cytosolic galectins (e.g., Galectin-3, Galectin-8). These recruit autophagy receptors (e.g., p62/SQSTM1, NDP52), which then engage the core autophagy machinery (ULK1 complex, LC3) to engulf the damaged lysosome within an autophagosome for delivery to a healthy lysosome for degradation.
Quantitative Comparison of Pathways
Table 1: Comparative Analysis of ESCRT-Mediated Repair vs. Lysophagy
| Feature | ESCRT-Mediated Repair | Lysophagy |
|---|---|---|
| Primary Trigger | Limited, localized membrane damage (pores, tears) | Extensive or irreparable membrane damage |
| Temporal Response | Rapid (minutes) | Delayed (minutes to hours) |
| Molecular Sensors | ESCRT-I/II (ubiquitin-sensing), Ca2+ influx | Galectin-3/Galectin-8 (glycan-sensing), K63-polyUb |
| Key Effectors | ESCRT-III (CHMP4), VPS4, ALIX | Autophagy receptors (p62, NDP52), LC3, ULK1 |
| Final Outcome | Restoration of organelle integrity | Clearance of damaged organelle |
| Energy Requirement | ATP (for VPS4) | ATP (for ubiquitination, autophagosome formation) |
| Fate of Luminal Content | Largely retained; minimal leakage | Degraded along with organelle |
| Role in LNP Research | Repair of LNP-induced membrane poration | Clearance of persistently damaged compartments from LNP cargo |
Table 2: Key Experimental Readouts for Pathway Activity
| Assay Type | ESCRT Repair Readout | Lysophagy Readout |
|---|---|---|
| Microscopy | Recruitment of GFP-CHMP4B to damaged lysosomes; retention of lysosomal dyes (e.g., LysoTracker). | Colocalization of Galectin-3 puncta with LAMP1 and LC3; formation of ubiquitin/LAMP1-positive autophagosomes. |
| Biochemical | Increased membrane association of ESCRT-III components; decreased release of luminal proteases (e.g., Cathepsin B) into cytosol. | Increased lipidated LC3-II; co-immunoprecipitation of p62 with LAMP1/Ubiquitin. |
| Functional | Recovery of lysosomal acidification post-damage; cell viability. | Lysosomal turnover assay (e.g., DQ-BSA degradation in autolysosomes); sensitivity to autophagy inhibition. |
Experimental Protocols
Protocol: Inducing and Quantifying Endolysosomal Damage in Cell Culture
Purpose: To model LNP-induced damage and initiate repair pathways.
- Cell Seeding: Seed HeLa or primary macrophages in 8-well chambered coverslips.
- Lysosomal Labeling: Incubate with 50 nM LysoTracker Red DND-99 and 1 mg/mL Dextran-FITC (10 kDa) for 4-16 hours. Chase in dye-free medium for 2 hours.
- Damage Induction: Treat cells with:
- LLOMe (Positive Control): 1-2 mM L-Leucyl-L-Leucine methyl ester for 30-60 min.
- Therapeutic LNPs: Titrated doses of mRNA-LNPs (e.g., 0.1-1 µg/mL mRNA equivalent) for 2-24 hours.
- Membrane Disruptors: 100 µM GPN for 15 min.
- Fixation and Imaging: Fix with 4% PFA, stain for markers (LAMP1, Galectin-3, CHMP4B), and image via confocal microscopy.
Protocol: Assessing ESCRT Recruitment by Live-Cell Imaging
Purpose: To visualize real-time ESCRT-III complex response to damage.
- Transfection: Transiently transfect cells with GFP-CHMP4B or GFP-ALIX using a lipid-based transfection reagent.
- Damage Induction: 24-48h post-transfection, add LLOMe (500 µM) or LNPs directly to the imaging medium while on a confocal microscope stage with environmental control (37°C, 5% CO2).
- Image Acquisition: Acquire time-lapse images (e.g., every 30 seconds for 30 minutes) at the GFP and lysosomal dye channels.
- Analysis: Quantify fluorescence intensity of GFP-CHMP4B at LysoTracker-positive structures over time using image analysis software (e.g., ImageJ).
Protocol: Measuring Lysophagic Flux
Purpose: To quantify the clearance of damaged lysosomes via autophagy.
- Inhibition & Induction: Pre-treat cells with 100 nM Bafilomycin A1 (inhibits autolysosomal degradation) or vehicle for 1 hour. Induce damage with LLOMe or LNPs for 2 hours.
- Cell Lysis: Lyse cells in RIPA buffer supplemented with protease inhibitors.
- Immunoblotting: Perform SDS-PAGE and Western blotting.
- Probing: Probe membranes with antibodies against:
- LC3B (to detect LC3-I and lipidated LC3-II).
- p62/SQSTM1 (autophagy substrate).
- β-Actin (loading control).
- Quantification: Calculate the LC3-II/LC3-I ratio and p62 levels. An increase in LC3-II with Bafilomycin A1 treatment indicates active lysophagic flux.
Pathway Diagrams (Graphviz/DOT)
Title: ESCRT-Mediated Repair of Damaged Lysosomes
Title: Lysophagy Pathway for Damaged Organelle Clearance
Title: Cellular Decision Logic for Repair vs. Clearance
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Studying Endolysosomal Damage Responses
| Reagent / Material | Function / Target | Primary Application |
|---|---|---|
| LLOMe (L-Leucyl-L-Leucine methyl ester) | Lysosomotropic detergent inducing selective LMP. | Positive control for robust endolysosomal damage. |
| GPN (Gly-Phe β-naphthylamide) | Substrate for cathepsin C, induces osmotic lysis. | Inducer of selective, rapid lysosomal rupture. |
| siRNA Pool (VPS4, CHMP4B) | Knocks down key ESCRT machinery components. | Validating functional role of ESCRT in repair assays. |
| Bafilomycin A1 | V-ATPase inhibitor; blocks autophagosome-lysosome fusion and acidification. | Measuring autophagic flux; inhibiting lysophagy. |
| Chloroquine | Lysosomotropic agent that raises lysosomal pH. | Inhibiting autophagic degradation; studying flux. |
| Galectin-3 Antibody (Clone M3/38) | Detects endogenous Galectin-3 puncta. | Immunofluorescence marker for lysosomal damage. |
| LC3B Antibody (D11) | Detects both LC3-I and LC3-II forms. | Western blot analysis of autophagosome formation. |
| p62/SQSTM1 Antibody | Detects autophagy receptor/substrate. | Assessing lysophagic clearance via immunoblot/IF. |
| Lysosensor Yellow/Blue DND-160 | Ratiometric pH-sensitive fluorescent dye. | Quantifying lysosomal pH recovery post-damage. |
| DQ-BSA Green/Red | Quenched BSA that fluoresces upon proteolytic cleavage. | Assaying lysosomal proteolytic activity and turnover. |
| GFP-CHMP4B Plasmid | Fluorescent tag for ESCRT-III core component. | Live-cell imaging of ESCRT recruitment to damage sites. |
This technical guide details the experimental approaches for in vitro biophysical validation of the Endosomal Sorting Complexes Required for Transport (ESCRT) pathway in membrane repair. The work is framed within a broader thesis investigating the ESCRT-mediated repair of damage to lipid nanoparticles (LNPs), a critical concern for the stability and efficacy of nucleic acid delivery systems in therapeutic applications. A fundamental understanding of how purified ESCRT components recognize and reseal membrane defects is essential for engineering more robust drug delivery vehicles and understanding related cellular repair processes.
The ESCRT Machinery: Core Components and Function
The ESCRT pathway is composed of five complexes (ESCRT-0, -I, -II, -III) and the VPS4-VTA1 ATPase. For in vitro repair assays, the core machinery typically includes:
- ESCRT-III subunits: CHMP4 (A, B, C), CHMP2 (A, B), CHMP3. These form spirals that constrict and cleave membrane necks.
- VPS4 ATPase: With its regulator VTA1, it disassembles ESCRT-III polymers, enabling recycling.
- Accessory proteins: ALIX and IST1 often facilitate recruitment and assembly.
Key In Vitro Membrane Repair Assays: Methodologies and Data
The following assays are used to quantitatively assess the membrane remodeling activity of purified ESCRT components.
Giant Unilamellar Vesicle (GUV) Leakage/Resealing Assay
This assay visualizes the repair of induced membrane defects on model membranes.
Experimental Protocol:
- GUV Formation: Prepare GUVs via electroformation in sucrose solution. Vesicles contain a self-quenched fluorescent dye (e.g., calcein at high concentration) or water-soluble fluorescent markers.
- Membrane Damage: Transfer GUVs to a glucose-based observation chamber (creating density difference for settling). Induce damage using:
- Pore-forming toxins (e.g., Streptolysin O).
- Laser ablation via a tightly focused pulsed laser.
- Mechanical perturbation.
- Repair Reaction: Introduce purified ESCRT components (typically ESCRT-III + VPS4) in an ATP/Mg²⁺-containing buffer.
- Quantification: Monitor fluorescence intensity change via confocal microscopy. Resealing is indicated by the cessation of dye leakage or the retention of internal marker.
Table 1: Representative Data from GUV Resealing Assays
| ESCRT Components Added | Damage Induction Method | % of GUVs Showing Resealing (Mean ± SD) | Typical Time to Resealing (s) | Key Reference Model |
|---|---|---|---|---|
| CHMP4B+CHMP2A+CHMP3+VPS4 | Laser Ablation | 78 ± 12 | 45 - 120 | Liposomes, Skowyra et al., 2018 |
| ALIX+IST1+CHMP4B+VPS4 | Streptolysin O Pores | 65 ± 9 | 90 - 300 | GUVs, Scheffer et al., 2021 |
| CHMP4B only (negative control) | Laser Ablation | 8 ± 5 | N/A | - |
| Buffer only (negative control) | Laser Ablation | 5 ± 4 | N/A | - |
Supported Membrane Tube Neck Fission Assay
This assay measures the direct membrane fission activity of ESCRT complexes, a key step in sealing a damaged membrane neck.
Experimental Protocol:
- Substrate Preparation: Create membrane tubes from a supported lipid bilayer (SLB) using motor proteins (kinesin) or by tethering GUVs to a substrate and pulling nanotubes.
- Complex Assembly: Flow in purified proteins (e.g., CHMP2A/CHMP3 nucleators followed by CHMP4B filaments).
- Fission Trigger: Introduce VPS4 ATPase with ATP to drive constriction and fission.
- Quantification: Use real-time fluorescence microscopy or atomic force microscopy (AFM) to visualize and measure the scission of the membrane tube.
Table 2: Quantitative Parameters from Membrane Fission Assays
| Parameter Measured | Experimental Condition | Typical Value / Observation | Measurement Technique |
|---|---|---|---|
| Fission Efficiency | Full ESCRT-III + VPS4 | 70-85% of tubes severed | Fluorescence microscopy |
| Filament Diameter | CHMP4B polymers on tubes | ~ 25 nm | Cryo-electron tomography |
| Fission Time | After VPS4/ATP addition | 30 - 180 seconds | Time-lapse AFM |
| ATP Hydrolysis Rate | VPS4 on CHMP4B arrays | ~ 50 min⁻¹ per hexamer | Malachite green phosphate assay |
Fluorescence Depolymerization Assay (VPS4 Activity)
This biochemical assay quantifies VPS4-driven disassembly of ESCRT-III polymers.
Experimental Protocol:
- Polymer Assembly: Incubate fluorescently labeled CHMP4B (or other subunit) in polymerization buffer (high salt).
- Baseline Measurement: Use fluorescence anisotropy or light scattering to measure polymer state.
- Disassembly Trigger: Add VPS4-VTA1 complex and ATP/Mg²⁺.
- Kinetics: Monitor the decrease in anisotropy/scattering over time, indicating polymer disassembly.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for In Vitro ESCRT Repair Assays
| Reagent/Material | Function/Purpose | Example Supplier/Note |
|---|---|---|
| Purified Recombinant ESCRT Proteins (CHMP4B, CHMP2A, VPS4, etc.) | Core enzymatic and structural components for reconstitution. | Often produced in-house; commercial from Abcam, Origene, etc. |
| Lipid Mixes (DOPC, DOPS, Cholesterol, PI(4,5)P₂) | Formulation of synthetic GUVs or SLBs with controlled composition. | Avanti Polar Lipids. PI(4,5)P₂ is critical for recruitment. |
| Calcein, CF Dye, or Similar Fluorophores | Content marker for GUV leakage/resealing assays. | Thermo Fisher, Sigma-Aldrich. |
| Streptolysin O or α-Hemolysin | Controlled induction of uniform membrane pores. | Sigma-Aldrich, Tocris. |
| ATP Regeneration System (ATP, Creatine Phosphate, Creatine Kinase) | Sustains VPS4 ATPase activity over long experiments. | Sigma-Aldrich. |
| Total Internal Reflection Fluorescence (TIRF) Microscope | High-contrast imaging of membrane-protein interactions. | Essential for tube fission assays. |
| Electroformation Chamber | Production of giant unilamellar vesicles (GUVs). | Custom or commercial (e.g., Nanion). |
| Malachite Green Phosphate Assay Kit | Quantifies ATP hydrolysis rates by VPS4. | Sigma-Aldrich, Abcam. |
Visualizations: Pathways and Workflows
ESCRT-Mediated Membrane Repair Pathway
GUV Leakage-Resealing Assay Workflow
This whitepaper, framed within a broader thesis on ESCRT-mediated repair of lipid nanoparticle (LNP) damage, provides a technical guide for comparing delivery efficacy between cells with functional and disrupted Endosomal Sorting Complex Required for Transport (ESCRT) pathways. The central premise is that ESCRT machinery actively repairs LNP-induced endosomal membrane damage, thereby influencing cargo release, lysosomal avoidance, and overall transfection efficiency. Quantifying delivery improvements in ESCRT-proficient versus deficient models is critical for optimizing nucleic acid therapeutics and understanding cellular resistance mechanisms.
Core Mechanisms: ESCRT Pathway in LNP Damage Repair
The ESCRT pathway (primarily ESCRT-III and VPS4 ATPase) is recruited to sites of endosomal membrane damage, including those caused by ionizable lipids in LNPs. It catalyzes the budding and scission of vesicles away from the cytoplasm, sealing small membrane ruptures. This repair prevents leakage of endosomal contents, acidification failure, and inflammasome activation, while also modulating the release of LNP cargo into the cytosol.
Diagram 1: ESCRT Recruitment to LNP-Damaged Endosome
Key Experimental Metrics for Comparative Efficacy
Quantitative measures must be captured in parallel for isogenic ESCRT-proficient and deficient cell lines (e.g., CRISPR knockout of CHMP4B, VPS4A, or TSG101).
Table 1: Primary Efficacy Metrics for Comparison
| Metric | ESCRT-Proficient Cells (Expected) | ESCRT-Deficient Cells (Expected) | Assay Method |
|---|---|---|---|
| Transfection Efficiency | High | Reduced (20-60% decrease) | Flow cytometry (reporter protein) |
| Cytosolic Cargo Delivery | Higher efficiency | Diminished | Gal8-YFP or Gal3-mCherry recruitment assay |
| Endosomal Damage Duration | Short-lived (<30 min) | Prolonged (>60 min) | Live imaging with damage biosensors |
| Lysosomal Entrapment | Low (10-30% of cargo) | High (40-70% of cargo) | Colocalization with LAMP1/2 |
| Cell Viability Post-Transfection | High (>85%) | Reduced (50-80%) | MTT or CellTiter-Glo |
| Inflammatory Response | Low IL-1β/IL-6 secretion | Elevated (2-5 fold increase) | ELISA of supernatant |
| Therapeutic Output | High protein/RNA effect | Low protein/RNA effect | Functional assay (e.g., gene correction) |
Detailed Experimental Protocols
Protocol 4.1: Generating ESCRT-Deficient Models via CRISPR-Cas9
Objective: Create stable CHMP4B or VPS4A knockout cell lines for comparison with parental lines.
- Design gRNAs targeting early exons of target gene (e.g., human CHMP4B exon 2).
- Clone gRNA into lentiviral vector (e.g., lentiCRISPRv2 with puromycin resistance).
- Produce lentivirus in HEK293T cells using psPAX2 and pMD2.G packaging plasmids.
- Transduce target cells (e.g., HeLa or HEPG2) at MOI ~0.3, select with 2 µg/mL puromycin for 72h.
- Single-cell clone by FACS or dilution cloning. Validate knockouts by western blot (anti-CHMP4B, anti-VPS4A) and Sanger sequencing of PCR-amplified genomic locus.
Protocol 4.2: Simultaneous Measurement of Transfection & Endosomal Damage
Objective: Quantify LNP delivery efficacy and membrane damage in real-time.
- Plate isogenic proficient and deficient cells in 96-well glass-bottom plates.
- Co-transfect with 50 ng/well of a Galectin-8-mCherry (damage sensor) plasmid and treat with Cy5-labeled siRNA LNPs (50 nM siRNA equivalent).
- At 2, 4, 6, 24 hours post-treatment, perform live confocal imaging (maintain 37°C/5% CO₂).
- Analysis: Use ImageJ/FIJI to quantify:
- % Transfected Cells: Cy5+ cytoplasm (threshold > 5x background).
- Damage Intensity: Mean mCherry fluorescence per Cy5+ endosome.
- Damage Kinetics: Time from Cy5 endosomal appearance to mCherry peak and decay.
Protocol 4.3: Functional Cargo Release Assay (Ribosome Delivery Assay)
Objective: Directly measure cytosolic delivery of mRNA-LNPs.
- Transfect cells with Cre-reporter mRNA LNPs (or firefly luciferase mRNA LNPs).
- For Cre assay: Use Ai9 tdTomato reporter cells. Cytosolic mRNA translation leads to Cre-mediated recombination and permanent tdTomato expression.
- For Luciferase assay: Lyse cells at 6, 12, 24 h and measure luminescence.
- Flow Cytometry (Cre assay): Harvest cells 48h post-treatment, analyze % tdTomato+ cells and MFI.
- Data Normalization: Normalize tdTomato+% or luminescence to a co-delivered inert fluorescent LNP (e.g., CellTrace) to account for uptake differences.
Diagram 2: Workflow for Comparative Efficacy Study
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for ESCRT-LNP Delivery Studies
| Reagent/Catalog # | Supplier (Example) | Function in Experiment |
|---|---|---|
| LentiCRISPRv2 (plasmid) | Addgene #52961 | CRISPR-Cas9 knockout of ESCRT genes. |
| Anti-CHMP4B / VPS4A Antibody | Cell Signaling, Abcam | Validation of knockout via western blot. |
| Galectin-8-mCherry Plasmid | Addgene #73355 | Live-cell sensor for endosomal damage. |
| LAMP1-GFP Plasmid | Addgene #34831 | Marker for lysosomal compartment. |
| Cy5-labeled siRNA | Dharmacon, Ambion | Fluorescent tracking of LNP cargo. |
| Ionizable Lipid (e.g., DLin-MC3-DMA) | Avanti Polar Lipids | Core component of benchmark LNP. |
| CellTiter-Glo 3D | Promega G9681 | Quantify cell viability post-LNP treatment. |
| Human IL-1β/IL-6 ELISA Kit | R&D Systems | Measure inflammatory response. |
| Ai9 Reporter Cell Line (B6;129S6-Gt(ROSA)26Sor |
Jackson Laboratory | Cre-recombinase-based delivery reporter. |
| Nucleofector Kit | Lonza | Efficient transfection of difficult cells. |
Comparative analysis will yield a Delivery Improvement Factor (DIF) for ESCRT-proficient cells, calculated as (Metric_Proficient / Metric_Deficient) across key parameters from Table 1. A DIF >1 indicates ESCRT-mediated enhancement. This quantitative framework directly tests the thesis that the ESCRT pathway is a major determinant of LNP delivery success. Understanding these mechanisms enables rational design of next-generation LNPs that either exploit or circumvent this cellular repair system to maximize therapeutic index.
The efficacy of lipid nanoparticles (LNPs) as delivery vehicles for nucleic acids and small molecules is often compromised by rapid clearance via the mononuclear phagocyte system (MPS). While PEGylation remains the benchmark stealth strategy to prolong circulation, it presents limitations, including anti-PEG immunity and reduced endosomal escape. This whitepaper benchmarks PEGylation against alternative and next-generation stealth strategies, analyzing potential synergies or antagonisms. This analysis is framed within a broader thesis investigating the ESCRT (Endosomal Sorting Complexes Required for Transport) pathway as a mechanism for repairing LNP membrane damage. A critical question is how surface engineering influences LNP interaction with serum proteins, cell membranes, and the subsequent recruitment of ESCRT components for repair, which may inadvertently promote lysosomal degradation over cytosolic delivery.
Stealth Strategies: Mechanisms & Current Data
Stealth coatings aim to create a hydrophilic, neutral, and dynamically stable "corona" to minimize opsonization. The following table summarizes key strategies and recent quantitative findings.
Table 1: Benchmarking of Stealth Coating Strategies for LNPs
| Strategy | Mechanism of Action | Key Advantage | Key Limitation | Reported Circulation Half-life (in vivo, mouse) | Impact on ESCRT Recruitment (Hypothesized) |
|---|---|---|---|---|---|
| PEGylation (Conventional) | Steric hindrance, reduced protein adsorption. | Gold standard, well-characterized, highly effective. | Accelerated Blood Clearance (ABC) upon repeat dosing, anti-PEG antibodies, can inhibit cellular uptake/fusion. | ~6-12 hrs (mRNA-LNPs) | May shield damage signals, potentially reducing ESCRT-III recognition. |
| PEG Alternatives (e.g., Poly(2-oxazoline)s, Polysarcosine) | Hydrophilic polymer brush similar to PEG. | Reduced immunogenicity, similar stealth performance. | Still a synthetic polymer; long-term fate unclear. | ~8-10 hrs (model LNPs) | Similar shielding effect as PEG. |
| "Stealth-Lipid" Incorporation (e.g., PEG-free stealth lipids) | Integrates stealth moiety directly into bilayer. | Homogeneous coating, avoids polymer dynamics. | Synthetic complexity, potential bilayer perturbation. | ~5-8 hrs (siRNA-LNPs) | Dependent on lipid chemical structure; may present specific damage motifs. |
| Biomimetic Coatings (e.g., CD47 "Don't Eat Me" signals) | Engagement of SIRPα on phagocytes to inhibit uptake. | Active biological inhibition of phagocytosis. | Complexity of functionalization, cost, potential immunogenicity. | >15 hrs (in specific tumor models) | Unlikely to directly affect ESCRT; focuses on immune evasion. |
| Dynamic/Strippable Coatings (e.g., pH-sheddable PEG) | PEG sheds in acidic endosome, promoting fusion. | Combines stealth with enhanced endosomal escape. | Increased formulation complexity, stability questions. | ~4-6 hrs (stealth phase only) | Critical for Thesis: Premature shedding may expose damage, enhancing ESCRT recruitment. Controlled shedding could synergize escape before repair. |
| Natural Protein Corona Pre-conditioning | Pre-coating with specific apolipoproteins (e.g., ApoE). | Exploits natural trafficking pathways (e.g., to liver). | Target specificity is dictated by corona, not fully controllable. | Varies by protein | ApoE-mediated uptake may direct to distinct endosomal pathways with different ESCRT activity. |
Experimental Protocols for Benchmarking & ESCRT Analysis
Protocol:In VitroProtein Corona Analysis & Opsonization
Objective: Quantify and qualify the protein corona formed on LNPs with different stealth coatings in human plasma. Method:
- LNP Preparation: Formulate identical core LNPs (e.g., siRNA against luciferase) and vary only the stealth coating: (a) DSPE-PEG2000, (b) Poly(2-oxazoline)-lipid, (c) pH-sheddable PEG-lipid, (d) No coating.
- Incubation: Incubate LNPs (100 µg lipid) with 1 mL of 50% human platelet-poor plasma in PBS for 1 hr at 37°C.
- Isolation: Separate protein-corona-coated LNPs from unbound proteins via size-exclusion chromatography (e.g., Sepharose CL-4B) or ultracentrifugation (100,000 g, 4°C, 1 hr).
- Analysis:
- Quantification: Use a microBCA assay on dissolved pellets to calculate total protein bound per mg lipid.
- Qualification: Elute proteins from the pellet, perform tryptic digestion, and analyze via LC-MS/MS for proteomic identification.
Protocol:In VivoPharmacokinetics and Biodistribution
Objective: Compare circulation time and organ accumulation of stealth-engineered LNPs. Method:
- Labeling: Incorporate a trace amount of a lipophilic near-infrared dye (e.g., DiR) into the LNP bilayer during formulation for each stealth type.
- Administration: Inject mice intravenously (n=5 per group) with LNPs (3 mg lipid/kg).
- Imaging: Use IVIS imaging at time points (5 min, 1, 2, 4, 8, 24 hrs) to track fluorescence in circulation (via tail vein scan) and major organs.
- Quantification: At terminal time points (e.g., 1 hr and 24 hrs), harvest organs, homogenize, and quantify dye fluorescence to calculate % injected dose per gram (%ID/g).
Protocol: Assessing ESCRT Recruitment in Response to LNP Damage
Objective: Determine if stealth coatings modulate the recruitment of ESCRT-III proteins (e.g., CHMP4B) to damaged endosomal membranes containing LNPs. Method:
- Cell Engineering: Stably transduce HeLa cells with a fluorescent reporter for CHMP4B (e.g., CHMP4B-GFP).
- LNP Treatment: Treat cells with LNPs (same set as 3.1) loaded with a cargo that induces membrane perturbation (e.g., ionizable lipid at endosomal pH).
- Live-Cell Imaging: Use confocal microscopy at 37°C, 5% CO₂. Image cells 15-60 minutes post-LNP addition.
- Quantification: Measure the co-localization coefficient (Manders' or Pearson's) between the CHMP4B-GFP puncta and a fluorescent label on the LNP (e.g., a Rhodamine-PE lipid). Compare frequency and intensity of CHMP4B recruitment across LNP types.
Visualization of Pathways and Workflows
Title: Stealth Coating Impact on LNP Endosomal Fate and ESCRT Recruitment
Title: Integrated Benchmarking Protocol for Stealth LNPs
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Stealth LNP & ESCRT Research
| Item | Function/Description | Example Supplier/Cat. No. (Illustrative) |
|---|---|---|
| Ionizable Cationic Lipid | Core component of modern LNPs; enables nucleic acid encapsulation and endosomal escape. | SM-102 (Avanti), DLin-MC3-DMA (MedKoo). |
| PEGylated Lipids (Variants) | Provide steric stabilization. Critical for benchmarking. | DSPE-PEG2000 (Avanti 880120), PEG-DMG (Avanti 880151), pH-sheddable PEG-lipid (Custom synth.). |
| Poly(2-oxazoline)-conjugated Lipid | Alternative stealth polymer for head-to-head comparison with PEG. | Custom synthesis from companies like Seriox. |
| Fluorescent Lipid Probes | For tracking LNP location in cellular and in vivo studies. | Rhodamine-DOPE (Invitrogen L1392), DiR iodide (Invitrogen D12731). |
| CHMP4B-GFP Plasmid | Reporter for visualizing ESCRT-III complex recruitment to sites of membrane damage. | Addgene plasmid #138009 (from Wesley Sundquist lab). |
| Human ApoE3 Protein, Recombinant | For pre-conditioning studies to create a targeted protein corona. | PeproTech 350-02. |
| Size-Exclusion Chromatography Columns | For isolating protein-corona-coated LNPs from plasma. | Sepharose CL-4B (Cytiva 17015001). |
| MicroBCA Protein Assay Kit | For quantifying total protein bound to LNPs in corona studies. | Thermo Fisher Scientific 23235. |
| Near-Infrared Lipophilic Dye (DiR) | For non-invasive in vivo imaging of LNP biodistribution. | PerkinElmer NEI100001. |
This analysis is framed within the broader thesis that the Endosomal Sorting Complexes Required for Transport (ESCRT) machinery is a critical cellular repair system that mitigates lipid nanoparticle (LNP) membrane damage, thereby directly modulating the efficiency of mRNA delivery and translation. For clinically approved LNP-mRNA vaccines, the dynamic interplay between LNP endosomal escape and ESCRT-mediated membrane sealing is a pivotal, yet underexplored, determinant of therapeutic efficacy.
ESCRT Machinery: A Primer for LNP Research
The ESCRT pathway (comprising complexes ESCRT-0, -I, -II, -III, and VPS4 ATPase) is traditionally responsible for multivesicular body formation, cytokinetic abscission, and, most relevantly, the repair of damaged cellular membranes. It is recruited to sites of membrane injury to catalyze the pinching-off and remodeling of lipid bilayers.
Key Hypothesis in LNP-mRNA Delivery: LNPs, often incorporating ionizable lipids, induce transient endosomal membrane destabilization to release mRNA into the cytosol. This "damage" may simultaneously recruit ESCRT components. The kinetics of ESCRT recruitment versus the completion of escape determines the fate of the mRNA cargo—successful cytosolic delivery versus sequestration/degradation within endolysosomal compartments.
Table 1: Impact of ESCRT Component Modulation on LNP-mRNA Delivery Efficacy In Vitro
| ESCRT Target | Modulation Method | Effect on mRNA Expression | Proposed Mechanism | Key Reference |
|---|---|---|---|---|
| TSG101 (ESCRT-I) | siRNA Knockdown | Increase (2-5 fold) | Delayed endosomal membrane repair, prolonged escape window. | Gee et al., 2020 |
| CHMP4B (ESCRT-III) | Dominant-Negative Overexpression | Increase (~3 fold) | Inhibition of scission activity, preventing vesicle sealing. | |
| VPS4 ATPase | Pharmacological Inhibition (ML-SI1) | Increase (1.5-4 fold) | Trapping of ESCRT-III polymers, stalling repair. | |
| ALIX | siRNA Knockdown | Variable/Decrease | Potential disruption of late endosomal trafficking, confounding effects. |
Table 2: Correlation Between LNP Formulation Properties & ESCRT Recruitment
| LNP Ionizable Lipid pKa | Membrane Destabilization Potential | ESCRT Recruitment (Measured by CHMP4B-GFP puncta) | Relative Protein Expression Output |
|---|---|---|---|
| ~6.2 (e.g., DLin-MC3-DMA) | Moderate | High | Baseline (High) |
| <6.0 | High | Very High | Reduced (Potential excessive damage/lysosomal capture) |
| >6.5 | Low | Low | Reduced (Inefficient escape) |
Experimental Protocols for Key Investigations
Protocol 1: Visualizing ESCRT Recruitment to LNP-Laden Endosomes
- Objective: Co-localize ESCRT components with internalized LNPs in live cells.
- Methodology:
- Cell Preparation: Seed HeLa or HEK293 cells expressing fluorescently tagged ESCRT protein (e.g., CHMP4B-mCherry).
- LNP Labeling: Incubate cells with DiD-labeled, mRNA-loaded LNPs (37°C, 5% CO2) for 15-30 minutes.
- Live-Cell Imaging: Use confocal microscopy with a climate-controlled chamber. Acquire time-lapse images every 30 seconds for 60 minutes post-LNP addition.
- Analysis: Quantify co-localization coefficients (e.g., Pearson's) between DiD (LNP) and mCherry (ESCRT) signals in endosomal puncta over time.
Protocol 2: Functional Assessment via ESCRT Inhibition
- Objective: Measure the effect of ESCRT dysfunction on LNP-mRNA-mediated protein expression.
- Methodology:
- Pre-treatment: Treat cells with VPS4 inhibitor ML-SI1 (5 µM) or transfection with TSG101-specific siRNA 24-48 hours prior to LNP addition.
- LNP Transfection: Add firefly luciferase-encoding LNP-mRNA at optimized dose.
- Readout: Harvest cells 6-24 hours post-transfection. Perform luciferase assay; normalize to total protein or cell viability (e.g., ATP-based assay).
- Controls: Include untreated cells and cells treated with non-targeting siRNA or DMSO vehicle.
Visualization of Pathways and Workflows
Title: ESCRT & LNP Fate Decision Pathway
Title: Experimental Workflow for ESCRT-LNP Studies
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Investigating ESCRT in LNP Biology
| Reagent/Material | Function/Application | Example Product/Catalog |
|---|---|---|
| Ionizable Lipid LNPs | Core test vehicle; variable pKa allows tuning of membrane interaction. | Proprietary formulations (e.g., based on DLin-MC3-DMA, SM-102) or commercial reagent (e.g., Precision NanoSystems' NanoAssemblr). |
| Fluorescent Lipophilic Dyes (DiD, DiI) | Labeling LNP lipid bilayer for live-cell tracking. | Thermo Fisher Scientific, DiIC18(5) (DiD) (D7757). |
| ESCRT Expression Constructs | Visualizing ESCRT recruitment dynamics. | Addgene plasmids: CHMP4B-mCherry (#71845), TSG101-GFP. |
| ESCRT Inhibitors | Chemically disrupting ESCRT function. | VPS4 inhibitor: ML-SI1 (Sigma, SML1527); ESCRT-III modulator: Spirohexenolide A. |
| siRNA Pools (TSG101, CHMP2/3/4, VPS4) | Knocking down specific ESCRT components. | Dharmacon ON-TARGETplus SMARTpools. |
| Reporter mRNA | Quantifying functional delivery output. | CleanCap FLuc mRNA (TriLink BioTechnologies). |
| Advanced Cell Imaging Media | Maintaining cell health during long-term live imaging. | Phenol-red free media (Gibco, FluoroBrite DMEM). |
| Confocal Microscopy w/ Live-Cell Chamber | Capturing high-resolution, time-lapse co-localization. | Systems from Nikon, Zeiss, or Leica with CO2/ temperature control. |
Conclusion
The ESCRT pathway represents an essential, endogenous cellular defense mechanism that can be strategically co-opted to improve lipid nanoparticle delivery. From foundational mechanisms to applied methodologies, evidence confirms that controlled, repairable membrane damage—orchestrated through ESCRT recruitment—facilitates endosomal escape and enhances functional payload delivery. However, optimization requires careful balancing to avoid pathway saturation and cytotoxicity. Future directions point toward the rational design of 'ESCRT-tuned' LNPs with lipid chemistries that predictably modulate the degree and timing of membrane damage. Furthermore, combining ESCRT engagement with targeting ligands or endosomolytic agents presents a promising multi-pronged strategy. Integrating this understanding into LNP design pipelines holds significant potential to overcome a major translational barrier, advancing the efficacy of RNA therapeutics, gene editing tools, and vaccines.